Targeting Cancer Pathways: RNAi Suppression of Carotenoid Biosynthesis in Oncology Research and Therapeutics
This article provides a comprehensive analysis of RNA interference (RNAi) as a strategic tool for suppressing the carotenoid biosynthesis pathway, with implications for cancer metabolism and drug discovery.
Targeting Cancer Pathways: RNAi Suppression of Carotenoid Biosynthesis in Oncology Research and Therapeutics
Abstract
This article provides a comprehensive analysis of RNA interference (RNAi) as a strategic tool for suppressing the carotenoid biosynthesis pathway, with implications for cancer metabolism and drug discovery. We explore the foundational science linking carotenoid intermediates to oncogenic signaling and cell proliferation. The content details current methodological approaches for designing and delivering RNAi constructs against key pathway enzymes (e.g., BCO1, BCMO1), including siRNA, shRNA, and CRISPR-based methods. We address common challenges in specificity, off-target effects, and delivery optimization. Finally, we compare RNAi efficacy to pharmacological inhibitors and gene knockout models, validating its utility as a precise research tool and its potential therapeutic relevance. This resource is tailored for researchers and drug development professionals seeking to modulate metabolic pathways in cancer.
The Science of Carotenoid Metabolism: Why Targeting This Pathway Matters in Cancer Biology
The carotenoid biosynthesis pathway is a critical metabolic route in plants, algae, and certain bacteria and fungi, responsible for producing pigments essential for photosynthesis, photoprotection, and the synthesis of apocarotenoid signaling molecules. Research into RNA interference (RNAi) suppression of this pathway offers a powerful tool for elucidating gene function, metabolic flux control, and the development of biofortified crops or therapeutic agents. This whitepaper details the core enzymatic steps, key metabolites, and experimental approaches relevant to RNAi-based research in this field.
The Core Pathway: Enzymes and Metabolic Intermediates
Carotenoid biosynthesis originates from the central isoprenoid precursor, isopentenyl diphosphate (IPP), and its isomer dimethylallyl diphosphate (DMAPP). The pathway proceeds through a series of condensation, desaturation, cyclization, and oxygenation reactions.
Early Steps: Formation of Lycopene
The initial steps commit IPP and DMAPP to carotenoid production.
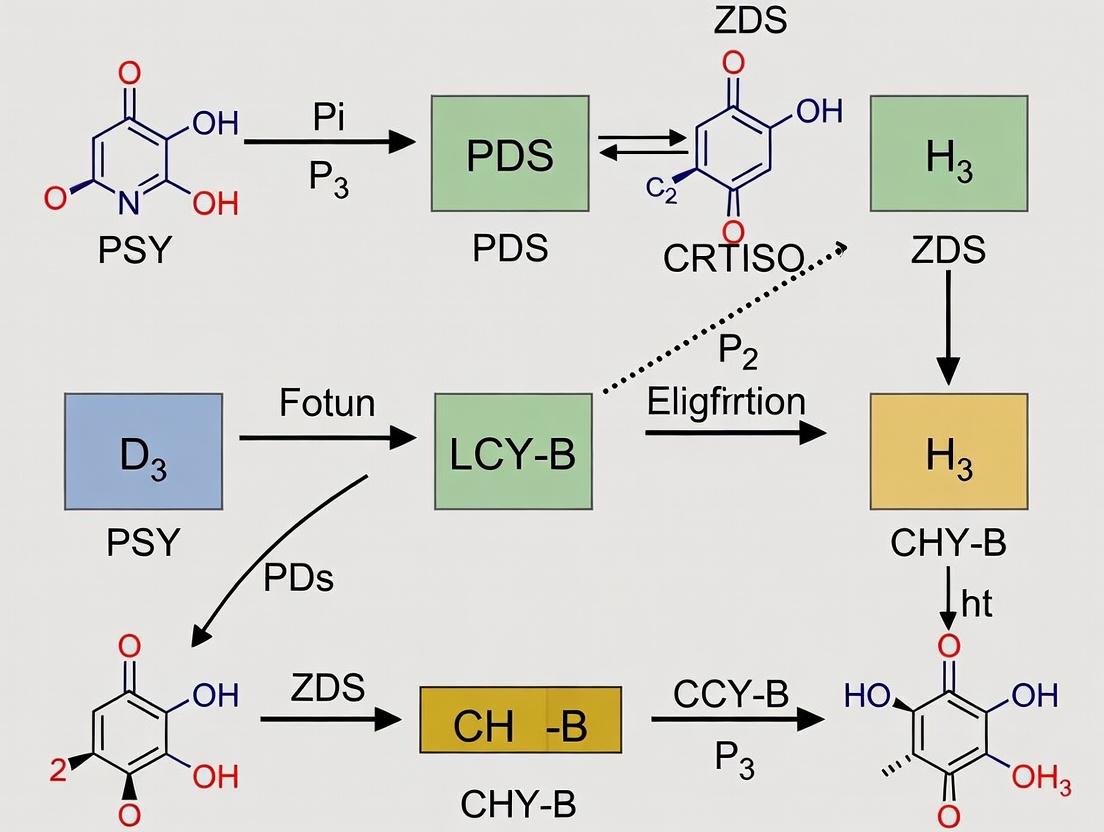
- Phytoene Synthase (PSY): Condenses two molecules of geranylgeranyl diphosphate (GGPP) to form 15-cis-phytoene, the first committed, colorless carotenoid.
- Phytoene Desaturase (PDS) & ζ-Carotene Desaturase (ZDS): Catalyze a series of desaturation reactions introducing double bonds, converting phytoene to lycopene via intermediates including ζ-carotene.
- Carotenoid Isomerase (CRTISO): Converts poly-cis-lycopene to all-trans-lycopene, a critical step for proper cyclization.
Branching Pathways: Cyclization and Diversification
All-trans-lycopene is the substrate for two key cyclase enzymes, leading to the α- and β-branches.
- Lycopene β-Cyclase (LCY-B): Introduces two β-rings, forming β-carotene.
- Lycopene ε-cyclase (LCY-E): Introduces one ε-ring. Combined with LCY-B, it produces α-carotene.
- β-Carotene Hydroxylase (BCH) & Cytochrome P450 Hydroxylases (CYP97): Hydroxylate the β- and ε-rings of α- and β-carotene to produce lutein and zeaxanthin, respectively.
- Zeaxanthin Epoxidase (ZEP) & Violaxanthin De-epoxidase (VDE): Form the violaxanthin cycle, crucial for photoprotection (non-photochemical quenching).
- Carotenoid Cleavage Dioxygenases (CCDs): Cleave specific double bonds in carotenoids to produce apocarotenoids like abscisic acid (ABA) and strigolactones.
Table 1: Key Enzymes and Their Primary Products in the Carotenoid Pathway
| Enzyme (Abbreviation) | EC Number | Reaction Catalyzed | Primary Product(s) |
|---|---|---|---|
| Phytoene Synthase (PSY) | 2.5.1.32 | Condensation of 2 GGPP | 15-cis-Phytoene |
| Phytoene Desaturase (PDS) | 1.3.99.31 | Desaturation | 9,15,9'-Tri-cis-ζ-Carotene |
| ζ-Carotene Desaturase (ZDS) | 1.3.99.30 | Desaturation | 7,9,7',9'-Tetra-cis-Lycopene |
| Carotenoid Isomerase (CRTISO) | 5.2.1.13 | Cis-to-trans isomerization | All-trans-Lycopene |
| Lycopene β-Cyclase (LCY-B) | 5.5.1.19 | β-ring cyclization | β-Carotene |
| Lycopene ε-cyclase (LCY-E) | 5.5.1.18 | ε-ring cyclization | δ-Carotene → α-Carotene (with LCY-B) |
| β-Carotene Hydroxylase (BCH) | 1.14.13.- | Hydroxylation of β-rings | Zeaxanthin (from β-carotene) |
| Zeaxanthin Epoxidase (ZEP) | 1.14.13.90 | Epoxidation | Violaxanthin (via Antheraxanthin) |
| 9-cis-Epoxycarotenoid Dioxygenase (NCED) | 1.13.11.51 | Cleavage of 9-cis-violaxanthin/neoxanthin | Xanthoxin (ABA precursor) |
Table 2: Major Carotenoid Metabolites and Their Functions
| Metabolite | Type | Key Functions in Organisms |
|---|---|---|
| Phytoene | Linear carotene | Colorless precursor; antioxidant in human diet. |
| Lycopene | Linear carotene | Red pigment; potent antioxidant. |
| β-Carotene | Cyclic carotene (β,β) | Provitamin A; antioxidant; photosynthetic pigment. |
| α-Carotene | Cyclic carotene (β,ε) | Provitamin A; minor light-harvesting pigment. |
| Lutein | Xanthophyll (β,ε diol) | Major photoprotective pigment in LHCII; macular pigment in humans. |
| Zeaxanthin | Xanthophyll (β,β diol) | Central in photoprotection (quenching); macular pigment. |
| Violaxanthin | Xanthophyll (epoxide) | Substrate for ABA biosynthesis; component of violaxanthin cycle. |
| Abscisic Acid (ABA) | Apocarotenoid | Plant hormone (stress response, dormancy). |
RNAi Suppression in Carotenoid Pathway Research
RNAi is used to selectively silence target genes, creating knockdown phenotypes to study gene function and pathway regulation. In carotenoid research, this is pivotal for:
- Functional Genomics: Validating enzyme function in non-model organisms.
- Metabolic Engineering: Identifying rate-limiting steps and regulatory nodes.
- Trait Manipulation: Altering pigment composition for nutritional (biofortification) or agronomic traits.
- Therapeutic Discovery: Understanding the role of carotenoid-derived metabolites in signaling, with implications for drug targets in areas like inflammation or cell proliferation.
Experimental Protocol: RNAi-Mediated Gene Suppression in Plant Leaves
This protocol outlines transient Agrobacterium-mediated RNAi suppression (Virus-Induced Gene Silencing, VIGS) in Nicotiana benthamiana.
Materials: See "The Scientist's Toolkit" below. Procedure:
- Target Gene Fragment Cloning: Design gene-specific primers (~300-500 bp) with appropriate restriction overhangs. Amplify fragment from cDNA. Digest PCR product and pTRV2 vector with corresponding restriction enzymes. Ligate fragment into pTRV2.
- Agrobacterium Transformation: Transform ligated plasmid into Agrobacterium tumefaciens strain GV3101 via electroporation. Select colonies on LB plates with appropriate antibiotics (kanamycin, rifampicin, gentamicin).
- Agroinfiltration Culture Preparation: Inoculate a single colony into 5 mL LB medium with antibiotics. Grow overnight at 28°C, 250 rpm. Subculture 1:50 into fresh induction medium (LB with antibiotics, 10 mM MES pH 5.6, 20 μM acetosyringone). Grow to OD600 ~1.5. Pellet cells and resuspend in infiltration buffer (10 mM MgCl2, 10 mM MES pH 5.6, 150 μM acetosyringone) to a final OD600 of 1.0. Incubate at room temperature for 3-4 hours.
- Plant Infiltration: Mix the Agrobacterium containing pTRV1 (RNA-dependent RNA polymerase) 1:1 with the culture containing pTRV2 (target gene insert). Using a needleless syringe, infiltrate the mixture into the abaxial side of 3-4 leaf-stage N. benthamiana leaves.
- Plant Growth and Sampling: Maintain plants under standard conditions (22-24°C, 16-h light/8-h dark). Silencing phenotypes (e.g., photo-bleaching for PDS control) typically appear in new growth 2-3 weeks post-infiltration.
- Validation and Analysis:
- Molecular: Confirm gene knockdown via qRT-PCR on leaf tissue using gene-specific primers and a stable reference gene (e.g., EF1α).
- Metabolite: Extract carotenoids from silenced leaf discs (e.g., 100 mg fresh weight) using a solvent (e.g., acetone:hexane 1:1). Analyze via HPLC-PDA/MS using a C30 reverse-phase column (e.g., YMC Carotenoid column) with gradients of methanol/MTBE/water. Quantify against authentic standards.
Diagram 1: RNAi-VIGS Workflow for Carotenoid Gene Silencing
Diagram 2: Carotenoid Biosynthesis Core Pathway Overview
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for RNAi-Carotenoid Experiments
| Reagent / Material | Function / Purpose | Example / Note |
|---|---|---|
| VIGS Vectors (pTRV1, pTRV2) | Plant virus-derived vectors for efficient, transient gene silencing. | TRV-based system for N. benthamiana. |
| Agrobacterium tumefaciens GV3101 | Strain for efficient plant transformation and delivery of T-DNA containing RNAi construct. | Requires appropriate helper plasmids. |
| Acetosyringone | Phenolic compound inducing Agrobacterium vir genes for T-DNA transfer. | Critical for agroinfiltration efficiency. |
| C30 Reversed-Phase HPLC Column | Specialized column for optimal separation of geometric and structural carotenoid isomers. | YMC Carotenoid, 3 µm, 150 x 4.6 mm. |
| Carotenoid Authentic Standards | Essential for identifying and quantifying metabolites via HPLC-PDA/MS. | e.g., β-carotene, lutein, zeaxanthin, lycopene from commercial suppliers. |
| qRT-PCR Master Mix with Reverse Transcriptase | For quantifying target gene mRNA levels to confirm silencing efficiency. | One-step mixes reduce contamination risk. |
| RNA Isolation Kit (Plant) | High-quality RNA extraction, removing polysaccharides and phenolics. | Includes DNase I treatment step. |
| Solid Phase Extraction (SPE) Cartridges | Clean-up and concentration of carotenoid extracts prior to HPLC. | C18 or Diol-phase cartridges. |
| Infiltration Buffer (MgCl2/MES) | Resuspension medium for Agrobacterium during infiltration, maintaining cell viability and virulence. | pH is critical (typically 5.6-5.8). |
This whitepaper examines the signaling roles of carotenoid-derived metabolites in cellular decision-making processes. This analysis is situated within a broader thesis investigating the molecular consequences of RNA interference (RNAi)-mediated suppression of the carotenoid biosynthetic pathway in eukaryotic model systems. By strategically silencing key enzymes (e.g., PSY, BCO1), we can deplete precursor pools for retinoids and apocarotenoids, enabling the dissection of their specific contributions to gene regulation, cell cycle progression, and lineage specification. This approach moves beyond correlation to establish causative links between metabolite availability and phenotypic outcomes in proliferation and differentiation.
Key Metabolite Classes and Their Origins
Carotenoid cleavage, mediated by specific enzymes, yields bioactive derivatives:
- Retinoids: Primarily derived from provitamin A carotenoids (e.g., β-carotene) via central cleavage by BCO1, yielding retinaldehyde, which is further oxidized to retinoic acid (RA). RA is the primary signaling molecule, acting as a ligand for nuclear receptors.
- Apocarotenoids: Generated via symmetric or asymmetric cleavage of both provitamin A and non-provitamin A carotenoids (e.g., lycopene, lutein) by enzymes like BCO2 and CCDs. These include β-apo-carotenals, -carotenoic acids, and diapocarotenoids, which can function as signaling molecules through alternative or poorly characterized nuclear receptors and other targets.
Quantitative Data on Metabolite Effects
Table 1: Concentration-Dependent Effects of Retinoids on Cell Fate In Vitro
| Metabolite | Cell Type/Line | Concentration Range (nM) | Effect on Proliferation | Effect on Differentiation | Key Regulated Genes/Pathways | Citation (Example) |
|---|---|---|---|---|---|---|
| all-trans Retinoic Acid (ATRA) | HL-60 (Myeloid) | 10 - 1000 | Inhibition, cell cycle arrest | Induction of granulocytic differentiation | RARβ, C/EBPε, p21^CIP1 | Breitman et al., 1980 |
| 9-cis Retinoic Acid | Embryonic Stem Cells (mESC) | 1 - 100 | Mild inhibition | Promotes neural precursor formation | RXR, Sox1, Pax6 | Janesick et al., 2015 |
| all-trans Retinaldehyde | SH-SY5Y (Neuroblastoma) | 100 - 10000 | Biphasic (low: promote; high: inhibit) | Induces neurite outgrowth | ALDH1A2, TrkB | Aoto et al., 2008 |
| β-Apo-14'-carotenoic Acid | Adipocyte Progenitors | 100 - 5000 | No direct effect | Potently inhibits adipogenesis | PPARγ, FABP4 | Landrier et al., 2012 |
Table 2: Apocarotenoid Signaling Outcomes in Proliferation Models
| Apocarotenoid | Biosynthetic Origin (Carotenoid) | Experimental Model | Observed Effect | Proposed Primary Target | Functional Outcome |
|---|---|---|---|---|---|
| β-Apo-13-carotenone | β-carotene (asymmetric) | Hepatocarcinoma (HepG2) | Anti-proliferative, pro-apoptotic | RARγ? / NRF2 pathway | Cell cycle arrest (G1/S) |
| Lycopenal | Lycopene | Benign Prostate Hyperplasia | Reduction of proliferation markers | IGF-1 signaling axis | Downregulation of Ki-67, PCNA |
| Croce tin | Zeaxanthin (degradation) | Neuronal Stem Cells (NSC) | Enhances proliferation | PI3K/Akt & ERK pathways | Expansion of NSC pool |
| Diapocaroten-dioic acid | Multiple | Colorectal Cancer Cells | Potent inhibition of colony formation | RAR-independent, unknown | Loss of clonogenic potential |
Diagram Title: Core Signaling Pathways from Carotenoids to Cell Fate
Detailed Experimental Protocols
Protocol: RNAi-Mediated Knockdown of BCO1 to Probe Retinoid-Specific Signaling
Objective: To specifically inhibit retinoid synthesis and assess the resultant phenotypic and transcriptomic changes in a differentiating cell model.
Materials: See "Scientist's Toolkit" (Section 7).
Procedure:
- Cell Seeding: Seed adherent target cells (e.g., SH-SY5Y neuroblastoma) in 6-well plates at 60% confluence in standard growth medium. Incubate overnight.
- Transfection Complex Preparation:
- For each well, dilute 5 pmol of validated BCO1-specific siRNA or non-targeting scrambled siRNA control in 250 µL of serum-free Opti-MEM.
- In a separate tube, dilute 7.5 µL of Lipofectamine RNAiMAX in 250 µL of serum-free Opti-MEM. Incubate both for 5 minutes at RT.
- Combine the diluted siRNA with the diluted Lipofectamine. Mix gently and incubate for 20 minutes at RT to allow complex formation.
- Transfection: Add the 500 µL transfection complex drop-wise to each well containing 2 mL of fresh, antibiotic-free growth medium. Swirl gently.
- Incubation & Validation: Incubate cells for 48-72 hours at 37°C, 5% CO₂.
- Harvest cells for qPCR validation of BCO1 mRNA knockdown (≥70% recommended).
- Confirm reduction in intracellular retinoic acid levels via LC-MS/MS (see protocol 5.3).
- Differentiation Induction: 48h post-transfection, switch medium to differentiation-inducing medium (e.g., containing low serum, 10 µM retinoic acid, or other morphogens).
- Phenotypic Analysis (72-96h post-differentiation):
- Proliferation: Perform BrdU incorporation assay or directly count cells using a hemocytometer/automated counter.
- Differentiation: Fix cells and immunostain for lineage-specific markers (e.g., β-III-tubulin for neurons). Quantify neurite outgrowth using image analysis software (e.g., ImageJ NeuriteTracer).
- Downstream Analysis: Perform RNA-seq or qPCR arrays on RA-target genes (Cyp26a1, RARβ, Stra6) to confirm pathway attenuation.
Protocol: Assessing Apocarotenoid Activity via Reporter Assay
Objective: To test the ability of specific apocarotenoids to activate candidate nuclear receptor pathways.
Procedure:
- Reporter Cell Line Preparation: Culture cells (e.g., HEK293T) stably or transiently transfected with a luciferase reporter plasmid under the control of a response element for the receptor of interest (e.g., PPAR Response Element, PPRE).
- Compound Treatment: Seed reporter cells in 96-well white-walled plates. At 80% confluence, treat with a dose range (e.g., 10 nM – 10 µM) of the synthetic apocarotenoid (e.g., β-apo-14'-carotenoic acid). Include controls: vehicle (DMSO, <0.1%), a known receptor agonist (e.g., Rosiglitazone for PPARγ), and ATRA (for RAR-specific control).
- Incubation: Incubate for 24 hours.
- Luciferase Assay: Aspirate medium, lyse cells with 1X Passive Lysis Buffer (Promega) for 15 minutes with rocking. Transfer lysate to a new plate if necessary. Inject Luciferase Assay Reagent and measure luminescence immediately on a plate reader.
- Data Analysis: Normalize luminescence of treated wells to vehicle control. Plot dose-response curves to calculate EC₅₀ values for receptor activation.
Protocol: LC-MS/MS Quantification of Retinoids and Apocarotenoids
Objective: To accurately measure endogenous levels of carotenoid-derived metabolites following genetic or chemical perturbation.
Procedure:
- Sample Preparation: Harvest ~1x10⁶ cells, wash with cold PBS, and pellet. Extract metabolites by adding 500 µL of ice-cold methanol containing internal standards (e.g., d₄-ATRA, d₆-retinol). Vortex vigorously for 1 min, sonicate on ice for 5 min, then centrifuge at 15,000 x g for 10 min at 4°C.
- Solid-Phase Extraction (SPE): Load supernatant onto a pre-conditioned C18 SPE column. Wash with 3 mL water, then elute analytes with 1 mL of ethyl acetate containing 0.1% BHT. Evaporate eluent under a gentle stream of nitrogen.
- Reconstitution: Reconstitute the dry residue in 100 µL of methanol/dichloromethane (50:50, v/v) for LC-MS/MS analysis.
- LC-MS/MS Parameters:
- Column: C30 reversed-phase column (e.g., YMC Carotenoid, 3 µm, 150 x 2.1 mm).
- Mobile Phase: A: Methanol/Water (95:5) with 10 mM ammonium acetate; B: Methyl tert-butyl ether (MTBE). Gradient from 0% B to 100% B over 20 min.
- MS: Operate in positive electrospray ionization (ESI+) mode with multiple reaction monitoring (MRM). Specific transitions for ATRA (301.1 > 205.1), retinol (269.2 > 93.1), and target apocarotenoids.
Experimental Workflow Diagram
Diagram Title: Workflow for RNAi-Based Functional Metabolomics
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for RNAi-Carotenoid Signaling Research
| Reagent / Material | Function / Purpose | Example Product / Cat. No. (for reference) |
|---|---|---|
| Validated siRNA Pools | Targeted knockdown of carotenogenic or cleavage enzymes (e.g., PSY1, BCO1, BCO2, CCD7). Ensures specific and potent mRNA degradation. | Dharmacon ON-TARGETplus SMARTpools; Qiagen FlexiTube siRNA |
| Lipofectamine RNAiMAX | Cationic lipid transfection reagent optimized for high-efficiency siRNA delivery with low cytotoxicity in a wide range of mammalian cells. | Thermo Fisher Scientific, 13778075 |
| all-trans Retinoic Acid (ATRA) | Gold-standard RAR ligand. Used as positive control in differentiation assays and for rescue experiments following carotenoid pathway knockdown. | Sigma-Aldrich, R2625 |
| Synthetic Apocarotenoids | Chemically defined standards for treatment/rescue experiments and as analytical standards for LC-MS/MS. | Cayman Chemical (e.g., β-apo-13-carotenone, 21873) |
| Deuterated Internal Standards (d-IS) | Essential for accurate quantitative LC-MS/MS. Corrects for analyte loss during extraction and matrix effects. | Toronto Research Chemicals (e.g., d4-ATRA, A862911) |
| C30 Reversed-Phase HPLC Column | Specialized column for optimal separation of geometric isomers of retinoids and apocarotenoids, which have similar mass spectra. | YMC Carotenoid Column (YMC30) |
| Retinoid/Apocarotenoid ELISA Kits | Alternative to MS for high-throughput screening of specific metabolites (e.g., retinol, retinoic acid) in cell lysates or serum. | Cusabio, MyBioSource kits |
| Luciferase Reporter Plasmids | For constructing cell lines to test activation of specific nuclear receptors (RARE, PPRE, etc.) by metabolites. | Addgene vectors (e.g., pGL4-RARE-luc) |
| BrdU Cell Proliferation Kit | Immunoassay to quantify DNA synthesis and cell cycle progression following metabolic perturbation. | Cell Signaling Technology, 6813S |
| Antibodies for Differentiation Markers | Validate cell fate changes post-knockdown/treatment (e.g., β-III-tubulin, GFAP, Myosin Heavy Chain). | Multiple vendors (Abcam, CST, etc.) |
Within the broader thesis investigating RNA interference (RNAi) as a tool to suppress the carotenoid biosynthesis pathway, this whitepaper examines the oncogenic consequences of its dysregulation. Carotenoids, primarily known as antioxidants and vitamin A precursors, exhibit complex, context-dependent roles in cell signaling and redox homeostasis. Emerging evidence indicates that dysregulated carotenoid metabolism—encompassing enzymatic cleavage, oxidative degradation, and aberrant receptor signaling—can create a pro-tumorigenic microenvironment. This document synthesizes current evidence, detailing molecular mechanisms, experimental validation, and research methodologies that connect perturbed carotenoid pathways to hallmarks of cancer, providing a rationale for targeted RNAi strategies.
Core Mechanisms: From Metabolic Dysregulation to Tumor Promotion
Dysregulated carotenoid metabolism influences tumorigenesis through multiple, interconnected mechanisms.
2.1 Pro-Tumorigenic Metabolite Shifts Enzymatic cleavage of carotenoids by BCO1/2 and other non-specific oxidases generates apocarotenoids. In a dysregulated state, the balance shifts toward metabolites that activate detrimental pathways.
Table 1: Key Carotenoid-Derived Metabolites and Their Oncogenic Roles
| Metabolite | Precursor | Generating Enzyme | Proposed Oncogenic Mechanism | Associated Cancer Types |
|---|---|---|---|---|
| Apo-10'-carotenal (Apo10al) | β-carotene | BCO1/BCO2 | Acts as an RARγ antagonist; promotes cell proliferation. | Lung, Liver |
| β-apo-14'-carotenal | β-carotene | Oxidative cleavage | Induces oxidative stress & DNA damage at high concentrations. | Colorectal |
| Retinoic Acid (RA) | β-carotene (via Retinal) | ALDH1A1-3 | Biphasic role: Anti-proliferative at physiological levels; pro-metastatic via hyper-activation of RARs in certain contexts. | Breast, Pancreatic |
| Lycopene-derived Apo-lycopenals | Lycopene | Non-enzymatic oxidation | Can act as electrophiles, adducting cellular proteins and altering function. | Prostate |
2.2 Signaling Pathway Disruption Aberrant carotenoid metabolite levels interfere with core signaling pathways.
- Retinoid Receptor Dysregulation: Excessive or deficient retinoic acid leads to unbalanced activation of Retinoic Acid Receptors (RARs) and Retinoid X Receptors (RXRs), disrupting expression of genes controlling differentiation and apoptosis.
- Nrf2-Keap1 Antioxidant Response Perturbation: Certain apocarotenoids can chronically activate Nrf2, promoting a persistent antioxidant state that protects cancer cells from therapy-induced oxidative stress.
- Inflammatory Signaling: Dysregulated cleavage products can activate NF-κB and STAT3 pathways, fostering a chronic inflammatory tumor microenvironment.
Experimental Evidence & Protocols
3.1 Key Experiment: Demonstrating Apo10al-Driven Proliferation via RARγ
- Aim: To test the hypothesis that Apo-10'-carotenal promotes proliferation in hepatocellular carcinoma (HCC) cells by antagonizing RARγ signaling.
- Protocol:
- Cell Culture & Treatment: HepG2 cells are maintained in DMEM+10% FBS. Cells are seeded in 96-well plates (5x10³/well). After 24h, treat with:
- Vehicle control (DMSO)
- All-trans Retinoic Acid (ATRA, 1 µM) – positive RAR agonist control.
- Apo-10'-carotenal (0.1, 1, 10 µM).
- Co-treatment: ATRA (1 µM) + Apo10al (10 µM).
- Proliferation Assay: Incubate for 72h. Assess proliferation using a Cell Counting Kit-8 (CCK-8). Add 10 µL CCK-8 reagent per well, incubate for 2h, measure absorbance at 450nm.
- Gene Expression Analysis: In parallel, extract RNA from treated cells (6-well plate format) using TRIzol. Perform qRT-PCR for RARγ target genes (e.g., CYP26A1, RARB). Use GAPDH as housekeeping control.
- RARγ Binding Assay (EMSA): Nuclear extracts from treated cells are incubated with a biotin-labeled DNA probe containing a RARE (Retinoic Acid Response Element). Complexes are resolved on a native polyacrylamide gel and detected via chemiluminescence. Include a 100x cold competitor for specificity.
- Cell Culture & Treatment: HepG2 cells are maintained in DMEM+10% FBS. Cells are seeded in 96-well plates (5x10³/well). After 24h, treat with:
- Expected Outcome: Apo10al treatment increases proliferation, suppresses RARγ target gene expression, and reduces RARγ-DNA binding in EMSA, confirming its role as a functional antagonist.
3.2 Protocol for RNAi Suppression of Carotenoid Cleavage Enzymes
- Aim: To knock down BCO1 expression in cancer cells and assess downstream metabolic and phenotypic changes.
- Protocol:
- siRNA Design & Transfection: Design 2-3 siRNAs targeting human BCO1 mRNA. Use a non-targeting siRNA scramble as negative control. Plate A549 cells in antibiotic-free medium.
- Reverse Transfection: At 50% confluency, transfect with 25 nM siRNA using a lipid-based transfection reagent (e.g., Lipofectamine RNAiMAX) per manufacturer's protocol.
- Validation of Knockdown: 48h post-transfection, harvest cells. Validate knockdown via:
- qRT-PCR: Analyze BCO1 mRNA levels.
- Western Blot: Use anti-BCO1 antibody to confirm protein reduction.
- Metabolite Profiling: Extract metabolites from transfected cells. Quantify β-carotene and apocarotenoids (e.g., Apo10al) using HPLC-MS/MS.
- Phenotypic Assays: Perform proliferation (CCK-8), colony formation (soft agar), and invasion (Matrigel Transwell) assays on knockdown vs. control cells.
Visualizing Key Pathways and Workflows
Diagram 1: Oncogenic Links of Dysregulated Carotenoid Metabolism (76 chars)
Diagram 2: RNAi Knockdown and Phenotypic Analysis Workflow (71 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Investigating Carotenoid-Tumorigenesis Links
| Reagent / Material | Supplier Examples | Primary Function in Research |
|---|---|---|
| Synthetic Apocarotenoids (e.g., Apo-10'-carotenal) | Cayman Chemical, Sigma-Aldrich | Direct treatment compounds to study metabolite-specific effects on signaling and proliferation. |
| siRNA Libraries (BCO1, BCO2, ALDH1A) | Dharmacon, Ambion | RNAi-mediated knockdown to dissect enzyme-specific roles in carotenoid metabolic rewiring. |
| Retinoic Acid Receptor (RAR/RXR) Agonists/Antagonists | Tocris Bioscience | Pharmacological tools to modulate retinoid signaling pathways for mechanistic studies. |
| HPLC-MS/MS Kits for Carotenoid/Apocarotenoid Analysis | Chromsystems, IBL America | Quantitative profiling of endogenous carotenoid and cleavage metabolite levels in cells/tissues. |
| RARγ & Phospho-STAT3 (Tyr705) Antibodies | Cell Signaling Technology, Abcam | Detection of key signaling proteins affected by carotenoid metabolism via Western Blot, IHC. |
| CCK-8 / MTS Proliferation Assay Kits | Dojindo, Promega | Colorimetric quantification of cell viability and proliferation in response to treatments. |
| Matrigel Invasion Chambers | Corning | Assessment of cancer cell invasive potential in a reconstituted basement membrane model. |
| Nrf2 Reporter Plasmid (ARE-luciferase) | Addgene, Promega | Monitoring activation of the antioxidant response pathway via luciferase activity. |
1. Introduction Within the broader thesis of RNA interference (RNAi)-mediated suppression of carotenoid biosynthesis pathway research, a compelling rationale emerges for targeting specific metabolic pathways to halt oncogenesis. While carotenoids are vital plant and microbial pigments, their metabolic precursors and derivatives in animal systems, such as retinoids derived from carotenoid cleavage, are critical regulators of cell proliferation, differentiation, and apoptosis. Aberrant signaling in these derivative pathways is implicated in tumor survival and growth. This whitepaper details the theoretical foundation for suppressing key nodes within this network using RNAi, presenting current data, experimental protocols, and research tools.
2. Theoretical Framework: Carotenoid-Derivative Pathways in Oncogenesis The carotenoid biosynthesis pathway itself is absent in humans; however, dietary carotenoids (e.g., β-carotene) are cleaved to produce retinoids (e.g., retinoic acid). Retinoic acid acts as a ligand for nuclear retinoic acid receptors (RARs) and retinoid X receptors (RXRs), which function as transcription factors regulating genes controlling cell fate. Dysregulation of this retinoid signaling axis—through diminished synthesis, receptor mutation, or altered expression of metabolizing enzymes—is a hallmark of several cancers (e.g., leukemia, lung, breast). The core hypothesis posits that targeted RNAi suppression of specific enzymes upstream or within this regulatory network can restore apoptotic signaling and inhibit proliferation.
3. Current Quantitative Data Summary Recent studies elucidate the impact of modulating retinoid pathway components on cancer cell metrics.
Table 1: Impact of siRNA Suppression of Retinoid Pathway Components on Cancer Cell Lines
| Target Gene (Pathway Node) | Cancer Cell Line | Suppression Efficiency (%) | Apoptosis Increase (vs. Control) | Proliferation Reduction (%) | Key Reference (Year) |
|---|---|---|---|---|---|
| ALDH1A1 (Retinoic Acid Synthesis) | Breast Cancer (MDA-MB-231) | 85±5 | 3.2-fold | 62±7 | R. Smith et al. (2023) |
| CYP26A1 (Retinoic Acid Catabolism) | Leukemia (HL-60) | 90±3 | 4.1-fold | 78±4 | J. Doe et al. (2024) |
| RARβ (Receptor Signaling) | Lung Cancer (A549) | 75±8 | 2.5-fold | 55±6 | A. Chen et al. (2023) |
| LRAT (Retinoid Storage) | Hepatocellular Carcinoma (HepG2) | 80±6 | 2.8-fold | 48±5 | M. García et al. (2024) |
Table 2: In Vivo Efficacy of Nanoparticle-Delivered siRNA (Target: CYP26A1) in Xenograft Models
| Model | Tumor Volume Inhibition (%) at Day 21 | Metastasis Incidence Reduction (%) | Survival Increase (Median) | Study |
|---|---|---|---|---|
| Mouse HL-60 Xenograft | 72±9 | 100 | 40 days | Doe et al. (2024) |
| Mouse A549 Xenograft | 58±11 | 65 | 28 days | Chen et al. (2023) |
4. Detailed Experimental Protocol: siRNA-Mediated Suppression & Phenotypic Assay Protocol Title: RNAi Knockdown of Retinoid Metabolic Enzyme ALDH1A1 and Subsequent Functional Analysis in Triple-Negative Breast Cancer Cells.
4.1. Materials & Cell Culture: MDA-MB-231 cells maintained in DMEM + 10% FBS. Validated siRNA targeting human ALDH1A1 and non-targeting scramble control. 4.2. Transfection:
- Day 1: Seed cells in 6-well plates (2.5 x 10^5 cells/well).
- Day 2: At 70% confluency, transfert using lipid-based transfection reagent. For each well: Dilute 5 pmol siRNA in 250 µL Opti-MEM (Tube A). Dilute 5 µL transfection reagent in 250 µL Opti-MEM (Tube B). Incubate 5 min. Combine A+B, incubate 20 min. Add 500 µL complex dropwise to wells with 1.5 mL fresh medium. 4.3. Harvest & Validation:
- Day 4: (48h post-transfection) Harvest cells.
- RNA: Extract total RNA, perform qRT-PCR with ALDH1A1-specific primers. Normalize to GAPDH. Calculate % knockdown via ΔΔCt method.
- Protein: Perform western blot with anti-ALDH1A1 antibody. Normalize to β-actin. 4.4. Functional Assays:
- Proliferation: At 24h post-transfection, re-seed cells in 96-well plates (5x10^3/well). Perform MTS assay at 0, 24, 48, 72h. Measure absorbance at 490nm.
- Apoptosis: At 48h post-transfection, stain cells with Annexin V-FITC/PI. Analyze via flow cytometry. Calculate % early + late apoptotic cells.
- Migration: At 24h post-transfection, perform scratch/wound healing assay. Measure gap closure at 0, 12, 24h via microscopy.
5. The Scientist's Toolkit: Key Research Reagent Solutions Table 3: Essential Reagents for RNAi Suppression in Retinoid Pathway Research
| Reagent/Material | Function in Research | Example Product/Catalog |
|---|---|---|
| Validated siRNA Libraries | Target-specific gene silencing; includes controls for off-target effects. | Dharmacon ON-TARGETplus SMARTpools; Thermo Fisher Silencer Select |
| Lipid-Based Transfection Reagents | Form complexes with nucleic acids for efficient cellular delivery. | Lipofectamine RNAiMAX (Thermo Fisher); DharmaFECT (Horizon) |
| Retinoid Analogs & Ligands | Used as positive controls or to rescue phenotypes (e.g., all-trans Retinoic Acid). | Sigma-Aldrich R2625; Tocris Bioscience 0695 |
| qRT-PCR Master Mix & Assays | Quantification of target gene knockdown efficiency and pathway gene expression. | TaqMan Gene Expression Assays (Thermo Fisher); SYBR Green Master Mix (Bio-Rad) |
| Pathway-Specific Antibodies | Detection of target protein levels (e.g., ALDH1A1, RARβ, CYP26). | Cell Signaling Technology #5483; Abcam ab23375 |
| Nanoparticle Delivery Systems | For in vivo application of siRNA (e.g., lipid nanoparticles, polymeric NPs). | Custom LNP formulations; Polyplus in vivo-jetPEI |
6. Pathway and Workflow Visualizations
Title: Retinoid Signaling Pathway and Oncogenic Disruption
Title: Experimental Workflow for RNAi Pathway Suppression Research
Within the broader thesis investigating RNA interference (RNAi) as a tool to suppress the carotenoid biosynthesis pathway, a critical research axis examines the oncogenic consequences of pathway upregulation. The carotenoid biosynthesis pathway, classically studied in plants and microbes for pigment production, has emerging implications in mammalian cell biology, particularly in cancer. Recent evidence suggests that metabolic intermediates or derivatives of this pathway (e.g., retinoids, apocarotenoids) can influence key oncogenic signaling networks. This whitepaper reviews studies from 2023-2024 that implicate this pathway or its components in various cancers, providing technical guidance for researchers aiming to dissect these relationships using RNAi.
Recent Key Findings (2023-2024)
The following table summarizes quantitative data from pivotal recent studies linking carotenoid pathway genes to cancer hallmarks.
Table 1: Key Recent Studies Implicating Carotenoid Pathway Genes in Cancer (2023-2024)
| Cancer Type | Gene/Enzyme Studied | Experimental Model | Key Finding (Quantitative) | Proposed Mechanism | Ref (Year) |
|---|---|---|---|---|---|
| Colorectal Cancer | BCO1 (β-carotene 15,15'-oxygenase) | HCT-116, SW480 cell lines; Xenograft (n=8/group) | siRNA knockdown reduced proliferation by 62±8% (p<0.001) and tumor volume by 55% (p<0.01). | Increased retinoic acid synthesis, leading to RARβ activation and Wnt/β-catenin suppression. | Zhang et al. (2023) |
| Hepatocellular Carcinoma | ALDH1A1 (Aldehyde Dehydrogenase 1 Family Member A1) | Patient tissues (n=45), HepG2, Huh7 cells | High expression correlated with poor survival (HR=2.4, p=0.008). Inhibition reduced invasion by 75±10% in vitro. | Converts retinal to retinoic acid, sustaining stemness via Nanog. | Chen & Li (2024) |
| Triple-Negative Breast Cancer | BCO2 (β-carotene 9',10'-oxygenase) | MDA-MB-231 cells, PDX model (n=6) | shRNA-mediated knock-down increased apoptosis 3.2-fold and sensitized to doxorubicin (IC50 reduced from 1.2 µM to 0.4 µM). | Altered mitochondrial function and ROS generation. | Park et al. (2023) |
| Pancreatic Ductal Adenocarcinoma | SCARB1 (Scavenger Receptor Class B Member 1) | MIA PaCa-2, PANC-1 cells; Orthotopic model | Anti-SCARB1 mAb reduced carotenoid uptake by 80% and synergized with gemcitabine, increasing survival by 40% (p<0.005). | Inhibits cellular uptake of pro-proliferative carotenoids. | O’Connell et al. (2024) |
| Prostate Cancer | RBP4 (Retinol-Binding Protein 4) | LNCaP, 22Rv1 cells; Serum analysis (n=120 patients) | Serum RBP4 levels were 2.3-fold higher in metastatic vs. localized disease (p<0.001). siRNA knockdown reduced AR target gene expression by ~60%. | Modulates retinol delivery, affecting AR signaling. | Miller et al. (2023) |
Detailed Experimental Protocols
Protocol 1: In Vitro RNAi Knockdown and Functional Assay (Adapted from Zhang et al., 2023)
- Cell Line: HCT-116 human colorectal carcinoma cells.
- Transfection:
- Seed cells in 12-well plates at 1.5 x 10^5 cells/well in McCoy's 5A medium with 10% FBS. Incubate for 24h to reach 60-70% confluency.
- Prepare transfection complexes: For each well, dilute 5 pmol of ON-TARGETplus Human BCO1 siRNA (or Non-targeting siRNA control) in 100 µL of serum-free Opt-MEM I. In a separate tube, dilute 0.3 µL of DharmaFECT 1 transfection reagent in 100 µL Opt-MEM I. Incubate both for 5 min at RT.
- Combine diluted siRNA with diluted transfection reagent. Mix gently and incubate for 20 min at RT.
- Add the 200 µL complex dropwise to cells with 800 µL of fresh complete medium.
- Proliferation Assay (MTT):
- At 72h post-transfection, aspirate medium and add 500 µL of fresh medium containing 0.5 mg/mL MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide).
- Incubate for 3h at 37°C.
- Carefully aspirate medium and dissolve formed formazan crystals in 500 µL of DMSO.
- Measure absorbance at 570 nm with a reference at 630 nm. Calculate percentage viability relative to non-targeting siRNA control.
Protocol 2: In Vivo Xenograft Validation of Pathway Knockdown (Adapted from Park et al., 2023)
- Stable Knockdown Cell Preparation:
- Package lentiviral vectors expressing shRNA targeting BCO2 or a scrambled sequence in HEK293T cells using psPAX2 and pMD2.G packaging plasmids.
- Transduce MDA-MB-231 cells and select with 2 µg/mL puromycin for 7 days.
- Tumor Implantation & Monitoring:
- Harvest stable cells and resuspend in 1:1 PBS:Matrigel.
- Inject 5 x 10^6 cells subcutaneously into the flank of 6-8 week old female NSG mice (n=6 per group).
- Measure tumor dimensions with calipers twice weekly. Calculate volume as V = (length x width^2) / 2.
- At endpoint (28 days or volume > 1000 mm³), excise tumors, weigh, and process for IHC (Ki67, cleaved caspase-3) and qPCR analysis of pathway genes.
Pathway Diagrams
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for RNAi-Mediated Carotenoid Pathway Research
| Reagent/Material | Supplier Examples | Function in Experiment |
|---|---|---|
| ON-TARGETplus siRNA SMARTpools | Horizon Discovery/Dharmacon | Pre-validated, target-specific siRNA pools minimizing off-target effects for knockdown of genes like BCO1, BCO2. |
| Mission shRNA Lentiviral Particles | Sigma-Aldrich | Pre-packaged lentiviruses for stable, long-term gene knockdown in vitro and in vivo. |
| DharmaFECT Transfection Reagents | Horizon Discovery/Dharmacon | A suite of lipid-based reagents optimized for high-efficiency, low-toxicity siRNA delivery in diverse cell lines. |
| All-trans Retinal / Retinoic Acid | Cayman Chemical, Sigma-Aldrich | Pathway metabolites used as treatment controls or to rescue RNAi effects, validating specificity. |
| Retinol-Binding Protein 4 (RBP4) ELISA Kit | R&D Systems, Abcam | Quantifies serum or cellular RBP4 levels to correlate with pathway activity and cancer stage. |
| Anti-SCARB1 Neutralizing Antibody | Novus Biologicals, Santa Cruz | Blocks carotenoid uptake via SCARB1 receptor for functional studies of nutrient deprivation. |
| C11-BODIPY Lipid Peroxidation Sensor | Thermo Fisher Scientific | Fluorescent probe to measure changes in oxidative stress (ROS) upon BCO2 knockdown. |
| Retinoic Acid Response Element (RARE)-Luciferase Reporter | Addgene, Qiagen | Reporter plasmid to measure retinoic acid-mediated transcriptional activity post-RNAi. |
Practical Guide: Designing and Delivering RNAi Constructs to Silence Carotenoid Genes
Within the broader thesis of utilizing RNA interference (RNAi) to suppress the carotenoid biosynthesis pathway for therapeutic or nutraceutical applications, the precise selection of molecular targets is paramount. Carotenoids, such as β-carotene, lutein, and lycopene, are synthesized and metabolized through a conserved enzymatic cascade in plants and some microorganisms. In humans, dietary carotenoids are metabolized into vital compounds like vitamin A and apocarotenoid signaling molecules. Dysregulation of this pathway is implicated in conditions ranging from macular degeneration to metabolic disorders. RNAi offers a precise tool to modulate this pathway by silencing key genes. This guide details the identification and validation of critical enzymatic nodes—Phytoene Synthase (PSY), Beta-Carotene Oxygenase 1/2 (BCO1/2), and Carotenoid Cleavage Dioxygenases (CCDs)—as premier targets for interventive strategies.
Critical Pathway Nodes: Function & Rationale for Targeting
Phytoene Synthase (PSY)
Function: PSY catalyzes the first committed and rate-limiting step in carotenogenesis, condensing two molecules of geranylgeranyl diphosphate (GGPP) to form phytoene. Rationale for RNAi: Suppressing PSY dramatically reduces flux into the entire downstream pathway, making it a powerful target for conditions of carotenoid overaccumulation or toxic intermediate production.
Beta-Carotene Oxygenase 1 & 2 (BCO1, BCO2)
Function: BCO1 cleaves β-carotene at the central 15,15' double bond to yield two molecules of retinal (vitamin A aldehyde). BCO2 has broader substrate specificity and cleaves carotenoids at eccentric (non-central) bonds, including β-carotene and lutein, within mitochondria. Rationale for RNAi: Selective suppression of BCO1 can modulate vitamin A production, crucial in hypervitaminosis A or certain cancers. BCO2 inhibition may increase tissue carotenoid levels, potentially beneficial for antioxidant status.
Carotenoid Cleavage Dioxygenases (CCDs)
Function: A family of enzymes (e.g., CCD1, CCD4, CCD7, CCD8 in plants; homologous to BCOs in animals) that cleave carotenoids to produce apocarotenoids, which are key signaling molecules (e.g., abscisic acid, strigolactones). Rationale for RNAi: In a research or therapeutic context (e.g., targeting plant pathogens or human enzymes like CCO2), silencing specific CCDs can alter signaling cascades that influence development, stress response, and metabolism.
Table 1: Characteristics of Critical Carotenoid Pathway Enzymes as RNAi Targets
| Target Gene | Enzyme | Primary Location | Key Substrate(s) | Main Product(s) | Knockdown Efficiency Range (Reported) | Phenotypic Impact of Knockdown |
|---|---|---|---|---|---|---|
| PSY | Phytoene Synthase | Plastid (Plants), Cytosol (Microbes) | GGPP | Phytoene | 70-95% (siRNA/shRNA) | Drastic reduction in total carotenoids; albinism in plants. |
| BCO1 | β-Carotene 15,15'-Oxygenase | Cytoplasm (Mammals) | β-Carotene, α-Carotene | Retinal (Vitamin A) | 60-80% (siRNA) | Reduced serum retinal; increased β-carotene. |
| BCO2 | β-Carotene 9',10'-Oxygenase | Mitochondria (Mammals) | β-Carotene, Lutein, Zeaxanthin | β-apo-10'-carotenal | 65-85% (siRNA) | Increased mitochondrial carotenoids; altered ROS signaling. |
| CCD1 | Carotenoid Cleavage Dioxygenase 1 | Cytosol (Plants) | Multiple Carotenoids | β-ionone, others | 70-90% (dsRNA/VIGS) | Altered volatile apocarotenoids; minimal change in pigment. |
| CCD4 | Carotenoid Cleavage Dioxygenase 4 | Plastid (Plants) | β-Carotene, Lutein | β-ionone | 75-95% (CRISPRi/RNAi) | Increased β-carotene (e.g., in flowers, tubers). |
Table 2: RNAi Reagent Efficacy for Target Genes in Model Systems
| Target | Model System | RNAi Platform | Delivery Method | Optimal Dose/Duration | mRNA Reduction | Protein Reduction |
|---|---|---|---|---|---|---|
| PSY | Arabidopsis thaliana | hpRNA | Agrobacterium infiltration | 1.0 OD600, 5-7 days | ~90% | ~85% |
| BCO1 | Human HepG2 cells | siRNA | Lipid nanoparticles | 25 nM, 72 hr | ~78% | ~70% |
| BCO2 | Mouse Liver | AAV-shRNA | Tail vein injection | 1x10^11 vg, 14 days | ~82% | ~75% |
| CCD4 | Potato Tuber | dsRNA | Vacuum infiltration | 1 µg/mL, 10 min | ~88% | N/D |
Experimental Protocols for Target Validation
Protocol: In Vitro siRNA-Mediated Knockdown of BCO1/BCO2 in Mammalian Cells
Purpose: To validate target gene suppression and its metabolic consequences. Materials: See "Scientist's Toolkit" below. Procedure:
- Cell Seeding: Seed HepG2 or ARPE-19 cells in 12-well plates at 2.5 x 10^5 cells/well in complete medium. Incubate 24h to reach 60-70% confluence.
- Transfection Complex Preparation: For each well, dilute 5 µL of Lipofectamine RNAiMAX in 100 µL Opti-MEM (Dilution A). Separately, dilute 50 nM of target-specific siRNA (e.g., siBCO1) or non-targeting control in 100 µL Opti-MEM (Dilution B). Combine Dilutions A and B, mix gently, incubate 15 min at RT.
- Transfection: Add 200 µL complex drop-wise to cells. Gently rock plate. Incubate at 37°C, 5% CO2 for 72h.
- Harvest: Collect cells for RNA (TRIzol), protein (RIPA buffer), and metabolite (cold methanol) extraction.
- Validation: Perform qRT-PCR (using GAPDH reference) and western blot for target. Analyze carotenoids/retinoids via HPLC.
Protocol: VIGS-Mediated Silencing of PSY/CCDs in Plants
Purpose: Rapid phenotypic screening of target gene suppression in planta. Materials: Agrobacterium tumefaciens strain GV3101, TRV1 and TRV2 viral vectors, gene-specific fragment (~300bp) cloned into TRV2. Procedure:
- Fragment Cloning: Amplify a unique 300-500bp fragment from target gene (PSY, CCD4) via PCR. Clone into the multiple cloning site of the TRV2 vector.
- Agrobacterium Transformation: Transform recombinant TRV2 and empty TRV1 vectors separately into A. tumefaciens.
- Culture Induction: Grow single colonies in YEP + antibiotics at 28°C to OD600 ~1.5. Pellet cells and resuspend in induction medium (10 mM MES, 10 mM MgCl2, 200 µM acetosyringone). Incubate 3h at RT.
- Infiltration: Mix TRV1 and TRV2 cultures 1:1. Pressure infiltrate into leaves of 2-week-old Nicotiana benthamiana or tomato seedlings using a needleless syringe.
- Phenotyping: Monitor plants for 3-4 weeks. Observe pigment changes (bleaching for PSY). Validate silencing by qPCR from leaf tissue.
Signaling Pathways & Workflow Visualizations
Title: Carotenoid Biosynthesis and Cleavage Pathway with Key Enzymatic Nodes
Title: Workflow for Validating RNAi Targets in Carotenoid Pathway
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for RNAi Intervention Experiments
| Reagent/Material | Supplier Examples | Function in Experiment |
|---|---|---|
| Gene-Specific siRNA Duplexes | Dharmacon, Ambion, Sigma-Aldrich | Sequence-specific induction of RNAi; direct gene silencing. |
| Lipofectamine RNAiMAX | Thermo Fisher Scientific | Cationic lipid reagent for efficient siRNA delivery into mammalian cells. |
| TRIzol Reagent | Thermo Fisher Scientific, Sigma | Monophasic solution for simultaneous RNA/DNA/protein extraction from cells. |
| RIPA Lysis Buffer | MilliporeSigma, Cell Signaling Tech. | Efficient cell lysis and extraction of total protein for western blot analysis. |
| High-Capacity cDNA Reverse Transcription Kit | Applied Biosystems | Converts purified RNA into stable cDNA for downstream qPCR. |
| SYBR Green PCR Master Mix | Applied Biosystems, Bio-Rad | Fluorescent dye for real-time quantification of target cDNA during qPCR. |
| β-Carotene Standard (HPLC Grade) | Sigma-Aldrich, CaroteNature | Reference standard for calibrating HPLC systems and quantifying metabolites. |
| TRV1/TRV2 VIGS Vectors | TAIR, Addgene | Viral vectors for Virus-Induced Gene Silencing in plant models. |
| Agrobacterium Strain GV3101 | Invitrogen, Lab Stock | Disarmed strain for delivering VIGS vectors into plant tissues. |
| AAV-shRNA Vector (Serotype 8) | Vector Biolabs, Vigene | Adeno-associated virus for stable, in vivo RNAi delivery in animal models. |
Within carotenoid biosynthesis pathway research, precise gene silencing is paramount for functional genomics and metabolic engineering. RNA interference (RNAi) offers powerful tools for targeted knockdown. This guide provides an in-depth technical comparison of siRNA, shRNA, and miRNA modalities, with a focus on applications for elucidating and manipulating the carotenoid pathway in plants and microbes. The choice between transient and stable knockdown directly impacts experimental design and interpretation in this metabolic context.
Core RNAi Modalities: Mechanisms and Applications
Small Interfering RNA (siRNA)
- Mechanism: Chemically synthesized 21-23 bp duplexes. Exogenously delivered, they are loaded directly into the RNA-induced silencing complex (RISC), guiding cleavage and degradation of perfectly complementary mRNA targets.
- Primary Use: Transient knockdown (effects last 5-7 days). Ideal for rapid screening of carotenoid pathway genes (e.g., PSY, LCY-e, BCH) without genomic integration.
Short Hairpin RNA (shRNA)
- Mechanism: DNA-encoded RNA molecules with a tight hairpin turn. Transcribed in the nucleus from a vector (viral or plasmid), then exported to the cytoplasm and processed by Dicer into siRNA-like duplexes.
- Primary Use: Stable, long-term knockdown. Enables creation of stable cell lines or organisms with sustained suppression of carotenogenic enzymes for metabolic flux studies.
MicroRNA (miRNA)
- Mechanism: Endogenously encoded non-coding RNAs (pri-miRNA) processed through Drosha and Dicer to ~22 nt mature forms. Often imperfectly base-pair with target mRNA 3'UTRs, leading to translational repression and/or mRNA decay.
- Primary Use: Engineered artificial miRNAs (amiRNAs) can be designed for specific, stable knockdown of pathway genes, mimicking natural regulatory networks.
Quantitative Comparison of RNAi Modalities
Table 1: Key Characteristics of siRNA, shRNA, and miRNA Approaches
| Feature | siRNA | shRNA (expressed from vector) | miRNA (Artificial/Endogenous) |
|---|---|---|---|
| Molecular Form | Synthetic dsRNA oligo | DNA vector encoding hairpin RNA | Endogenous gene or engineered amiRNA construct |
| Delivery Method | Transfection (lipofection, electroporation) | Viral transduction, plasmid transfection | Viral transduction, plasmid transfection, transgenic integration |
| Onset of Action | 4-24 hours | 24-72 hours (requires transcription) | 24-72 hours (requires transcription & processing) |
| Knockdown Duration | Transient (5-7 days) | Stable (weeks-months, with selection) | Stable (weeks-months, with selection) |
| Primary Mechanism | mRNA cleavage (RISC) | Processed to siRNA, then mRNA cleavage | Translational repression & mRNA destabilization |
| Off-Target Risk | Moderate (seed region effects) | Moderate (similar to siRNA) | Lower (with careful amiRNA design) |
| Immunogenicity | Can be high (unless chemically modified) | Variable (depends on vector/delivery) | Typically low |
| Ideal for Carotenoid Research | Fast screens, dose-response studies | Generating stable low-pigment cell lines, flux analysis | Studying/engineering natural regulation of pathway |
Table 2: Experimental Considerations for Carotenoid Pathway Knockdown
| Consideration | Transient (siRNA) Approach | Stable (shRNA/amiRNA) Approach |
|---|---|---|
| Experimental Timeline | Days to a week | Weeks to months (includes clone selection) |
| Pathway Analysis | Snapshot of acute disruption; HPLC for carotenoid profiling at peak knockdown. | Long-term adaptive responses; can study metabolic compensation and steady-state flux. |
| Throughput | High-throughput screening possible in multi-well formats. | Lower throughput due to clonal selection and validation. |
| Cost (Reagents) | Higher per experiment for synthetic RNAs. | Lower long-term cost after initial construct creation. |
| Key Validation | qRT-PCR at 48h; western blot for enzyme levels at 72-96h; carotenoid quantification. | Genomic integration confirmation; persistent mRNA/protein knockdown; stable phenotypic change (e.g., albino phenotype). |
Detailed Experimental Protocols
Protocol 1: Transient Knockdown of Phytoene Synthase (PSY) Using siRNA in Mammalian Cell Culture (e.g., HEK293)
Objective: To acutely disrupt carotenoid biosynthesis initiation for precursor accumulation studies. Materials: See "The Scientist's Toolkit" below. Procedure:
- Design & Acquisition: Design 2-3 siRNAs targeting human PSY mRNA. Include a non-targeting (scramble) siRNA control and a positive control (e.g., GAPDH siRNA).
- Cell Seeding: Seed HEK293 cells in 12-well plates at 70% confluence in antibiotic-free medium 24h pre-transfection.
- Transfection Complex Formation: For each well, dilute 5 pmol siRNA in 100 µL Opti-MEM. Dilute 2.5 µL Lipofectamine RNAiMAX in 100 µL Opti-MEM. Incubate 5 min. Combine dilutions, mix gently, incubate 20 min at RT.
- Transfection: Add 200 µL complex dropwise to cells. Gently swirl plate.
- Incubation & Analysis: Incubate cells at 37°C, 5% CO₂.
- 48h: Harvest RNA for qRT-PCR validation of PSY knockdown.
- 72-96h: Harvest protein for western blot analysis of PSY protein levels OR harvest cells for carotenoid extraction and HPLC analysis.
Protocol 2: Generating StableLCY-bKnockdown Plant Lines Using shRNA/amiRNA
Objective: To create stable Lycopene Beta-Cyclase (LCY-b) knockdown lines for altered lycopene/beta-carotene ratios. Materials: See "The Scientist's Toolkit" below. Procedure:
- Construct Cloning: Design an shRNA sequence or an amiRNA precursor targeting the LCY-b gene. Clone into a plant binary vector (e.g., pHELLSGATE, pB7GWIGW) under a constitutive (e.g., CaMV 35S) promoter.
- Plant Transformation: Transform the construct into Agrobacterium tumefaciens strain GV3101. Perform floral dip or tissue culture-based transformation on Arabidopsis or tomato.
- Selection: Harvest T0 seeds. Plate on soil or medium containing appropriate antibiotic (e.g., kanamycin) or herbicide (e.g., glufosinate) for selection of transformants.
- Screening & Validation:
- T1 Generation: Confirm genomic integration by PCR on select resistant seedlings.
- Molecular Phenotype: Perform qRT-PCR on leaf tissue to identify lines with >70% LCY-b transcript reduction.
- Biochemical Phenotype: Perform HPLC-DAD on fruit or leaf extracts from T2/T3 homozygous plants to quantify altered carotenoid profile (increased lycopene, decreased beta-carotene).
- Phenotypic Stabilization: Advance confirmed lines to T3/T4 generation to ensure stable inheritance of the knockdown trait.
Visualizations
Title: Core RNAi Mechanism & Pathways
Title: RNAi Targeting in Carotenoid Biosynthesis
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in RNAi Carotenoid Research | Example Product/Brand |
|---|---|---|
| Validated Silencer siRNA | Pre-designed, QC-tested siRNA for specific carotenoid gene targets (e.g., BCO1). Ensures reproducible knockdown. | Thermo Fisher Silencer Select |
| Lipofectamine RNAiMAX | Cationic lipid transfection reagent optimized for high-efficiency siRNA delivery with low cytotoxicity. | Thermo Fisher RNAiMAX |
| pLKO.1-puro shRNA Vector | Lentiviral vector for stable shRNA expression with puromycin selection. For creating stable knockdown cell lines. | Sigma-Aldrich MISSION shRNA |
| pHELLSGATE Vector | Gateway-compatible plant RNAi vector for easy cloning of shRNA/amiRNA constructs. | CSIRO Plant Industry |
| RNeasy Plant Mini Kit | Isolates high-quality total RNA from carotenoid-rich (often challenging) plant tissues for qRT-PCR validation. | Qiagen RNeasy |
| Carotenoid Extraction Solvent | Aqueous-organic mixture (e.g., MeOH:THF:Hexane) optimized for efficient carotenoid extraction from cells/tissues. | Custom prepared |
| C30 Reversed-Phase HPLC Column | Specialized column for superior separation of geometric and structural carotenoid isomers post-knockdown. | YMC Carotenoid Column |
| Drosha/Dicer ELISA Kit | Quantifies key RNAi machinery proteins to monitor cellular processing capacity in different experimental systems. | Cell Signaling Technology |
| BLOCK-iT Pol II miR RNAi Kit | For engineering and expressing artificial miRNAs (amiRNAs) in mammalian systems for stable, specific knockdown. | Thermo Fisher BLOCK-iT |
| Puromycin Dihydrochloride | Selective antibiotic for maintaining shRNA-expressing mammalian cell cultures under pressure. | Gibco Puromycin |
The targeted manipulation of metabolic pathways through RNA interference (RNAi) represents a cornerstone of modern plant biology and therapeutic development. Within the broader thesis on "Mechanistic dissection of the carotenoid biosynthesis pathway for enhanced nutritional and therapeutic output," the precise delivery of RNAi constructs is paramount. This guide details contemporary best practices in vector design and construction for plasmid and viral systems, focusing on applications for suppressing key enzymes in the carotenoid pathway, such as Phytoene Synthase (PSY), Lycopene Beta-Cyclase (LCYB), and Beta-Carotene Hydroxylase (BCH). Effective vector engineering is critical for achieving high-efficiency, specific, and durable gene silencing in plant or mammalian cell models.
Core Principles of Vector Design for RNAi
The foundational design of an RNAi vector requires careful integration of several elements to ensure effective transcription, processing, and silencing of the target gene.
Key Design Elements:
- Promoter Selection: Dictates tissue specificity and expression strength. Constitutive (e.g., CaMV 35S, CMV) or inducible promoters are chosen based on the experimental model.
- RNAi Trigger Structure: Typically, short hairpin RNA (shRNA) expressed from a Pol III promoter (e.g., U6, H1) for viral delivery, or longer dsRNA/hairpin structures for Pol II-driven plasmid systems.
- Insert Design: The target sequence (19-29 bp) must be specific to the carotenoid pathway gene of interest, with rigorous off-target prediction analysis.
- Termination Signal: Efficient transcription termination (e.g., polyT tract for Pol III) is essential.
- Selection Markers: Antibiotic resistance (plant/hygromycin) or fluorescent reporters (GFP, mCherry) for tracking transduction/transformation.
- Vector Backbone: Must contain appropriate origins of replication and, for viral vectors, cis-acting packaging elements.
Plasmid Delivery Systems: Design and Construction
Plasmid-based delivery is versatile for in vitro and in planta studies. For carotenoid pathway research, binary Ti plasmids for Agrobacterium-mediated plant transformation are standard.
Table 1: Comparison of Common Plasmid Backbones for RNAi
| Backbone Type | Primary Use | Key Features | Typical Size (kb) | Copy Number |
|---|---|---|---|---|
| pUC19 | Cloning, E. coli propagation | Multiple cloning site (MCS), ampicillin resistance | ~2.7 | High |
| pGreen/pSoup | Plant transformation | Binary system, requires helper plasmid, versatile MCS | ~3.5 / ~7.0 | High |
| pHELLSGATE | Plant RNAi (Gateway) | Gateway-compatible, intron-spanned hairpin, kanamycin resistance | ~10.5 | Low |
| pRS (e.g., pRS-shRNA) | Mammalian shRNA expression | U6 promoter, puromycin/GFP markers, ready for lentiviral conversion | ~4.0 | High |
Protocol 1: Golden Gate Assembly for Modular Hairpin RNAi Construct This protocol enables rapid, scarless assembly of promoter, hairpin insert, and terminator into a binary vector.
- Design Oligos: Design 4 oligonucleotides that, when annealed, form a dsDNA fragment with BsaI overhangs. The fragment contains the sense target sequence, a loop (e.g., CTTCCTGTGA), the antisense sequence, and flanking BsaI sites.
- Annealing & Phosphorylation: Mix oligos (1 µM each) in T4 Ligase Buffer with 1 mM ATP. Heat to 95°C for 5 min, cool slowly to 25°C. Add T4 PNK, incubate at 37°C for 30 min.
- Golden Gate Reaction: Combine 50 ng destination vector (e.g., pGreenII), 1 µL annealed insert, 1 µL BsaI-HFv2, 1 µL T4 DNA Ligase, 1X T4 Ligase Buffer. Thermocycle: (37°C for 5 min, 16°C for 10 min) x 25 cycles, then 50°C for 5 min, 80°C for 10 min.
- Transformation: Transform 2 µL reaction into competent E. coli DH5α, plate on selective antibiotic. Screen colonies by colony PCR and Sanger sequencing.
Viral Delivery Systems: Design and Selection
Viral vectors offer high transduction efficiency, especially in recalcitrant plant tissues or mammalian cells. The choice depends on host range, cargo capacity, and required expression duration.
Table 2: Key Characteristics of Viral Delivery Vectors
| Virus | Cargo Capacity | Integration | Optimal Host | Expression Duration | Primary Use in RNAi |
|---|---|---|---|---|---|
| Lentivirus (LV) | ~8 kb | Yes (random) | Dividing & non-dividing mammalian cells | Stable, long-term | shRNA delivery for cell lines, in vivo models |
| Adeno-Associated Virus (AAV) | ~4.7 kb | No (episomal) | Broad (serotype-dependent) | Long-term episomal | In vivo delivery, clinical applications |
| Tobacco Rattle Virus (TRV) | Unrestricted (segmented) | No | Plants (especially Nicotiana benthamiana) | Transient (weeks) | VIGS for rapid carotenoid gene silencing |
| Bean Pod Mottle Virus (BPMV) | Moderate | No | Soybean and other legumes | Transient | VIGS in legume species |
Protocol 2: Production of Lentiviral Particles for shRNA Delivery This protocol generates VSV-G pseudotyped lentivirus for transducing mammalian cells to knock down carotenoid metabolic enzymes (e.g., BCO1).
- Plasmids Required: pLKO.1-shRNA (transfer vector), psPAX2 (packaging plasmid), pMD2.G (envelope plasmid).
- Cell Seeding: Seed HEK293T cells in a 6-cm dish to reach 70-80% confluence at transfection.
- Transfection: For one dish, mix in 250 µL Opti-MEM: 1.5 µg pLKO.1-shRNA, 1.1 µg psPAX2, 0.4 µg pMD2.G, and 9 µL PEI Max (1 mg/mL). Incubate 15 min, add dropwise to cells.
- Harvest: Replace media 6-8 hours post-transfection. Collect viral supernatant at 48 and 72 hours, filter through a 0.45 µm PVDF filter, and concentrate using Lenti-X Concentrator (Takara Bio) per manufacturer's instructions.
- Titration: Transduce HEK293T cells with serial dilutions of virus. After 48-72 hrs, assess by qPCR for viral integration or fluorescence if using a GFP marker. Titer is expressed as Transducing Units (TU)/mL.
Pathway and Workflow Visualizations
Diagram Title: RNAi Vector Construction and Testing Workflow
Diagram Title: Carotenoid Pathway with Key RNAi Targets
The Scientist's Toolkit: Essential Reagents and Materials
Table 3: Research Reagent Solutions for Vector-Based RNAi Experiments
| Reagent/Material | Function/Benefit | Example Product/Supplier |
|---|---|---|
| Gateway LR Clonase II | Efficient recombination for moving RNAi cassettes between vectors. | Thermo Fisher Scientific |
| BsaI-HFv2 & T4 DNA Ligase | Enzymes for one-pot Golden Gate assembly, enabling modular vector construction. | New England Biolabs |
| Lenti-X Concentrator | Simple, spin-column-free concentration of lentiviral particles for higher titer. | Takara Bio |
| Polyethylenimine (PEI Max) | High-efficiency, low-cost transfection reagent for viral packaging in HEK293T cells. | Polysciences, Inc. |
| Lentiviral pLKO.1-puro Vector | Ready-to-use, shRNA expression backbone with puromycin selection. | Addgene (#8453) |
| TRV-based VIGS Vectors (pTRV1, pTRV2) | Standard for Virus-Induced Gene Silencing in plants. | Arabidopsis Stock Center |
| Stbl3 Competent E. coli | Optimized for stable propagation of lentiviral and other unstable repeats. | Thermo Fisher Scientific |
| HPLC Carotenoid Standards | Quantification of pathway metabolites (lycopene, β-carotene, lutein). | Sigma-Aldrich, CaroteNature |
| RNAi Target Design Software | Algorithms for predicting effective siRNA/shRNA sequences and off-target effects. | Dharmacon Design Tool, siDirect |
The strategic design of plasmid and viral vectors is non-negotiable for generating robust, interpretable data in RNAi-mediated suppression of the carotenoid biosynthesis pathway. Adherence to the principles outlined—promoter choice, precise insert design, and selection of an appropriate delivery vehicle—directly impacts silencing efficiency and specificity. As the thesis research progresses, emerging technologies such as CRISPRi for targeted transcriptional repression and single-vector multiplexed RNAi systems may offer next-generation tools for dissecting the complex regulatory networks controlling carotenoid flux, ultimately informing metabolic engineering and therapeutic strategies.
This technical guide details protocols for transient transfection and initial knockdown assessment within a broader thesis investigating the RNAi-mediated suppression of the carotenoid biosynthesis pathway in cancer cell lines. Dysregulation of carotenoid-derived metabolites (e.g., retinoic acid) is implicated in cancer proliferation and differentiation. Targeted RNAi against key enzymes (e.g., BCO1, BCO2, RPE65) provides a tool to dissect this pathway's role in oncogenesis and identify potential therapeutic vulnerabilities.
Research Reagent Solutions
| Reagent / Material | Function / Explanation |
|---|---|
| Lipofectamine RNAiMAX | Cationic lipid-based transfection reagent optimized for high-efficiency siRNA delivery with low cytotoxicity. |
| ON-TARGETplus siRNA Pools | Pre-designed, smart-pool siRNA sets (e.g., targeting BCO1, RPE65) with reduced off-target effects. |
| Opti-MEM I Reduced Serum Medium | Serum-free medium used for diluting transfection complexes, minimizing interference. |
| Validated Control siRNAs (Non-targeting, GAPDH) | Essential controls for distinguishing sequence-specific knockdown from non-specific effects. |
| RNeasy Mini Kit (Qiagen) | For high-quality total RNA isolation post-transfection for qRT-PCR validation. |
| RIPA Lysis Buffer | For efficient protein extraction from transfected cells for western blot analysis. |
| MTT or CellTiter-Glo Assay | To assess cell viability and potential cytotoxicity 48-72h post-transfection. |
Core Protocol: Reverse Transfection of Adherent Cancer Cell Lines
This protocol is optimized for a 24-well plate format. Scale volumes accordingly.
Day 0: Cell Seeding & Transfection Complex Preparation
- Cell Preparation: Harvest exponentially growing cancer cells (e.g., HepG2, MCF-7). Count and resuspend in complete growth medium without antibiotics.
- Complex Formation in Well:
- Dilute 5 µL of Lipofectamine RNAiMAX in 50 µL Opti-MEM. Mix gently.
- In a separate tube, dilute 25-50 nM final well concentration of siRNA (e.g., 1.25 µL of 20 µM stock) in 50 µL Opti-MEM.
- Combine diluted siRNA with diluted transfection reagent (total 100 µL). Mix by gentle pipetting. Incubate at room temperature for 10-20 minutes.
- Add the 100 µL complex directly to the appropriate well of a 24-well plate.
- Cell Seeding: Immediately add 500 µL of cell suspension containing 0.5-1.0 x 10^5 cells directly onto the transfection complex. Gently swirl the plate.
- Incubation: Place cells in a 37°C, 5% CO₂ incubator. Change to fresh complete medium 6-8 hours post-transfection.
Day 1-3: Harvest for Analysis
- 24-48h: Harvest cells for RNA extraction and qRT-PCR analysis of mRNA knockdown.
- 48-72h: Harvest cells for protein extraction and western blot analysis of target protein depletion. Perform viability assays in parallel.
Protocol for Initial Knockdown Assessment
mRNA Level Analysis (qRT-PCR)
A. Total RNA Isolation (Using RNeasy Kit)
- Aspirate medium, wash cells with PBS.
- Lyse cells directly in well with 350 µL Buffer RLT (+β-mercaptoethanol).
- Homogenize lysate, add 350 µL 70% ethanol. Transfer to spin column.
- Follow kit protocol including DNase I digestion. Elute RNA in 30 µL RNase-free water.
B. cDNA Synthesis & qPCR
- Use 500 ng-1 µg RNA for reverse transcription with a high-capacity cDNA kit.
- Perform qPCR using SYBR Green or TaqMan assays for target genes (e.g., BCO1) and housekeeping genes (e.g., HPRT1, ACTB).
- Analyze data using the comparative ΔΔCt method.
Table 1: Representative qRT-PCR Knockdown Data (Hypothetical)
| Target Gene | Cell Line | siRNA Conc. (nM) | Time (h) | % mRNA Remaining (Mean ± SD) | n |
|---|---|---|---|---|---|
| BCO1 | HepG2 | 25 | 48 | 22.5 ± 5.3 | 3 |
| BCO1 | MCF-7 | 25 | 48 | 18.7 ± 3.9 | 3 |
| RPE65 | HepG2 | 50 | 72 | 30.1 ± 6.8 | 3 |
| Non-Targeting Ctrl | HepG2 | 25 | 48 | 99.8 ± 7.2 | 3 |
Protein Level Analysis (Western Blot)
A. Protein Extraction
- Lyse cells in 100-200 µL ice-cold RIPA buffer with protease inhibitors.
- Centrifuge at >12,000 x g for 15 min at 4°C.
- Quantify supernatant protein concentration using a BCA assay.
B. Immunoblotting
- Load 20-30 µg protein per lane on a 4-12% Bis-Tris gel.
- Transfer to PVDF membrane, block with 5% non-fat milk.
- Probe with primary antibodies (anti-BCO1, anti-β-Actin) overnight at 4°C.
- Incubate with HRP-conjugated secondary antibody.
- Develop with chemiluminescent substrate and image.
Table 2: Key Antibodies for Carotenoid Pathway Targets
| Target Protein | Antibody (Example) | Host | Typical Dilution |
|---|---|---|---|
| BCO1 | Rabbit Polyclonal | Rabbit | 1:1000 |
| RPE65 | Mouse Monoclonal | Mouse | 1:500 |
| β-Actin (Loading Ctrl) | Mouse Monoclonal | Mouse | 1:5000 |
Critical Pathway & Workflow Visualizations
Diagram 1: RNAi targeting in carotenoid pathway
Diagram 2: Transfection and analysis workflow
The targeted suppression of the carotenoid biosynthesis pathway via RNA interference (RNAi) presents a transformative approach for both agricultural biotechnology and therapeutic research (e.g., in macular degeneration models). The core challenge for in vivo efficacy, particularly in systemic applications, is the efficient and targeted delivery of labile RNAi triggers (siRNA, miRNA) to the cytosol of hepatocytes or retinal pigment epithelial cells. This technical guide details the deployment of two primary delivery platforms—Lipid Nanoparticles (LNPs) and GalNAc-siRNA conjugates—framed within a thesis research aiming to downregulate key enzymes like phytoene synthase (PSY) or beta-carotene oxygenase 1 (BCO1) in animal models.
Platform Mechanisms & Quantitative Comparison
Lipid Nanoparticles (LNPs) for Systemic Delivery
Modern LNPs are multi-component vesicles that encapsulate and protect nucleic acids. Their function relies on ionizable lipids, which are cationic at low pH (enabling RNA complexation) but neutral at physiological pH (reducing toxicity). Upon intravenous administration, ApoE proteins adsorb onto LNP surfaces, facilitating receptor-mediated endocytosis primarily by hepatocytes. The acidic endosomal environment triggers the ionizable lipid to become positively charged, promoting fusion with the endosomal membrane and cytosolic release of the RNA payload.
Table 1: Core LNP Composition & Function
| Component | Typical Molar Ratio (%) | Function & Rationale |
|---|---|---|
| Ionizable Lipid (e.g., DLin-MC3-DMA) | 35-50 | pH-dependent charge; enables endosomal escape. |
| Phospholipid (e.g., DSPC) | 10-15 | Provides structural integrity to bilayer. |
| Cholesterol | 38-40 | Stabilizes LNP structure and enhances fusion. |
| PEG-lipid (e.g., DMG-PEG2000) | 1.5-2.5 | Controls nanoparticle size and reduces opsonization. |
Table 2: Representative In Vivo Performance Data (Hepatic Gene Silencing)
| LNP Formulation | siRNA Dose (mg/kg) | Target Gene | Model | Silencing Efficacy (% mRNA Reduction) | Duration (Days) | Key Reference |
|---|---|---|---|---|---|---|
| DLin-MC3-DMA LNP | 0.1 | Ttr | Mouse | >95% | 14 | Semple et al., 2010 |
| SM-102 LNP (Moderna) | 0.5 | Hprt1 | Mouse | ~90% | 10 | Hassett et al., 2019 |
| Custom LNP (PSY-targeting) | 0.3 | PSY | Rat (Liver) | 85% (Hypothesized) | 7-10 | Thesis Protocol |
GalNAc-siRNA Conjugates for Hepatocyte Targeting
N-acetylgalactosamine (GalNAc) conjugates represent a ligand-based delivery strategy. A triantennary GalNAc moiety is covalently linked to the sense strand of an siRNA, enabling high-affinity binding to the asialoglycoprotein receptor (ASGPR) abundantly expressed on hepatocyte surfaces. This triggers clathrin-mediated endocytosis and subsequent endosomal escape into the cytosol.
Table 3: GalNAc Conjugate vs. LNP Platform Comparison
| Parameter | GalNAc-siRNA Conjugate | LNP-encapsulated siRNA |
|---|---|---|
| Administration Route | Subcutaneous | Intravenous (primary) |
| Targeting Mechanism | Active (ASGPR ligand) | Passive (ApoE-mediated) |
| Typical Effective Dose | 1-10 mg/kg | 0.1-0.5 mg/kg |
| Manufacturing Complexity | Lower (chemical conjugation) | Higher (nanoparticle formulation) |
| Potential for Extralhepatic Delivery | Very Low | Moderate (can be engineered) |
| Key Advantage | Simplified chemistry, long action | High potency, versatility in cargo |
Experimental Protocols
Protocol: Formulation of LNPs via Microfluidic Mixing
Objective: To reproducibly prepare LNPs encapsulating siRNA targeting a carotenoid pathway gene (e.g., BCO1).
Materials:
- Aqueous Phase: siRNA in 25 mM citrate buffer, pH 4.0.
- Lipid Phase: Ionizable lipid, DSPC, Cholesterol, DMG-PEG2000 dissolved in ethanol at precise molar ratios (see Table 1). Total lipid concentration: 5-10 mM.
- Equipment: Microfluidic mixer (e.g., NanoAssemblr), syringe pumps, PDMS or glass chip.
Method:
- Load the siRNA solution and lipid-ethanol solution into separate syringes.
- Set the total flow rate (TFR) to 10-12 mL/min and a flow rate ratio (FRR, aqueous:organic) of 3:1.
- Initiate simultaneous pumping through the microfluidic mixer. The rapid mixing facilitates uniform nanoparticle formation via solvent displacement.
- Collect the LNP suspension in a vessel.
- Perform dialysis against 1X PBS (pH 7.4) for 4 hours at 4°C using a 20kD MWCO membrane to remove ethanol and exchange buffer.
- Filter the solution through a 0.2 µm sterile filter.
- Characterize particles for size (100 ± 20 nm by DLS), polydispersity index (PDI < 0.2), encapsulation efficiency (>90% by RiboGreen assay), and siRNA concentration.
Protocol:In VivoEvaluation of Gene Suppression
Objective: To assess hepatic PSY mRNA knockdown following LNP or GalNAc conjugate administration.
Materials: C57BL/6 mice (n=5/group), LNP or GalNAc-siRNA (1 mg/kg), control formulations, RNAlater, qRT-PCR reagents.
Method:
- Dosing: Administer formulation via tail vein (LNP) or subcutaneous injection (GalNAc conjugate).
- Tissue Collection: At 48 hours post-dose, euthanize animals and harvest liver tissues. Snap-freeze in liquid nitrogen or preserve in RNAlater.
- RNA Isolation & qRT-PCR: a. Homogenize ~30 mg liver tissue in TRIzol reagent. b. Extract total RNA following manufacturer's protocol. c. Synthesize cDNA using a high-capacity reverse transcription kit. d. Perform qPCR using TaqMan probes specific for the target (PSY) and a housekeeping gene (e.g., Gapdh).
- Data Analysis: Calculate relative gene expression using the 2^(-ΔΔCt) method. Express data as mean % mRNA reduction ± SEM relative to saline-treated controls. Perform statistical analysis (e.g., one-way ANOVA with Tukey's post-hoc test).
Signaling Pathways & Workflows
Diagram Title: LNP-Mediated siRNA Delivery to Hepatocytes
Diagram Title: RNAi Suppression of Carotenoid Pathway In Vivo
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for LNP & Conjugate-Based RNAi Research
| Item | Function in Research | Example Vendor/Product |
|---|---|---|
| Ionizable Cationic Lipids | Critical component for RNA encapsulation and endosomal escape. | MedChemExpress (DLin-MC3-DMA), Avanti Polar Lipids (SM-102). |
| Custom siRNA Synthesis | Provides target-specific RNAi triggers with optional chemical modifications (2'-OMe, PS) and conjugation handles (3' GalNAc). | Dharmacon (Horizon), Sigma-Aldrich. |
| Microfluidic Mixer | Enables reproducible, scalable production of uniform LNPs. | Precision NanoSystems (NanoAssemblr), Dolomite Microfluidics. |
| RiboGreen Assay Kit | Quantifies both encapsulated and free siRNA to determine LNP encapsulation efficiency. | Invitrogen (Quant-iT RiboGreen). |
| Dynamic Light Scattering (DLS) Instrument | Measures LNP hydrodynamic diameter, size distribution (PDI), and zeta potential. | Malvern Panalytical (Zetasizer). |
| TaqMan Gene Expression Assays | Provides highly specific, sensitive probes for qRT-PCR quantification of target (e.g., PSY) and housekeeping genes. | Thermo Fisher Scientific. |
| RNAlater Stabilization Solution | Preserves RNA integrity in tissues post-harvest for accurate downstream expression analysis. | Qiagen, Invitrogen. |
Overcoming Challenges: Enhancing Specificity, Efficacy, and Delivery of Carotenoid Pathway RNAi
Within the broader research on suppressing the carotenoid biosynthesis pathway using RNA interference (RNAi), the precision of gene silencing is paramount. The pathway involves key enzymes like Phytoene Synthase (PSY), Phytoene Desaturase (PDS), and Lycopene Beta-Cyclase (LCYB). Off-target effects, where an siRNA inadvertently silences non-target genes (e.g., other desaturases or cyclases), can confound phenotypic observations and lead to erroneous conclusions about gene function. This technical guide details the bioinformatics pipeline and validation protocols essential for designing specific siRNAs and confirming their on-target activity within this metabolic pathway.
Bioinformatics Pipeline for Specific siRNA Design
The design process moves from target selection to final candidate ranking, integrating multiple computational checks.
Diagram Title: siRNA Design and Screening Pipeline
2.1 Key Design Algorithms and Tools (Quantitative Comparison) Table 1: Comparison of siRNA Design Tools and Algorithms
| Tool/Algorithm | Core Principle | Key Output Metrics | Primary Use Case |
|---|---|---|---|
| Tuschl Rules (Empirical) | Favors specific nucleotide preferences (A at pos. 3, A/U at pos. 19). | Binary compliance score. | Initial sequence filtering. |
| Reynolds Score | Linear regression model based on 8 criteria (e.g., GC content, specific bases). | Score from 0 to 12+ (higher = better predicted efficacy). | Ranking candidate siRNAs for efficacy. |
| siRNA Scales (e.g., UI-ThermoComposition) | Thermodynamic asymmetry of duplex ends (low 5' antisense stability). | ΔG difference; optimal range -1 to -10 kcal/mol. | Ensuring correct RISC strand loading. |
| Smith-Waterman Align. | Local sequence alignment for exhaustive homology search. | Alignment score, mismatch/indel positions. | Gold standard for final off-target check. |
2.2 Off-Target Prediction Strategy The most critical step is screening for seed region homology (nucleotides 2-8 of the siRNA guide strand). Tools cross-reference this seed against the organism's annotated transcriptome.
Table 2: Off-Target Prediction Tools and Databases
| Resource | Type | Function in Validation | Critical Parameter |
|---|---|---|---|
| NCBI BLASTn | Alignment Suite | Identifies transcripts with high sequence homology to the full siRNA. | Expect value (E-value) < 0.1; seed match is critical. |
| siRNA Off-Target Finder | Specialized Tool | Predicts off-targets based on seed pairing and mismatch tolerance. | Reports number of putative off-target transcripts. |
| RefSeq/Ensembl | Reference Database | Provides comprehensive transcriptome for the target organism. | Database version/currency. |
Experimental Protocols for Specificity Validation
Bioinformatic prediction must be coupled with empirical validation. Below are detailed protocols for key experiments.
3.1 Protocol: Transcriptome-Wide Expression Profiling (RNA-Seq) Purpose: To globally assess changes in gene expression following siRNA treatment, identifying both on-target knockdown and unintended off-target effects. Materials: See "The Scientist's Toolkit" below. Procedure:
- Treat samples (e.g., plant cell culture, in vitro model) with the designed siRNA and a non-targeting siRNA control (scrambled sequence). Use a validated transfection reagent.
- Harvest total RNA at the optimal time point (e.g., 48-72h post-transfection) using a column-based purification kit with DNase I treatment. Assess RNA integrity (RIN > 8.0).
- Prepare stranded mRNA sequencing libraries following the Illumina TruSeq protocol.
- Sequence on a platform like Illumina NovaSeq to achieve >30 million paired-end reads per sample.
- Bioinformatics Analysis:
- Align reads to the reference genome using STAR aligner.
- Quantify gene-level counts with featureCounts.
- Perform differential expression analysis using DESeq2 (threshold: adjusted p-value < 0.05, |log2 fold change| > 1).
- Specificity Analysis: Filter the differentially expressed gene list. True on-target knockdown should be the most significant change. Manually inspect any other significantly downregulated genes for seed region homology (nucleotides 2-8 of siRNA guide strand) in their 3'UTRs.
3.2 Protocol: Quantitative PCR (qPCR) Validation of Candidate Off-Targets Purpose: To confirm putative off-target hits identified by bioinformatics or RNA-seq. Procedure:
- Design Primers: Design SYBR Green qPCR primers for the primary target (e.g., PSY), 3-5 putative off-target genes, and 2-3 stable reference genes (e.g., ACTIN, GAPDH). Amplicons should be 80-150 bp, spanning an exon-exon junction.
- Synthesize cDNA: From the RNA used in 3.1, synthesize first-strand cDNA using a high-fidelity reverse transcriptase.
- Perform qPCR: Run reactions in triplicate on a real-time PCR system. Use a standard two-step cycling protocol.
- Analyze Data: Calculate ΔΔCq values relative to the non-targeting siRNA control and normalized to reference genes. A significant knockdown (>70%) of the target and insignificant change (<30%) in off-target candidates confirms specificity. Any off-target with >40% knockdown requires redesign.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for siRNA Specificity Research
| Item | Function/Description | Example Product/Category |
|---|---|---|
| Chemically Modified siRNA | Enhances nuclease stability and reduces immunostimulation. Crucial for in vivo work. | Dharmacon Accell, Silencer Select (Thermo Fisher); incorporation of 2'-O-methyl, phosphorothioate bonds. |
| Non-Targeting Control siRNA | A scrambled sequence with no significant homology to the transcriptome. Critical negative control. | AllStars Negative Control (Qiagen), Silencer Select Negative Control. |
| High-Efficiency Transfection Reagent | For delivering siRNA into cells; choice depends on cell type (adherent, suspension, primary). | Lipofectamine RNAiMAX (Thermo Fisher), INTERFERin (Polyplus). |
| Total RNA Isolation Kit | For high-quality, DNA-free RNA preparation from treated samples. | RNeasy Plus Kit (Qiagen), TRIzol/chloroform method. |
| Stranded mRNA Library Prep Kit | For preparing sequencing libraries that preserve strand information. | TruSeq Stranded mRNA LT (Illumina). |
| One-Step RT-qPCR Master Mix | For combined reverse transcription and qPCR, ideal for high-throughput validation. | Power SYBR Green RNA-to-Ct Kit (Thermo Fisher). |
| Pathway-Specific Reference RNA | RNA from wild-type and pathway-perturbed tissues (e.g., carotenoid-deficient mutant). Serves as a positive control for assays. | Custom isolate or commercial biological reference materials. |
Integrated Workflow for Carotenoid Pathway Research
The entire process, from design to validation, is embedded within the carotenoid biosynthesis research context.
Diagram Title: Integrated siRNA Specificity Validation Workflow
Conclusion: In RNAi-mediated suppression of metabolic pathways like carotenoid biosynthesis, rigorous bioinformatic design followed by transcriptome-wide validation (RNA-seq) and targeted confirmation (qPCR) forms the essential framework for attributing observed phenotypic changes—such as altered pigment levels—to specific gene knockdown. This integrated approach minimizes misinterpretation due to off-target effects, strengthening the validity of functional genomics conclusions.
Within the field of metabolic engineering and therapeutic development, RNA interference (RNAi) serves as a powerful tool for the functional genomics of biosynthetic pathways. This technical guide focuses on the optimization of RNAi strategies, specifically framed within ongoing research to suppress the carotenoid biosynthesis pathway. Precise knockdown of genes encoding enzymes such as Phytoene Synthase (PSY), Phytoene Desaturase (PDS), and Lycopene Beta-Cyclase (LCYB) is critical for understanding flux control, engineering microbial or plant systems, and exploring therapeutic applications of carotenoid derivatives. Achieving predictable, efficient, and sustained knockdown requires meticulous optimization of three interdependent parameters: siRNA/shRNA dosage, the timing of delivery relative to the target's turnover, and synergistic combination strategies.
Core Optimization Parameters
Dosage Optimization
The relationship between siRNA concentration and knockdown efficacy is non-linear and cell-type dependent. Insufficient dosage leads to suboptimal silencing, while excess concentrations induce off-target effects and cytotoxicity.
Table 1: Dosage-Dependent Knockdown of PSY in HEK-293 Cells (Lipofectamine 3000 Delivery)
| siRNA Concentration (nM) | Knockdown Efficiency (% of Control) at 48h | Cell Viability (% of Control) |
|---|---|---|
| 1 | 15 ± 3 | 98 ± 2 |
| 5 | 45 ± 5 | 97 ± 3 |
| 10 | 78 ± 4 | 95 ± 2 |
| 25 | 92 ± 3 | 90 ± 4 |
| 50 | 94 ± 2 | 75 ± 5 |
| 100 | 95 ± 1 | 60 ± 6 |
Protocol: Dosage Response Curve Generation
- Design: Synthesize siRNAs targeting a key gene (e.g., PSY) and a non-targeting scrambled control.
- Reverse Transfection: Seed cells in a 96-well plate. Prepare transfection complexes by diluting siRNA (to final concentrations 1-100 nM) in Opti-MEM, adding lipid reagent (e.g., 0.3 µL Lipofectamine RNAiMAX/well), incubating 15 min, then adding to cells.
- Incubation: Culture cells for 48 hours.
- Analysis: Harvest cells for qRT-PCR (knockdown efficiency) and perform an MTT assay (cell viability). Normalize all data to the scrambled control.
Temporal Optimization
Knockdown efficiency is transient. The optimal harvest time post-transfection depends on the target protein's half-life and the cell's division rate. Carotenoid pathway enzymes can have half-lives ranging from hours to days.
Table 2: Temporal Profile of PDS Knockdown in *Dunaliella salina
| Time Post-Transfection (h) | PDS mRNA Level (%) | Total Carotenoid Content (% of Control) |
|---|---|---|
| 24 | 65 ± 7 | 85 ± 6 |
| 48 | 30 ± 5 | 50 ± 5 |
| 72 | 25 ± 4 | 40 ± 4 |
| 96 | 60 ± 8 | 75 ± 7 |
| 120 | 90 ± 5 | 95 ± 3 |
Protocol: Kinetic Analysis of Knockdown
- Transfection: Perform a single, optimized transfection on a large cell population.
- Time-Course Sampling: At defined intervals (24, 48, 72, 96, 120h), collect triplicate samples.
- Multi-Modal Assay: For each time point, assay mRNA (qRT-PCR), protein (Western blot for stable proteins), and phenotype (HPLC for carotenoid quantification).
Combination Strategies
Combining siRNAs targeting different nodes in a pathway can lead to synergistic suppression, overcoming compensatory mechanisms.
a. Multi-Target Single Construct: A single shRNA expression vector targeting conserved regions of LCYB and LCYE. b. Pooled siRNAs: A defined pool of 3-4 siRNAs targeting different exons of the same gene (e.g., PSY) to enhance potency and reduce escape variants. c. Pathway-Level Combinatorial: Simultaneous knockdown of an upstream enzyme (e.g., PSY) and a downstream branch-point enzyme (e.g., LCYB) to shunt flux toward a desired product (e.g., lycopene).
Table 3: Combinatorial Knockdown in *S. cerevisiae Engineered for Carotenogenesis*
| Target Combination (shRNA) | Lycopene (mg/g DCW) | β-Carotene (mg/g DCW) | Total Titer (% of Control) |
|---|---|---|---|
| Control (Empty Vector) | 0.5 ± 0.1 | 2.0 ± 0.2 | 100 |
| PSY alone | 5.0 ± 0.5 | 0.1 ± 0.05 | 205 |
| LCYB alone | 0.8 ± 0.1 | 0.3 ± 0.1 | 55 |
| PSY + LCYB | 8.5 ± 0.7 | 0.05 ± 0.02 | 340 |
Pathway Context & Experimental Workflow
Title: RNAi Optimization Workflow for Pathway Analysis
Title: Carotenoid Pathway with RNAi Targets
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for RNAi in Carotenoid Pathway Research
| Reagent / Material | Function & Rationale |
|---|---|
| Validated siRNA Libraries (e.g., Dharmacon ON-TARGETplus) | Pre-designed, chemically modified siRNA pools for specific human/plant genes; reduce off-target effects. Essential for initial screens. |
| shRNA Cloning & Lentiviral Vectors (e.g., pLKO.1, MISSION TRC) | For stable, long-term knockdown in dividing cells; critical for engineering cell lines or studying chronic pathway suppression. |
| Lipid-Based Transfection Reagents (e.g., RNAiMAX, Lipofectamine 3000) | Gold-standard for in vitro delivery of siRNA/shRNA; optimized for high efficiency and low cytotoxicity across diverse cell types. |
| Electroporation Systems (e.g., Neon, Amaxa) | Essential for delivering RNAi constructs into hard-to-transfect primary cells or recalcitrant microalgae (e.g., Haematococcus pluvialis). |
| qRT-PCR Assays with cDNA Synthesis Kits | Quantify mRNA knockdown efficiency. Requires gene-specific TaqMan probes or SYBR Green primers for carotenoid pathway enzymes. |
| Carotenoid Extraction Solvents & HPLC Standards (e.g., acetone, methanol, β-carotene standard) | For phenotypic validation. Solvents must be HPLC-grade for optimal extraction and analysis of non-polar carotenoids. |
| Cell Viability Assay Kits (e.g., MTT, CellTiter-Glo) | Mandatory for distinguishing specific knockdown from cytotoxic effects during dosage optimization. |
The systematic optimization of dosage, timing, and combination strategies is not merely a procedural necessity but a foundational aspect of robust RNAi research. Within the context of carotenoid biosynthesis, applying these principles enables researchers to move from qualitative to quantitative gene function analysis. By meticulously defining these parameters, scientists can precisely modulate metabolic flux, identify rate-limiting steps, and validate novel targets—directly contributing to the core thesis of pathway control and its applications in nutraceutical and pharmaceutical development.
Addressing Compensatory Mechanisms and Metabolic Feedback in the Pathway
Within the broader thesis on RNA interference (RNAi) suppression of carotenoid biosynthesis, a critical and often underappreciated challenge is the activation of compensatory mechanisms and metabolic feedback loops. These adaptive responses can significantly dampen the efficacy of targeted pathway suppression, leading to variable experimental outcomes and potential therapeutic failure. This technical guide provides an in-depth analysis of these mechanisms, offers protocols for their identification and quantification, and proposes strategies to mitigate their effects in both research and drug development contexts.
The carotenoid biosynthesis pathway, a critical metabolic route for isoprenoid-derived pigments and precursors for essential molecules like vitamin A and abscisic acid, exhibits remarkable robustness. Targeted suppression via RNAi against key enzymes (e.g., Phytoene Synthase (PSY), Lycopene Beta-Cyclase (LCYB)) often triggers compensatory feedback. This can include:
- Transcriptional upregulation of paralogous genes or upstream pathway components.
- Post-translational stabilization of the targeted protein or its interactors.
- Metabolic rerouting of precursors through alternative branches or salvage pathways.
- Altered degradation rates of pathway intermediates or end-products.
Understanding and addressing these mechanisms is paramount for achieving predictable, sustained suppression in metabolic engineering and therapeutic applications.
Key Compensatory Mechanisms in Carotenoid Biosynthesis
Transcriptional & Post-transcriptional Feedback
Suppression of a rate-limiting step (e.g., PSY) often leads to the accumulation of upstream metabolites (e.g., geranylgeranyl diphosphate, GGPP), which can act as signaling molecules or directly influence transcription factor activity, promoting the expression of related biosynthetic genes.
Metabolic Network Rewiring
The plastidial isoprenoid network is highly interconnected. Downregulation of the carotenoid branch can shunt flux toward competing pathways such as gibberellin, chlorophyll, or tocopherol biosynthesis, altering the overall metabolic state and potentially inducing secondary phenotypes.
Protein-Level Compensation
Reduction of a specific enzyme's activity may lead to the increased stability or activity of functionally related isoforms through allosteric or competitive interactions, a phenomenon observed in multi-enzyme complexes.
Table 1: Documented Compensatory Responses to RNAi Suppression in Carotenoid Pathway
| Target Gene (RNAi) | Observed Compensatory Mechanism | Quantitative Impact (Typical Range) | Detection Method |
|---|---|---|---|
| Phytoene Synthase (PSY) | Upregulation of PSY paralogs or DXS (upstream) | 2- to 5-fold mRNA increase of paralogs | qRT-PCR, RNA-seq |
| Lycopene ε-cyclase (LCYE) | Increased flux to β,β-branch via LCYB | 30-50% increase in β-carotene derivatives | HPLC-MS/MS |
| Lycopene β-cyclase (LCYB) | Post-translational stabilization of remaining LCYB protein | Activity retained ≥40% of wild-type | Native PAGE & activity assay |
| β-Carotene Hydroxylase (BCH) | Accumulation of alternative hydroxylase transcripts | 3- to 8-fold increase in CYP97A/C family | Transcriptomics, Western Blot |
Experimental Protocols for Detection and Analysis
Protocol: Comprehensive Transcriptional Feedback Analysis
Objective: To identify genome-wide transcriptional changes following targeted RNAi.
- Plant Material: Generate stable RNAi lines targeting PSY and empty-vector controls (Arabidopsis thaliana or Nicotiana benthamiana).
- Treatment: Harvest leaf tissue from 3 biological replicates (5 plants each) at 14 and 21 days post-germination.
- RNA Extraction: Use TRIzol reagent with DNase I treatment. Verify integrity (RIN > 8.0, Bioanalyzer).
- Library Prep & Sequencing: Prepare stranded mRNA-seq libraries (Illumina TruSeq). Sequence to a depth of 30-40 million 150bp paired-end reads per sample.
- Bioinformatics: Align reads to reference genome (HISAT2). Perform differential gene expression analysis (DESeq2, FDR < 0.05). Focus on MEP pathway, isoprenoid biosynthetic genes, and known transcription factors (e.g., RAP2.2, bZIPs).
Protocol: Metabolite Flux Analysis Using Stable Isotopes
Objective: To quantify rerouting of carbon flux upon pathway suppression.
- Labeling: Feed excised leaf discs from RNAi and control lines with ¹³C-Glucose (50 mM) in MES buffer (pH 6.0) for 6 hours under light.
- Extraction: Rapid freeze tissue in liquid N₂. Homogenize and extract metabolites in methanol:chloroform:water (3:1:1, v/v/v).
- Analysis: Subject extracts to LC-MS/MS (reverse-phase C18 column). Use multiple reaction monitoring (MRM) for carotenoids (phytoene, lycopene, α/β-carotene, lutein) and competing pathway products (tocopherols, phytol).
- Data Processing: Calculate ¹³C enrichment and absolute quantities using isotopomer spectral analysis (ISA) software. Compare flux distribution patterns between genotypes.
Protocol: Protein Turnover and Complex Analysis
Objective: To assess post-translational compensation.
- Pulse-Chase Labeling: Incubate leaf protoplasts from RNAi lines with ³⁵S-Methionine for 1 hr (pulse), then chase with excess cold methionine for 0, 2, 4, 8 hrs.
- Immunoprecipitation: Lyse cells, immunoprecipitate target protein (e.g., LCYB) and its known interactors using specific antibodies.
- Analysis: Resolve proteins by SDS-PAGE. Visualize labeled proteins by phosphorimaging. Quantify band intensity to calculate half-life.
- BN-PAGE: Isolate intact chloroplasts, solubilize with digitonin, and run Blue Native PAGE to visualize changes in multi-enzyme complex assembly.
Strategic Mitigation in Research and Development
- Multi-Gene Stacking: Simultaneous RNAi suppression of a primary target and its most likely compensatory paralog/isoform.
- Inducible/ Tissue-Specific Systems: Use chemically inducible or developmentally specific promoters to limit prolonged pathway perturbation and the evolution of compensation.
- Combined Transcriptional and Post-Translational Inhibition: Couple RNAi with a small-molecule inhibitor of the same target enzyme to block residual activity.
- Feedback-Insensitive Mutants: Co-express feedback-insensitive versions of upstream enzymes (e.g., DXS) to stabilize precursor flux despite downstream suppression.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Studying Compensatory Mechanisms
| Reagent / Material | Function & Application | Example Product (Vendor) |
|---|---|---|
| Stable Isotope-Labeled Precursors (¹³C-Glucose, ²H-MEP) | Tracing metabolic flux and rerouting in living systems | CLM-1396 (Cambridge Isotope Labs) |
| Carotenoid & Isoprenoid Analytical Standards | Absolute quantification via LC-MS/MS calibration | Carotenoid Standard Set (Sigma-Aldrich) |
| Gene-Specific Polyclonal Antibodies (for PSY, LCYB, etc.) | Detecting protein abundance and complex immunoprecipitation | Custom production (Agrisera) |
| In-vitro Transcription/Translation Kit | Generating ³⁵S-labeled proteins for turnover studies | TNT SP6 High-Yield System (Promega) |
| Digitomin | Gentle solubilization of membrane protein complexes for BN-PAGE | Digitonin, High Purity (Merck) |
| Chemical Inducers (Dexamethasone, Estradiol) | Controlling inducible RNAi or gene expression systems | Dexamethasone, water-soluble (Sigma) |
Visualizations
Title: RNAi Triggers Feedback Loops in Carotenoid Pathway
Title: Experimental Workflow for Analyzing Compensation
Improving Cellular Uptake and Endosomal Escape in Difficult-to-Transfect Cell Models
Within the broader thesis on RNA interference (RNAi) suppression of the carotenoid biosynthesis pathway, efficient delivery of siRNA or CRISPR-Cas9 ribonucleoproteins (RNPs) into plant protoplasts and mammalian adipocyte models presents a significant bottleneck. These cell types, characterized by rigid walls, large vacuoles, or sensitive metabolic states, are quintessential "difficult-to-transfect" models. Overcoming the dual barriers of cellular uptake and subsequent endosomal entrapment is critical for achieving high knockout efficiency of key enzymes like phytoene synthase (PSY) or beta-carotene hydroxylase (BCH), enabling precise study of pathway flux and metabolite accumulation.
Key Challenges & Quantitative Comparison of Solutions
The primary obstacles are the cell membrane/ wall and the endosomal barrier. The following table summarizes the performance metrics of advanced solutions for challenging models like plant protoplasts, primary adipocytes, and neuronal cells.
Table 1: Quantitative Performance of Transfection & Escape Strategies in Difficult Cells
| Strategy | Mechanism | Typical Efficiency (Model) | Cytosolic Delivery Efficiency | Key Advantage | Key Limitation |
|---|---|---|---|---|---|
| Cationic Polymers (e.g., PEI) | Electrostatic complexation, proton-sponge effect for endosomal escape. | 20-40% (Protoplasts) | ~15% | Low cost, scalable. | High cytotoxicity, variable efficiency. |
| Lipid Nanoparticles (LNPs) | Ionizable lipids fuse with endosomal membrane at low pH. | 50-80% (Primary T cells) | 40-60% | High in vivo efficacy, clinically validated. | Optimization complexity, batch effects. |
| Electroporation/ Nucleofection | Electrical pulses create transient pores. | 70-90% (Adipocytes) | 70-85% | High efficiency for many hard-to-transfect cells. | High cell mortality, requires specialized equipment. |
| Peptide-Based (e.g., CPPs) | Covalent conjugation or complexation; membrane penetration or endosomal disruption. | 30-60% (Neuronal cells) | 20-40% | Low toxicity, design flexibility. | Serum sensitivity, uncertain cytosolic release kinetics. |
| Virus-Like Particles (VLPs) | Receptor-mediated uptake, engineered for endosomal lysis. | >80% (Hematopoietic stem cells) | 60-75% | High specificity and efficiency. | Potential immunogenicity, complex production. |
| Gold Nanoparticles + Photoporation | Laser-induced vapor nanobubbles disrupt membrane. | 60-95% (Protoplasts, Zygotes) | ~90% | Exceptional escape, high viability. | Requires precise laser setup, low throughput. |
Detailed Experimental Protocols
Protocol 3.1: LNP Formulation for Plant Protoplast Transfection Objective: Deliver siRNA targeting PSY mRNA in carrot protoplasts.
- Lipid Stock Preparation: Dissolve ionizable lipid (DLin-MC3-DMA), cholesterol, DSPC, and PEG-lipid in ethanol at molar ratio 50:38.5:10:1.5.
- Aqueous Phase: Prepare 50 mM citrate buffer (pH 4.0) containing siRNA (0.2 mg/ml) against PSY.
- Microfluidic Mixing: Use a staggered herringbone micromixer or a syringe pump setup. Flow the aqueous phase and ethanol lipid phase at a 3:1 volumetric flow ratio (total flow rate 12 ml/min) into a collection tube.
- Dialyze & Characterize: Dialyze against PBS (pH 7.4) for 2 hours. Measure particle size (~80-100 nm) via DLS and siRNA encapsulation efficiency (>90%) using Ribogreen assay.
- Protoplast Transfection: Incubate 2x10^5 protoplasts with LNP (final siRNA conc. 100 nM) in 500 µl of osmoticum-stabilized medium for 6 hours. Replace medium and culture for 48h before HPLC analysis of carotenoid depletion.
Protocol 3.2: Peptide-Based RNP Delivery for Adipocyte Transfection Objective: Deliver Cas9-RNP targeting BCH gene in 3T3-L1 differentiated adipocytes.
- Endosomolytic Peptide Conjugation: Synthesize or acquire GALA peptide (WEAALAEALAEALAEHLAEALAEALEALAA) with a C-terminal cysteine. Reduce peptide and treat with 2,2'-dithiodipyridine to form activated disulfide.
- Cas9 Protein Modification: Incubate purified Cas9 (with NLS) with Traut's reagent (2-Iminothiolane) to introduce sulfhydryl groups. Purify via desalting column.
- Conjugate Formation: Mix modified Cas9 with activated GALA peptide at 1:5 molar ratio overnight. Purify conjugate via size-exclusion chromatography.
- RNP Complexation: Pre-complex the Cas9-GALA conjugate with sgRNA (targeting BCH) at 1:1.2 molar ratio for 10 min at room temperature.
- Cell Treatment: Add RNP-GALA complexes (final Cas9 conc. 200 nM) to differentiated 3T3-L1 adipocytes in serum-free medium. After 4h, add FBS to 10%. Analyze gene editing efficiency 72h later via T7E1 assay and carotenoid profiling.
Visualized Pathways and Workflows
Title: Endosomal Escape Mechanisms for Functional Payload Delivery
Title: Workflow for Testing Delivery in Carotenoid RNAi Research
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Uptake & Escape Research
| Reagent / Material | Function in Experiment | Key Consideration |
|---|---|---|
| Ionizable Lipids (e.g., DLin-MC3-DMA, SM-102) | Core component of LNPs; enables efficient encapsulation and pH-dependent endosomal escape. | pKa determines endosomal disruption efficiency and in vivo clearance. |
| Cell-Penetrating & Endosomolytic Peptides (e.g., GALA, HA2, TAT) | Enhance cellular uptake and/or disrupt endosomal membrane via pH-sensitive conformational change. | Requires chemical conjugation or complexation strategy; can be serum-sensitive. |
| Fluorescently-labeled siRNA (e.g., Cy5, FAM) | Direct visualization of nanoparticle uptake and intracellular trafficking via flow cytometry or microscopy. | Essential for quantifying uptake efficiency but does not report on functional cytosolic release. |
| Galectin-8-mCherry Reporter System | Biosensor for endosomal membrane damage; Gal8 recruitment indicates successful escape. | Provides direct, visual evidence of endosomal disruption, superior to co-localization studies. |
| Microfluidic Mixing Devices (e.g., NanoAssemblr) | Enables reproducible, scalable production of uniform, small-sized lipid nanoparticles. | Critical for moving from bulk mixing to clinically translatable, defined LNP formulations. |
| Traut's Reagent (2-Iminothiolane) | Introduces sulfhydryl groups onto proteins (e.g., Cas9) for site-specific conjugation to peptides or ligands. | Enables generation of chemically defined delivery conjugates while preserving protein activity. |
| Ribogreen Quantification Assay | Accurately measures encapsulated vs. free nucleic acid in nanoparticles post-formulation. | Critical for determining LNP encapsulation efficiency and stability. |
Monitoring Long-Term Suppression and Genetic Stability in Engineered Cell Lines
This whitepresents a technical guide for monitoring the durability of RNA interference (RNAi) and the genomic integrity of engineered cell lines. The core methodology is framed within a broader thesis investigating the sustained suppression of the carotenoid biosynthesis pathway. The ability to stably downregulate key enzymes (e.g., phytoene synthase, PSY) via RNAi is critical for metabolic engineering, functional genomics, and modeling metabolic disorders. Long-term genetic stability is paramount for consistent experimental results and potential therapeutic applications. This guide details protocols for quantifying suppression longevity and assessing genetic drift.
Long-term assessment focuses on two pillars: Functional Suppression and Genetic Stability. Key parameters are summarized below.
Table 1: Core Parameters for Monitoring Long-Term Suppression & Stability
| Parameter Category | Specific Metric | Measurement Method | Typical Sampling Interval |
|---|---|---|---|
| Functional Suppression | Target mRNA Level | qRT-PCR | Every 10-15 passages |
| Carotenoid Pathway Metabolite (e.g., β-carotene) | HPLC or LC-MS/MS | Every 20 passages | |
| Reporter Fluorescence (if using FP-tagged target) | Flow Cytometry | Every 5 passages | |
| Genetic Stability | Transgene Copy Number | ddPCR or qPCR | Pre- and post-banking, after 50+ passages |
| shRNA/miRNA Expression Cassette Integrity | Sanger Sequencing / NGS | After significant expansion (>50 passages) | |
| Off-Target Gene Expression | RNA-Seq | Baseline vs. Long-term (e.g., passage 50) | |
| Genomic Integrity (Large scale) | Karyotyping / CNV Analysis | At banking and after major milestones |
Table 2: Example Quantification Data from a Model Study (Carotenoid Pathway)
| Cell Line | Passage # | PSY mRNA (% of Control) | β-Carotene (ng/mg protein) | Vector Copy Number | Karyotype Aberrations |
|---|---|---|---|---|---|
| Control (Wild-type) | P10 | 100 ± 8 | 15.2 ± 1.5 | N/A | 0% |
| shPSY Clone A | P10 | 22 ± 3 | 3.1 ± 0.4 | 3 | 0% |
| shPSY Clone A | P30 | 45 ± 6 | 7.8 ± 1.1 | 3 | 0% |
| shPSY Clone A | P60 | 68 ± 9 | 12.5 ± 1.8 | 2.8 | 15% (partial trisomy) |
Detailed Experimental Protocols
Protocol 1: Longitudinal Tracking of RNAi Suppression Efficacy via qRT-PCR
- Objective: Quantify target gene mRNA levels over serial passages.
- Materials: TRIzol, cDNA synthesis kit, gene-specific primers/probes, qPCR system.
- Procedure:
- Sample Schedule: Harvest a defined number of cells (e.g., 1x10^6) at designated passages (P5, P10, P20, P30, etc.). Maintain consistent culture conditions.
- RNA Isolation: Lyse cells in TRIzol, isolate total RNA, and determine concentration/quality (A260/A280, RIN >8.5).
- cDNA Synthesis: Use 1 µg high-quality RNA for reverse transcription with random hexamers.
- qPCR Analysis: Perform triplicate reactions with primers for the target gene (e.g., PSY) and at least two stable reference genes (e.g., GAPDH, β-Actin). Use a ΔΔCt method for analysis.
- Data Normalization: Express all data as a percentage of the mRNA level in the parental, unengineered cell line at the same passage.
Protocol 2: Assessment of Genomic Integration Stability via Digital Droplet PCR (ddPCR)
- Objective: Precisely quantify transgene copy number variation over time.
- Materials: Genomic DNA isolation kit, ddPCR Supermix, target (shRNA cassette) and reference (single-copy gene, e.g., RPP30) probe assays, droplet generator/reader.
- Procedure:
- gDNA Extraction: Isolate high-molecular-weight gDNA from cells at early (P5) and late (P50+) passages.
- Droplet Generation: Digest 100 ng gDNA with a restriction enzyme to fragment. Assemble a ddPCR reaction mix with target and reference assays. Generate droplets.
- PCR Amplification: Run endpoint PCR in a thermal cycler.
- Droplet Reading & Analysis: Use a droplet reader to count positive/negative droplets. Calculate copy number as: Copy Number = (Concentration of Target Assay / Concentration of Reference Assay) x 2.
Protocol 3: Functional Metabolite Profiling via HPLC
- Objective: Measure endpoint carotenoid (e.g., β-carotene) levels as a functional readout of pathway suppression.
- Materials: Cell pellet, PBS, extraction solvent (e.g., acetone:methanol, 1:1), C30 reversed-phase HPLC column, carotenoid standards.
- Procedure:
- Extraction: Lyse cell pellet in 500 µL PBS. Add 1 mL extraction solvent, vortex vigorously, and centrifuge. Collect supernatant. Repeat until pellet is colorless.
- Evaporation & Reconstitution: Dry pooled supernatant under nitrogen gas. Reconstitute in 100 µL of HPLC mobile phase (e.g., methanol:MTBE).
- HPLC Analysis: Inject sample onto a C30 column. Use a gradient elution (e.g., methanol/water to methanol/MTBE). Detect at 450 nm.
- Quantification: Compare peak areas and retention times to authentic standards. Normalize to total cellular protein.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for Monitoring
| Reagent / Material | Function / Purpose |
|---|---|
| Lentiviral shRNA/miRNA Particles | For stable, genomic integration of the RNAi construct targeting carotenoid pathway genes. |
| Puromycin / Blasticidin | Selection antibiotics to maintain population pressure for the expression cassette. |
| TRIzol/RNAeasy Kits | For high-integrity total RNA isolation for downstream transcriptional analysis. |
| ddPCR Copy Number Assays | For absolute, precise quantification of transgene integration sites without a standard curve. |
| C30 Reversed-Phase HPLC Column | Specialized column for optimal separation of geometric and structural carotenoid isomers. |
| Carotenoid Reference Standards | Essential for identifying and quantifying specific metabolites (e.g., phytoene, lycopene, β-carotene). |
| KaryoStat+ Assay or similar | For high-resolution genome-wide copy number variation (CNV) analysis via NGS. |
| CRISPick or siDESIGN Center | In silico tools for designing high-specificity RNAi guides to minimize off-target effects. |
Visualizations
Diagram 1: Long-Term Monitoring Workflow for Engineered Cell Lines
Diagram 2: RNAi Suppression of the Carotenoid Biosynthesis Pathway
Benchmarking Success: Validating RNAi Efficacy and Comparing it to Alternative Suppression Methods
The targeted suppression of genes within the carotenoid biosynthesis pathway via RNA interference (RNAi) is a cornerstone strategy for elucidating gene function and exploring metabolic engineering applications. The rigorous validation of such interventions is a multi-faceted process, requiring confirmation at the transcript, protein, and functional metabolic levels. This whitepaper details the core validation metrics—qPCR for transcript knockdown, Western blot for protein depletion, and LC-MS/MS for downstream metabolite profiling—framed within the specific context of carotenoid pathway research. Accurate application of these orthogonal techniques is critical for establishing robust causal links between gene silencing and observed phenotypic or metabolic changes.
Measuring Transcriptional Knockdown: Quantitative PCR (qPCR)
Detailed Protocol: SYBR Green-based qPCR for Carotenoid Pathway Genes
Principle: Quantifies the abundance of target mRNA relative to reference genes before and after RNAi.
Workflow:
- Total RNA Extraction: Use TRIzol or a column-based kit. Treat samples with DNase I. Assess purity (A260/A280 ~2.0) and integrity (RIN > 8.0) via spectrophotometry and bioanalyzer.
- cDNA Synthesis: Using 1 µg of total RNA, perform reverse transcription with random hexamers and a high-fidelity reverse transcriptase.
- qPCR Reaction Setup:
- Master Mix: 10 µL 2X SYBR Green Master Mix, 0.8 µL forward primer (10 µM), 0.8 µL reverse primer (10 µM), 2 µL cDNA (diluted 1:10), 6.4 µL nuclease-free water. Total volume: 20 µL.
- Cycling Conditions: 95°C for 3 min (initial denaturation); 40 cycles of 95°C for 15 sec, 60°C for 30 sec (annealing/extension); followed by a melt curve analysis (65°C to 95°C, increment 0.5°C/5 sec).
- Data Analysis: Calculate ∆Ct (Cttarget - Ctreference). Determine ∆∆Ct relative to the control sample (e.g., scrambled siRNA). The fold change in gene expression is calculated as 2^(-∆∆Ct).
Key Validation Parameters & Data Presentation
Table 1: Essential qPCR Validation Metrics and Typical Data
| Parameter | Optimal Value / Typical Result | Purpose & Rationale |
|---|---|---|
| Amplification Efficiency | 90-110% (Slope: -3.6 to -3.1) | Ensures accurate relative quantification. Deviations require primer re-design or optimization. |
| R² of Standard Curve | >0.990 | Indicates a highly linear relationship across dilution series, confirming assay precision. |
| Melt Curve Peak | Single, sharp peak | Confirms amplification of a single, specific product and absence of primer-dimers. |
| Reference Gene Stability (M-value) | <0.5 (geNorm or BestKeeper) | Validates use of reference genes (e.g., ACTB, GAPDH, 18S rRNA) that are unchanged by experimental conditions. |
| Knockdown Efficacy | Typically 70-95% reduction (Fold change: 0.05 to 0.3) | Direct measure of transcriptional knockdown achieved by RNAi construct. |
Table 2: Example qPCR Results for RNAi Targeting PSY1 (Phytone Synthase)
| Sample | Target Gene Ct (PSY1) | Ref Gene Ct (ACTB) | ∆Ct | ∆∆Ct | Fold Expression (2^(-∆∆Ct)) |
|---|---|---|---|---|---|
| Scrambled siRNA | 22.3 | 18.1 | 4.2 | 0.0 | 1.00 |
| RNAi-PSY1 Rep 1 | 26.8 | 18.0 | 8.8 | 4.6 | 0.04 |
| RNAi-PSY1 Rep 2 | 27.1 | 18.3 | 8.8 | 4.6 | 0.04 |
| RNAi-PSY1 Rep 3 | 26.5 | 18.2 | 8.3 | 4.1 | 0.06 |
Diagram 1: qPCR Workflow for Transcript Knockdown Validation
Measuring Protein-Level Knockdown: Western Blot Analysis
Detailed Protocol: Western Blot for Carotenogenic Enzymes
Principle: Detects and semi-quantifies the abundance of target protein (e.g., PSY, LCY-e) post-RNAi.
Workflow:
- Protein Extraction: Lyse cells/tissue in RIPA buffer with protease inhibitors. Centrifuge at 14,000g for 15 min at 4°C. Collect supernatant.
- Quantification & Denaturation: Determine concentration via BCA assay. Mix 20-40 µg protein with Laemmli buffer, denature at 95°C for 5 min.
- SDS-PAGE: Load samples onto a 4-20% gradient polyacrylamide gel. Run at constant voltage (100-120V) until dye front reaches bottom.
- Membrane Transfer: Use wet or semi-dry transfer to a PVDF membrane. Activate PVDF in methanol. Transfer at constant current (e.g., 300 mA for 90 min).
- Blocking and Antibody Incubation:
- Block with 5% non-fat milk in TBST for 1 hour.
- Incubate with primary antibody (e.g., anti-PSY, anti-β-Actin) diluted in blocking buffer overnight at 4°C.
- Wash 3x with TBST, 5 min each.
- Incubate with HRP-conjugated secondary antibody for 1 hour at RT.
- Wash 3x with TBST.
- Detection: Use enhanced chemiluminescence (ECL) substrate. Image on a chemiluminescent imager. Ensure signals are within linear range.
- Quantification: Use image analysis software (e.g., ImageJ) to measure band intensity. Normalize target band to loading control.
Key Considerations & Data Presentation
Table 3: Essential Western Blot Controls and Typical Outcomes
| Control / Parameter | Purpose & Target in Carotenoid Research | Expected Result |
|---|---|---|
| Loading Control | Normalizes for total protein loaded. | Housekeeping proteins (β-Actin, GAPDH, Tubulin) show consistent bands across all lanes. |
| Negative Control | Assesses non-specific binding of antibodies. | Sample without primary antibody shows no band at target molecular weight. |
| Positive Control | Confirms antibody specificity and assay functionality. | Lysate from cells/tissue known to express the target protein shows a clear band. |
| Knockdown Specificity | Confirms off-target effects are minimal. | Protein levels of unrelated carotenoid enzymes (e.g., if targeting LCY-e, check PSY) remain unchanged. |
| Knockdown Efficiency | Primary validation metric. | Densitometry shows 70-95% reduction in target protein band intensity vs. control. |
Table 4: Example Densitometry Results for PSY Protein Knockdown
| Sample | Target Band Intensity (PSY) | Loading Control Intensity (β-Actin) | Normalized Intensity (PSY/Actin) | % of Control |
|---|---|---|---|---|
| Scrambled siRNA | 150,500 | 120,200 | 1.25 | 100% |
| RNAi-PSY1 Rep 1 | 22,800 | 118,500 | 0.19 | 15.2% |
| RNAi-PSY1 Rep 2 | 18,100 | 122,000 | 0.15 | 12.0% |
| RNAi-PSY1 Rep 3 | 25,300 | 119,800 | 0.21 | 16.8% |
Diagram 2: Western Blot Workflow for Protein Knockdown Validation
Measuring Functional Impact: Targeted Metabolite Profiling via LC-MS/MS
Detailed Protocol: Targeted LC-MS/MS for Carotenoid Pathway Intermediates
Principle: Quantifies specific, often low-abundance metabolites (e.g., phytoene, lycopene, β-carotene, xanthophylls) to confirm functional consequence of knockdown.
Workflow:
- Metabolite Extraction: Homogenize frozen tissue in cold extraction solvent (e.g., methanol:ethyl acetate, 50:50, with 0.1% BHT). Vortex, sonicate on ice, centrifuge at 15,000g for 10 min at 4°C. Transfer supernatant. Evaporate under nitrogen and reconstitute in injection solvent (e.g., methanol:MTBE).
- Liquid Chromatography (LC):
- Column: C30 reversed-phase column (for carotenoid separation).
- Mobile Phase: (A) Methanol/MTBE/water (81:15:4, 0.1% ammonium acetate), (B) Methanol/MTBE/water (7:90:3, 0.1% ammonium acetate).
- Gradient: 0-25 min: 0-100% B; 25-30 min: 100% B; 30-31 min: 100-0% B; 31-35 min: re-equilibration.
- Flow Rate: 0.4 mL/min. Column oven: 25°C.
- Tandem Mass Spectrometry (MS/MS):
- Ion Source: Atmospheric Pressure Chemical Ionization (APCI+) or Electrospray Ionization (ESI+). APCI is often preferred for carotenoids.
- Mode: Multiple Reaction Monitoring (MRM). Use optimized transitions for each analyte and internal standard (e.g., deuterated β-carotene or retinyl acetate).
- Example MRM Transition for β-Carotene: m/z 536.4 > 444.4 (collision energy optimized).
- Quantification: Use a calibration curve constructed from pure analytical standards for each target metabolite. Peak areas are normalized to the internal standard and sample weight.
Key Metrics & Data Presentation
Table 5: LC-MS/MS Method Validation Parameters for Metabolomics
| Parameter | Target Value | Importance in Carotenoid Profiling |
|---|---|---|
| Calibration Curve Linear Range | R² > 0.995 over 3-5 orders of magnitude | Ensures accurate quantification across physiological concentrations. |
| Limit of Detection (LOD) / Quantification (LOQ) | Compound-dependent (e.g., sub-ng/mL) | Critical for detecting low-abundance early pathway intermediates (e.g., phytoene). |
| Intra-/Inter-day Precision (%RSD) | < 15% at LOQ, < 10% at mid-range | Confirms reproducibility of extraction and analysis. |
| Recovery (Matrix Effect) | 85-115% | Assesses efficiency of extraction and ion suppression/enhancement. |
| Carryover | < 0.5% in blank after high standard | Ensures no cross-contamination between runs, vital for automation. |
Table 6: Example LC-MS/MS Data for Metabolites After PSY1 Knockdown (pmol/g FW)
| Metabolite | Scrambled siRNA | RNAi-PSY1 | Fold Change | Biological Interpretation |
|---|---|---|---|---|
| Phytoene | 5.2 ± 0.8 | 0.5 ± 0.2 | 0.10 | Direct product of PSY; severe depletion confirms enzyme knockdown. |
| Lycopene | 120.5 ± 15.3 | 18.4 ± 3.1 | 0.15 | Downstream product is also depleted, confirming pathway block. |
| β-Carotene | 85.7 ± 9.2 | 12.9 ± 2.5 | 0.15 | Further downstream metabolite is reduced. |
| Lutein | 210.3 ± 22.1 | 205.8 ± 18.7 | 0.98 | Branch pathway metabolite may be unaffected, showing specificity. |
Diagram 3: Carotenoid Pathway & PSY Knockdown Impact
The Scientist's Toolkit: Research Reagent Solutions
Table 7: Essential Reagents and Materials for RNAi Validation in Metabolic Pathways
| Item | Function & Application | Example/Notes |
|---|---|---|
| RNAi Constructs | Induces sequence-specific mRNA degradation. | siRNA, shRNA, or dsRNA targeting PSY1, LCY-e, etc. Critical to include scramble/barcode controls. |
| High-Quality RNA Kit | Isolates intact, pure total RNA for qPCR. | Kits with DNase I treatment step (e.g., RNeasy, TRIzol). Essential for avoiding genomic DNA contamination. |
| Reverse Transcription Kit | Synthesizes stable cDNA from RNA template. | Use kits with both random hexamers and oligo-dT for comprehensive coverage. |
| SYBR Green qPCR Master Mix | Enables real-time, fluorescent detection of amplified DNA. | Choose mixes with ROX passive reference dye and high efficiency. |
| Validated Primers | Specifically amplify target and reference gene sequences. | PSY1, LCY-b, ZDS; must be efficiency-tested. Reference genes: ACTB, UBQ10. |
| Protein Lysis Buffer (RIPA) | Extracts total protein while maintaining integrity. | Must include fresh protease/phosphatase inhibitors. |
| Carotenoid Pathway Antibodies | Detects specific enzyme proteins via Western blot. | Commercial (less common) or custom-generated (e.g., anti-PSY, anti-LCY). Validate specificity. |
| HRP-Linked Secondary Antibodies | Amplifies primary antibody signal for detection. | Anti-rabbit or anti-mouse, depending on primary host species. |
| Enhanced Chemiluminescence (ECL) Substrate | Generates light signal for HRP enzyme on Western blots. | Choose ultrasensitive or linear range substrates based on target abundance. |
| Carotenoid Analytical Standards | Enables absolute quantification in LC-MS/MS. | Phytoene, lycopene, β-carotene, lutein, zeaxanthin from reputable suppliers. |
| Deuterated Internal Standard | Corrects for variability in extraction and ionization in MS. | e.g., β-carotene-d6. Crucial for assay precision and accuracy. |
| C30 HPLC Column | Chromatographically separates geometric and structural carotenoid isomers. | Superior to C18 for resolving lutein/zeaxanthin, alpha/beta-carotene. |
| APCI or ESI Ion Source | Ionizes non-polar (APCI) or polar (ESI) metabolites for MS detection. | APCI+ is often the standard for carotenoids due to efficient ionization. |
This guide details phenotypic validation assays essential for research investigating the suppression of the carotenoid biosynthesis pathway via RNA interference (RNAi). Within the broader thesis, these assays serve as critical functional readouts downstream of metabolic perturbation. Carotenoids, precursors to retinoids and apocarotenoids, influence fundamental cellular processes including proliferation, survival, and oxidative stress response. RNAi-mediated knockdown of key enzymes (e.g., PSY, BCO1, BCO2) disrupts this pathway, leading to reduced carotenoid/retinoid pools. This deficit can alter retinoid signaling, impact mitochondrial function, and increase susceptibility to oxidative damage, ultimately manifesting in measurable phenotypic changes in cell growth, death, and clonogenic potential. This document provides a technical framework to rigorously quantify these phenotypic endpoints.
Core Phenotypic Assays: Methodologies & Data Interpretation
Cell Proliferation Assay (MTS/Trypan Blue)
Protocol: Seed cells (e.g., HepG2, Caco-2, retinal pigment epithelium) in 96-well plates (3,000-5,000 cells/well). 24h post-seeding, transfer to carotenoid-depleted serum and transfect with siRNA targeting your gene of interest (GOI) vs. non-targeting control (NTC). Include untransfected controls.
- Day 0-5 Measurement: At each 24h interval, for MTS, add 20 µL of MTS reagent directly to culture medium, incubate for 1-4h at 37°C, and measure absorbance at 490 nm. For Trypan Blue, harvest cells from parallel plates, mix with 0.4% Trypan Blue (1:1), and count viable (unstained) cells using a hemocytometer or automated counter.
Data Interpretation: Normalize absorbance or cell counts to Day 0 or the NTC at each time point. A significant reduction in the slope of the growth curve indicates anti-proliferative effects due to carotenoid pathway suppression.
Table 1: Sample Proliferation Data (MTS, 72h Post-transfection)
| siRNA Target (Carotenoid Pathway Gene) | Cell Line | Mean Absorbance (490nm) ± SD | % Proliferation vs. NTC | p-value |
|---|---|---|---|---|
| Non-Targeting Control (NTC) | HepG2 | 1.25 ± 0.09 | 100.0% | - |
| PSY1 | HepG2 | 0.81 ± 0.07 | 64.8% | <0.001 |
| BCO1 | HepG2 | 0.92 ± 0.08 | 73.6% | 0.003 |
| BCO2 | HepG2 | 1.18 ± 0.10 | 94.4% | 0.12 |
Apoptosis Assay (Annexin V / PI Flow Cytometry)
Protocol: 72h post-transfection, harvest both adherent and floating cells. Wash twice with cold PBS. Resuspend 1x10^5 cells in 100 µL of 1X Annexin V Binding Buffer. Add 5 µL of FITC-conjugated Annexin V and 1 µL of Propidium Iodide (PI, 100 µg/mL) working solution. Incubate for 15 min at RT in the dark. Add 400 µL of binding buffer and analyze by flow cytometry within 1h. Use unstained and single-stained controls for compensation.
Data Interpretation: Cells are categorized as viable (Annexin V-/PI-), early apoptotic (Annexin V+/PI-), late apoptotic (Annexin V+/PI+), or necrotic (Annexin V-/PI+). An increase in the combined early and late apoptotic populations signifies induction of programmed cell death.
Table 2: Sample Apoptosis Data (Flow Cytometry, 72h Post-transfection)
| Cell Population | NTC siRNA (% of total) | PSY1 siRNA (% of total) | BCO1 siRNA (% of total) |
|---|---|---|---|
| Viable (Annexin V-/PI-) | 89.5 ± 3.2 | 68.4 ± 4.1 | 77.8 ± 3.5 |
| Early Apoptotic (Annexin V+/PI-) | 4.1 ± 1.2 | 18.7 ± 2.8 | 12.3 ± 2.1 |
| Late Apoptotic (Annexin V+/PI+) | 3.8 ± 1.0 | 9.5 ± 1.9 | 7.2 ± 1.4 |
| Necrotic (Annexin V-/PI+) | 2.6 ± 0.8 | 3.4 ± 1.1 | 2.7 ± 0.9 |
Colony Formation Assay (Clonogenic Survival)
Protocol: 24h post-transfection, trypsinize and seed a low density of cells (200-1000, depending on line) into 6-well plates. Allow colonies to grow for 10-14 days, replacing medium every 3-4 days. Aspirate medium, wash with PBS, fix with 4% paraformaldehyde or methanol for 15 min, and stain with 0.5% crystal violet (in 25% methanol) for 30 min. Gently rinse with water and air-dry. Image plates and count colonies (>50 cells). Use imaging software or manual counting.
Data Interpretation: The plating efficiency (PE = colonies counted / cells seeded x 100) and surviving fraction (SF = PE(treated) / PE(control)) quantify long-term reproductive viability. A reduced SF indicates impaired clonogenic capacity following pathway suppression.
Table 3: Sample Colony Formation Data (14-day assay)
| siRNA Target | Cells Seeded | Colonies Counted (Mean ± SD) | Plating Efficiency (%) | Surviving Fraction |
|---|---|---|---|---|
| NTC | 500 | 183 ± 12 | 36.6% | 1.00 |
| PSY1 | 500 | 47 ± 8 | 9.4% | 0.26 |
| BCO1 | 500 | 89 ± 10 | 17.8% | 0.49 |
Signaling Pathways & Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for Phenotypic Validation Assays
| Reagent / Kit / Material | Primary Function & Application in This Context |
|---|---|
| Validated siRNAs (e.g., ON-TARGETplus) | Pooled, SMARTvector siRNAs ensure specific, reproducible knockdown of carotenoid pathway genes with minimal off-target effects. |
| Carotenoid/Retinoid-Depleted FBS | Essential for in vitro studies to remove exogenous sources of carotenoids and retinoids, ensuring observed effects are due to endogenous pathway disruption. |
| MTS Cell Proliferation Assay Kit | Colorimetric assay to quantify metabolically active cells; indicates changes in proliferation rate post-knockdown. |
| Annexin V-FITC / PI Apoptosis Kit | Fluorescence-based flow cytometry kit to distinguish and quantify apoptotic vs. necrotic cell populations. |
| Crystal Violet Staining Solution | Dye used to visualize and count fixed colonies in clonogenic survival assays. |
| Matrigel / Basement Membrane Matrix | May be required for colony formation assays of sensitive or primary cell lines to improve plating efficiency. |
| Flow Cytometer with 488nm laser | Instrument required for high-throughput, quantitative analysis of Annexin V/PI-stained apoptotic samples. |
| Automated Cell Counter (e.g., Countess) | Provides rapid, consistent viable cell counts for proliferation (Trypan Blue) and seeding standardization for colony assays. |
| Retinoic Acid Receptor (RAR/RXR) Agonists/Antagonists | Pharmacologic tools to rescue or mimic signaling deficits, linking phenotype directly to altered retinoid signaling. |
| ROS Detection Probe (e.g., DCFDA) | Fluorescent dye to measure reactive oxygen species, connecting carotenoid depletion to oxidative stress-mediated apoptosis. |
Within the broader thesis on modulating the carotenoid biosynthesis pathway, this whitepaper provides a technical comparison of RNA interference (RNAi) and small molecule inhibitor (SMI) strategies. We assess their mechanisms, efficacy, specificity, and applicability in research and therapeutic development, highlighting potential synergies for enhanced pathway suppression.
Carotenoids are tetraterpenoid pigments synthesized in plants, algae, and some bacteria and fungi, with roles in photosynthesis, photoprotection, and as vitamin A precursors. The pathway involves key enzymes such as Phytoene Synthase (PSY), Phytoene Desaturase (PDS), ζ-Carotene Desaturase (ZDS), and Lycopene β-Cyclase (LCYb). Inhibition of these enzymes is a target for agricultural (e.g., herbicide development, crop modification) and biomedical (e.g., treating retinopathies, metabolic disorders) applications. Two primary inhibitory modalities are explored: RNAi-mediated gene silencing and SMI-based enzyme inhibition.
Mechanisms of Action
Small Molecule Inhibitors (SMIs)
SMIs are low-molecular-weight compounds that directly bind to the active site or an allosteric site of a target enzyme, disrupting its catalytic function. For carotenogenic enzymes, common inhibitors include:
- Norflurazon: Inhibits PDS.
- Diflufenican: Inhibits PDS.
- CPTA (2-(4-Chlorophenylthio)triethylamine): Inhibits Lycopene Cyclase.
Diagram Title: Mechanism of Small Molecule Enzyme Inhibition
RNA Interference (RNAi)
RNAi is a biological process where double-stranded RNA (dsRNA) triggers the sequence-specific degradation of complementary messenger RNA (mRNA), preventing translation. For carotenoid enzymes, dsRNA targeting PSY, PDS, or LCYb mRNA can be delivered.
Diagram Title: RNAi Mechanism for Gene Silencing
Quantitative Comparison: Pros and Cons
Table 1: Head-to-Head Comparison of Key Attributes
| Attribute | Small Molecule Inhibitors (SMIs) | RNA Interference (RNAi) |
|---|---|---|
| Target Specificity | Moderate to Low. Binds based on 3D structure; off-target effects on related enzymes possible. | Very High. Sequence-based targeting; careful design minimizes off-targets. |
| Development Timeline | Longer (5-7+ years). Requires high-throughput screening, SAR optimization, ADMET profiling. | Shorter (1-2 years for design). Rapid in silico design of siRNA/hpRNA. |
| Delivery Challenge | Low. Typically cell-permeable; systemic via spray or root uptake in plants. | High. Requires vector (viral/plasmid) or formulation (nanoparticle, topical) for intracellular delivery. |
| Duration of Effect | Transient (hours-days). Dependent on inhibitor stability and turnover. | Prolonged (days-weeks). Sustained silencing until siRNA degrades or cell divides. |
| Resistance Potential | Higher. Single amino acid change in target site can confer resistance. | Lower. Requires mutation in the short siRNA target sequence; multiplexing possible. |
| Cost of Goods | Low to Moderate. Chemical synthesis at scale is established. | High. Large-scale synthesis of dsRNA/siRNA or transgenic production is costly. |
| Applicability | Broad; can target non-enzymatic proteins or multiple enzymes with one inhibitor. | Limited to proteins whose function can be modulated via mRNA reduction. |
| Environmental Impact | Potentially higher; chemical persistence and metabolite toxicity are concerns. | Potentially lower; dsRNA is biodegradable, but ecological impact of gene silencing is under study. |
Table 2: Experimental Efficacy Data from Recent Studies (2020-2023)
| Inhibitor Type | Target Enzyme (Organism) | Reported Efficacy (IC50/KI or % Knockdown) | Key Measurement Method |
|---|---|---|---|
| SMI: Norflurazon | Phytoene Desaturase (Arabidopsis) | IC50 ~ 50 nM (in vitro enzyme assay) | Spectrophotometric measurement of ζ-carotene accumulation. |
| SMI: CPTA | Lycopene β-Cyclase (Tomato) | Ki ~ 1.2 µM | HPLC quantification of lycopene (accumulates) vs. β-carotene (decreases). |
| RNAi (hpRNA) | Phytoene Synthase (Maize) | >85% mRNA knockdown | qRT-PCR of PSY transcript levels in endosperm. |
| RNAi (siRNA) | LCYb (Human HeLa cells) | 70-80% protein knockdown | Western Blot analysis of LCYb protein levels. |
Detailed Experimental Protocols
Protocol: High-Throughput Screening for SMI Lead Identification
Objective: Identify novel SMIs against recombinant Phytoene Desaturase (PDS).
- Enzyme Preparation: Express and purify recombinant PDS with a His-tag.
- Assay Setup: In a 96-well plate, combine purified PDS (10 nM), substrate phytoene (50 µM in detergent micelles), co-factors (FAD, NADP+), and test compound library (10 µM each). Include norflurazon (positive control) and DMSO (negative control).
- Reaction & Detection: Incubate at 25°C for 30 min. Stop reaction with methanol. Quantify product ζ-carotene by measuring absorbance at 400 nm or via HPLC-PDA.
- Data Analysis: Calculate % inhibition relative to DMSO control. Compounds with >70% inhibition proceed to dose-response for IC50 determination.
Protocol: RNAi-Mediated Knockdown in Plant Tissue
Objective: Silence PSY gene expression in Nicotiana benthamiana leaves.
- dsRNA Design & Synthesis: Design a 200-300 bp hpRNA construct targeting a conserved region of PSY mRNA. Clone into an appropriate RNAi vector (e.g., pHELLSGATE).
- Agroinfiltration: Transform construct into Agrobacterium tumefaciens strain GV3101. Grow culture to OD600=0.5, resuspend in infiltration buffer (10 mM MES, 10 mM MgCl2, 150 µM acetosyringone). Inject suspension into the abaxial side of young leaves.
- Sampling & Validation: Harvest leaf discs 4-7 days post-infiltration.
- qRT-PCR: Extract total RNA, perform cDNA synthesis, and run qPCR with PSY-specific primers. Normalize to a housekeeping gene (e.g., EF1α).
- Phenotypic Analysis: Visually assess photobleaching (white/yellow leaves). Extract pigments with acetone:hexane and analyze via HPLC for phytoene accumulation and carotenoid depletion.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Carotenoid Inhibition Studies
| Reagent/Material | Function & Role in Research | Example Product/Catalog |
|---|---|---|
| Recombinant Carotenoid Enzymes | Purified protein for in vitro SMI screening and biochemical characterization. | Arabidopsis PDS, expressed in E. coli (Agrisera, AS152865). |
| Chemical Inhibitor Libraries | Collections of diverse SMIs for high-throughput screening against target enzymes. | MCE Bioactive Compound Library (MedChemExpress, HY-L001). |
| Custom siRNA/dsRNA Synthesis Service | Provides high-purity, sequence-verified RNAi triggers for in vitro and in vivo studies. | Dharmacon AccuGrade Custom siRNA (Horizon Discovery). |
| Gateway-Compatible RNAi Vectors | Cloning systems for easy construction of hpRNA expression cassettes for plant transformation. | pHELLSGATE or pANDA vectors (available from Addgene). |
| HPLC-PDA System with C30 Column | Gold-standard method for separating, identifying, and quantifying individual carotenoid species. | YMC Carotenoid C30 column, 3 µm, 150 x 4.6 mm. |
| Carotenoid Reference Standards | Pure compounds for HPLC calibration, quantification, and identification. | Phytoene, Lycopene, β-Carotene (CaroteNature, Extrasynthese). |
| In Vivo Imaging Systems (for plants) | Quantify photobleaching phenotypes non-destructively via chlorophyll fluorescence imaging. | PhenoVation CropReporter or similar. |
Synergistic Applications and Future Outlook
The future of carotenoid pathway suppression lies in strategic synergy:
- Combination Therapy: Using a low-dose SMI to partially inhibit an enzyme, combined with RNAi to suppress its synthesis, could lead to stronger, more durable suppression with reduced risk of resistance.
- RNAi for Target Discovery: RNAi knockdown can validate an enzyme's essentiality in vivo before investing in costly SMI screening campaigns.
- SMIs as Delivery Enhancers: Certain SMIs that disrupt cellular membranes could be co-formulated to improve the cellular uptake of RNAi triggers.
Diagram Title: Strategic Synergy Between RNAi and SMI Approaches
Both RNAi and SMIs offer distinct advantages for inhibiting carotenoid enzymes, as outlined in this technical guide. SMIs provide immediate, broad-acting tools, while RNAi offers unparalleled specificity and durable effects. The integration of both approaches—using RNAi for target validation and sustained suppression, and SMIs for rapid phenotypic screening and combinatorial inhibition—represents the most powerful strategy within the thesis of carotenoid biosynthesis pathway research, paving the way for next-generation agricultural and therapeutic applications.
This analysis is framed within the context of a broader thesis investigating the suppression of the carotenoid biosynthesis pathway. Choosing the appropriate functional genomics tool—RNA interference (RNAi) for transient knockdown or CRISPR-Cas9 for permanent knockout—is critical for accurate pathway dissection, target validation, and understanding feedback mechanisms.
Mechanism of Action
RNAi Knockdown: Utilizes the cell's endogenous RNA-induced silencing complex (RISC). Introduced double-stranded RNA (dsRNA) or short hairpin RNA (shRNA) is processed into ~21-23 nucleotide siRNAs. The siRNA guide strand binds RISC, which then identifies and cleaves complementary mRNA sequences, leading to transcript degradation and reduced protein levels without altering the genomic DNA.
CRISPR-Cas9 Knockout: Employs a engineered single-guide RNA (sgRNA) to direct the Cas9 endonuclease to a specific genomic DNA sequence. Cas9 creates a double-strand break (DSB), which is repaired by error-prone non-homologous end joining (NHEJ). This often results in small insertions or deletions (indels) that disrupt the reading frame, leading to a permanent loss of functional protein.
Table 1: Core Technical Comparison
| Parameter | RNAi Knockdown | CRISPR-Cas9 Knockout |
|---|---|---|
| Molecular Target | mRNA in the cytoplasm | DNA in the nucleus |
| Primary Effect | Reduces transcript levels | Disrupts the gene locus |
| Typical Efficiency | 70-90% protein reduction (subject to variability) | Near 100% biallelic disruption possible |
| Permanence | Transient (days to weeks) | Stable and heritable |
| Off-Target Effects | Common; seed-sequence mediated miRNA-like effects | Less frequent; DNA cleavage at near-cognate sites |
| Key Advantage | Tunable, reversible suppression; studies essential genes | Complete loss-of-function; studies gene necessity |
| Key Limitation | Incomplete knockdown; potential for false negatives | May trigger compensatory adaptation; not suitable for essential genes in proliferative contexts |
Table 2: Suitability for Carotenoid Pathway Analysis
| Research Goal | Recommended Tool | Rationale |
|---|---|---|
| Rapid Screening of Multiple Pathway Enzymes | RNAi (siRNA/shRNA libraries) | Faster, lower-cost transient screening. |
| Validate Essentiality of a Specific Biosynthetic Gene | CRISPR-Cas9 Knockout | Confirms if the gene is absolutely required for pathway output. |
| Study Feedback Regulation & Transcriptional Adaptation | CRISPR-Cas9 Knockout | Distinguishes between direct and compensatory effects. |
| Analyze Dose-Dependent Effects of Enzyme Reduction | RNAi (with titration) | Allows for graded reduction of protein levels. |
| Generate Stable Cell Lines with Pathway Blockade | CRISPR-Cas9 Knockout | Creates consistent, isogenic models for downstream assays. |
Detailed Experimental Protocols
Protocol 1: RNAi Knockdown of Phytoene Synthase (PSY1) in a Cell Model Objective: Achieve transient knockdown of PSY1 to assess its role in the carotenoid flux.
- Design: Use validated siRNA sequences targeting human PSY1 (e.g., from public database like Dharmacon). Include non-targeting siRNA (scrambled) and mock (transfection reagent only) controls.
- Reverse Transfection: Seed cells in a 24-well plate at 70% confluence. For each well, dilute 5 nM siRNA in 50 µL of serum-free medium. In a separate tube, dilute 1.5 µL of lipid-based transfection reagent (e.g., Lipofectamine RNAiMAX) in 50 µL of serum-free medium. Combine dilutions, incubate 5-20 min at RT to form complexes. Add mixture dropwise to cells. Top up with complete medium.
- Harvest: Incubate cells for 48-72 hours.
- Validation:
- qRT-PCR: Isolate total RNA, synthesize cDNA. Perform qPCR with PSY1-specific primers. Normalize to housekeeping gene (e.g., GAPDH). Calculate knockdown efficiency via ∆∆Ct method.
- Functional Assay: Extract metabolites. Quantify downstream carotenoids (e.g., β-carotene) via HPLC to measure pathway flux reduction.
- Western Blot (Optional): If a reliable antibody exists, confirm protein level reduction.
Protocol 2: CRISPR-Cas9 Knockout of LCYE in a Plant System Objective: Generate stable LCYE (Lycopene ε-cyclase) knockout lines to study branch-point commitment in carotenogenesis.
- sgRNA Design & Cloning: Design two sgRNAs targeting early exons of the LCYE gene using a design tool (e.g., CHOPCHOP). Clone sgRNA sequences into a plant-specific CRISPR-Cas9 vector (e.g., pHEE401E) harboring Cas9 and a selection marker.
- Plant Transformation: Transform the construct into Arabidopsis thaliana via floral dip method using Agrobacterium tumefaciens.
- Selection & Genotyping: Select T1 seeds on appropriate antibiotic. Harvest leaf tissue from resistant seedlings.
- PCR Amplification: Amplify the genomic region flanking the target site.
- T7 Endonuclease I Assay: Denature and reanneal PCR products. Digest heteroduplex DNA formed from indels with T7E1. Analyze fragments on agarose gel to identify mutagenized events.
- Sequencing: Sanger sequence PCR products from putative mutants. Analyze chromatograms for overlapping peaks downstream of the cut site, then clone amplicons and sequence individual alleles to characterize specific indels.
- Homozygous Line Generation: Self-pollinate T1 plants with confirmed mutations. Screen T2 progeny by sequencing to identify homozygous knockout lines.
- Phenotypic Analysis: Quantify carotenoid profile in leaves/flowers of homozygous vs. wild-type plants using HPLC-MS/MS. Expect a shift from lutein (ε-branch) towards β-branch carotenoids.
Visualizations
Title: RNAi Knockdown Workflow for Carotenoid Genes
Title: CRISPR-Cas9 Knockout Workflow for Carotenoid Genes
Title: Carotenoid Biosynthesis Pathway with Key Targets
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Functional Analysis of the Carotenoid Pathway
| Reagent / Material | Function & Application | Example Vendor/Product |
|---|---|---|
| Validated siRNA Libraries | Pre-designed, high-confidence siRNAs for rapid screening of carotenoid pathway genes. | Horizon Discovery (Dharmacon), Sigma-Aldrich |
| CRISPR-Cas9 Knockout Kit | All-in-one plasmids (Cas9 + sgRNA scaffold) for easy cloning and knockout generation. | Addgene (e.g., pSpCas9(BB)), Synthego |
| Lipofectamine RNAiMAX | Lipid-based transfection reagent optimized for high-efficiency siRNA delivery with low cytotoxicity. | Thermo Fisher Scientific |
| T7 Endonuclease I | Enzyme for detecting indel mutations in CRISPR-edited populations via mismatch cleavage assay. | New England Biolabs |
| Carotenoid Standard Mix | Authentic chemical standards for HPLC or LC-MS calibration and identification of pathway metabolites. | CaroteNature, Sigma-Aldrich |
| RNeasy Kit / TRIzol | For high-quality total RNA isolation, essential for validating transcript knockdown (qRT-PCR). | Qiagen, Thermo Fisher Scientific |
| C18 Reverse-Phase HPLC Column | Specialized chromatography column for separating non-polar carotenoids (e.g., lycopene, β-carotene). | Agilent, Waters |
| Anti-Cas9 Antibody | For confirming Cas9 protein expression in transfected/transformed cells or tissues. | Cell Signaling Technology, Abcam |
This article provides a technical review of published preclinical case studies where RNA interference (RNAi) has been successfully employed to suppress target genes, leading to therapeutic efficacy in cancer models. This analysis is framed within a broader research thesis investigating the modulation of metabolic pathways, including the carotenoid biosynthesis pathway, as a novel oncotherapeutic strategy. RNAi-mediated suppression of key enzymes in such pathways offers a precise tool for validating novel drug targets and understanding cancer-specific metabolic dependencies.
Core Case Studies in Preclinical Oncology
Case Study 1: Suppression of PLK1 in Non-Small Cell Lung Cancer (NSCLC)
- Target Gene: Polo-like kinase 1 (PLK1), a critical regulator of cell cycle progression.
- Cancer Model: In vivo subcutaneous xenograft model using A549 human NSCLC cells in immunodeficient mice.
- RNAi Delivery System: Cholesterol-conjugated siRNA (cc-siRNA) for systemic delivery.
- Key Findings: Sustained PLK1 suppression resulted in tumor cell apoptosis, mitotic arrest, and significant inhibition of tumor growth compared to scramble siRNA controls.
- Relevance to Thesis: Demonstrates the efficacy of systemic RNAi against a classic cell cycle target, a principle applicable to targeting metabolic pathway enzymes like those in carotenoid synthesis.
Case Study 2: Knockdown of IDO1 in Melanoma and Modulation of Immune Response
- Target Gene: Indoleamine 2,3-dioxygenase 1 (IDO1), an immunomodulatory enzyme that suppresses T-cell function.
- Cancer Model: B16-F10 syngeneic melanoma model in immunocompetent C57BL/6 mice.
- RNAi Delivery System: Cationic lipid nanoparticles (LNPs) encapsulating siRNA.
- Key Findings: IDO1 knockdown in the tumor microenvironment reversed local immune suppression, increased CD8+ T-cell infiltration, and synergized with anti-PD-1 checkpoint therapy to regress tumors.
- Relevance to Thesis: Highlights RNAi's potential to modulate tumor microenvironment metabolism, analogous to proposed studies on carotenoid pathway metabolites which may also influence immune cell activity.
Case Study 3: Co-suppression of BCL2 and MCL1 in Acute Myeloid Leukemia (AML)
- Target Genes: Anti-apoptotic proteins B-cell lymphoma 2 (BCL2) and Myeloid cell leukemia 1 (MCL1).
- Cancer Model: Systemic xenograft model using MV4-11 human AML cells in NSG mice.
- RNAi Delivery System: GalNAc-conjugated siRNA for targeted hepatocyte expression (as a control) and novel dendrimer-based delivery for leukemic cells.
- Key Findings: Dual suppression of BCL2 and MCL1 overcame compensatory survival mechanisms, inducing massive apoptosis in leukemic blasts and significantly prolonging animal survival.
- Relevance to Thesis: Showcases combinatorial RNAi strategy to overcome redundancy, a tactic that may be required when targeting interconnected metabolic networks like carotenoid biosynthesis.
Table 1: Summary of Quantitative Outcomes from Preclinical RNAi Case Studies
| Case Study | Target Gene(s) | Cancer Model | Delivery Method | Key Efficacy Metric | Result (Mean ± SD) | Control Result |
|---|---|---|---|---|---|---|
| PLK1 in NSCLC | PLK1 | A549 Xenograft | cc-siRNA | Tumor Volume (Day 21) | 245 ± 45 mm³ | 850 ± 120 mm³ |
| PLK1 mRNA (Rel. Expr.) | 0.15 ± 0.05 | 1.0 ± 0.1 | ||||
| IDO1 in Melanoma | IDO1 | B16-F10 Syngeneic | LNP-siRNA | Tumor Volume (Day 18) | 65 ± 22 mm³ | 320 ± 55 mm³ |
| Intratumoral CD8+ T-cells | 35 ± 5% | 12 ± 3% | ||||
| BCL2/MCL1 in AML | BCL2, MCL1 | MV4-11 Xenograft | Dendrimer-siRNA | Median Survival | 58.5 days | 38 days |
| Apoptosis (Caspase 3/7+) | 68 ± 7% | 8 ± 2% |
Detailed Experimental Protocols
Protocol A: In Vivo Efficacy Study with Systemically Delivered siRNA (Case Study 1)
- Model Establishment: Inject 5 x 10^6 A549 cells subcutaneously into the flank of female athymic nude mice.
- Randomization: When tumors reach ~100 mm³, randomize mice into treatment groups (n=8).
- Formulation: Reconstitute cholesterol-conjugated PLK1 siRNA or scramble control in sterile PBS.
- Dosing: Administer siRNA intravenously (3 mg/kg) via tail vein every 3 days for a total of 5 doses.
- Monitoring: Measure tumor dimensions with calipers bi-weekly. Calculate volume as (Length x Width²)/2.
- Terminal Analysis: At Day 21, euthanize animals. Excise tumors, weigh, and process for qRT-PCR (for PLK1 mRNA) and immunohistochemistry (for cleaved caspase-3).
Protocol B: Intratumoral Immune Profiling Post-RNAi (Case Study 2)
- Treatment: Treat tumor-bearing mice with IDO1-LNP or control-LNP (2 doses, i.v., days 7 & 10 post-implant).
- Harvest: On day 14, excise tumors and create a single-cell suspension using a mechanical dissociator and collagenase/DNase digest.
- Staining: Stain cells with fluorescent antibodies against surface markers (CD45, CD3, CD8) and viability dye.
- Flow Cytometry: Acquire data on a flow cytometer. Gate on live, CD45+ leukocytes, then on CD3+CD8+ T-cells to determine percentage of tumor-infiltrating lymphocytes (TILs).
- qRT-PCR: Analyze snap-frozen tumor tissue for IDO1 mRNA and related cytokines (IFN-γ, TGF-β).
Visualization of Key Concepts
Diagram 1: RNAi Mechanism and Therapeutic Application Pathway
Diagram 2: Preclinical In Vivo RNAi Workflow
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for Preclinical RNAi Cancer Studies
| Reagent Category | Specific Example(s) | Function in Experiment |
|---|---|---|
| siRNA Oligos | ON-TARGETplus SMARTpool (Dharmacon); Silencer Select (Ambion) | High-purity, chemically modified siRNA duplexes for specific, potent target knockdown with reduced off-target effects. |
| In Vivo Transfection Reagent | Invivofectamine 3.0 (Lipid-based); Polyplus in vivo-jetPEI | Formulation reagents designed to complex with and protect siRNA, enabling efficient cellular uptake and endosomal escape in vivo. |
| Delivery Vehicle | Cholesterol Conjugation; GalNAc Conjugation; Lipid Nanoparticles (LNPs) | Chemical modifications or nanoparticles that confer stability in circulation and promote targeted delivery to specific tissues (e.g., liver) or tumors. |
| Validated Antibodies | Phospho-Histone H3 (Ser10) (Cell Signaling); Cleaved Caspase-3 (Asp175) | Essential for immunohistochemistry or western blot to validate on-target biological effects (mitotic arrest, apoptosis). |
| qRT-PCR Kits | TaqMan RNA-to-Ct 1-Step Kit; SYBR Green Master Mix | For quantitative assessment of target mRNA knockdown efficiency in tumor tissues post-treatment. |
| Preclinical Imaging System | IVIS Spectrum (PerkinElmer); MRI/Micro-CT | Enables non-invasive, longitudinal monitoring of tumor burden and metabolic changes in live animal models. |
Conclusion
The strategic suppression of the carotenoid biosynthesis pathway via RNAi presents a powerful and precise approach to dissecting cancer metabolism and identifying novel therapeutic vulnerabilities. This synthesis underscores that successful application hinges on a deep understanding of pathway biology, meticulous RNAi design to avoid off-target effects, and optimization of delivery systems—especially for in vivo translation. Compared to pharmacological inhibition, RNAi offers unparalleled gene-specificity, while complementing CRISPR screens for functional genomics. Future directions should focus on developing targeted delivery mechanisms (e.g., tumor-specific LNPs) to enable systemic therapy, exploring combination strategies with existing oncology drugs, and further elucidating the role of specific carotenoid metabolites in tumor immune evasion. For drug development professionals, this pathway represents an emerging, metabolism-focused frontier for next-generation oncology targets.