Optimized Agrobacterium rhizogenes Root Transformation Protocol: A Step-by-Step Guide for Drug Discovery Researchers
This comprehensive protocol details an optimized method for Agrobacterium rhizogenes-mediated root transformation, a powerful technique for producing recombinant proteins and bioactive compounds in hairy root cultures.
Optimized Agrobacterium rhizogenes Root Transformation Protocol: A Step-by-Step Guide for Drug Discovery Researchers
Abstract
This comprehensive protocol details an optimized method for Agrobacterium rhizogenes-mediated root transformation, a powerful technique for producing recombinant proteins and bioactive compounds in hairy root cultures. Targeting researchers and drug development professionals, the article covers foundational principles, a detailed step-by-step methodology, troubleshooting for common issues, and validation strategies to ensure reliable, high-yield results for biomedical applications.
Understanding Agrobacterium rhizogenes: The Biology Behind Hairy Root Disease and Biotech Applications
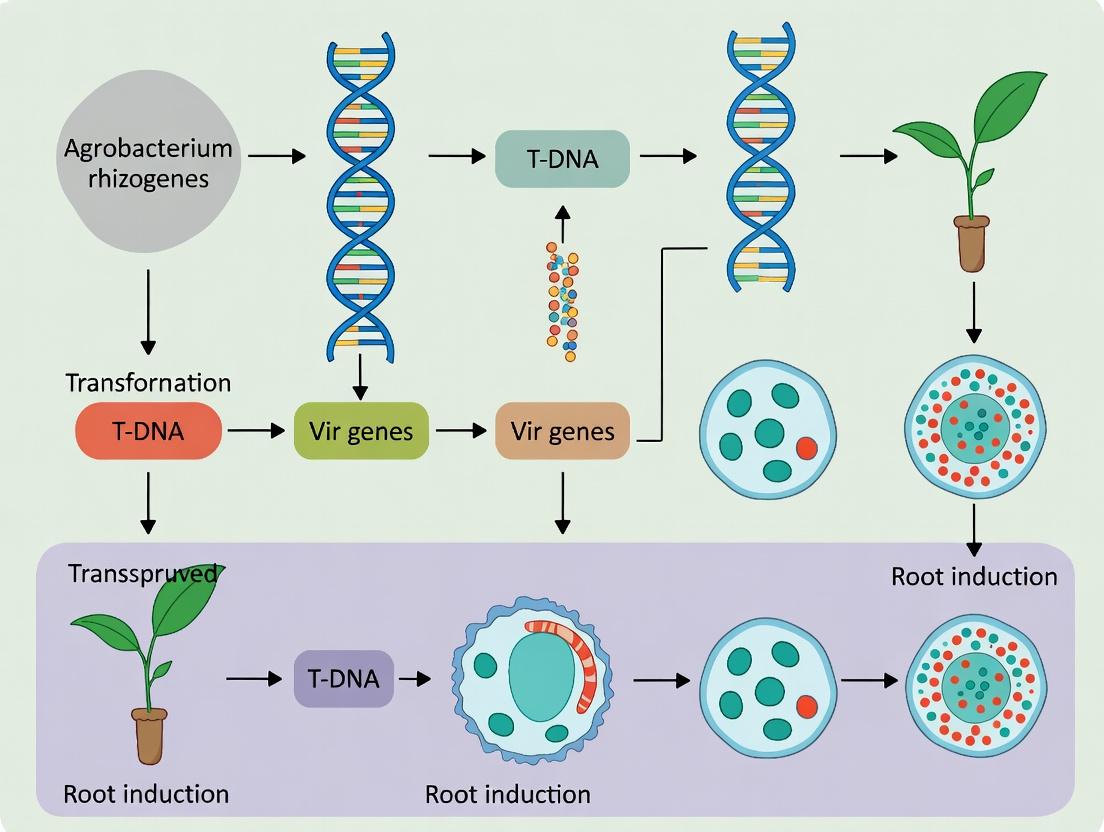
Agrobacterium rhizogenes is a gram-negative soil bacterium and a natural genetic engineer. It causes "hairy root" disease in dicotyledonous plants by transferring a segment of DNA (T-DNA) from its Root-inducing (Ri) plasmid into the plant genome. The integration and expression of T-DNA genes lead to the proliferation of adventitious roots at the infection site. These "hairy roots" are characterized by rapid growth, high lateral branching, and genetic stability.
Core Mechanism: The Ri Plasmid and Virulence System The key genetic components reside on the Ri plasmid, which shares functional homology with the Ti (Tumor-inducing) plasmid of A. tumefaciens.
- T-DNA (Transferred DNA): Defined by left and right border sequences (LB/RB). The core T-DNA contains root oncogenic loci (rol) genes (rolA, rolB, rolC, rolD), which are responsible for auxin and cytokinin perturbation leading to root initiation and growth.
- Virulence (Vir) Region: A set of operons (virA, virB, virC, virD, virE, virG, virH) that are activated by plant-derived phenolic compounds (e.g., acetosyringone) and sugars. The proteins process and transfer the T-DNA into the plant cell.
- Origin of Replication (Ori): For plasmid maintenance in the bacterium.
- Opine Catabolism Genes: Allow the bacterium to utilize opines, unique amino acid derivatives synthesized by the transformed plant, as a nutrient source.
Signaling Pathway for T-DNA Transfer
Title: A. rhizogenes Virulence Induction and T-DNA Transfer
Key Applications in Research and Industry
Hairy root cultures derived from A. rhizogenes transformation are versatile platforms.
Table 1: Primary Applications of Hairy Root Cultures
| Application Area | Specific Use | Key Advantage |
|---|---|---|
| Secondary Metabolite Production | Sustainable synthesis of plant-derived pharmaceuticals (alkaloids, terpenoids, phenolics), flavors, and dyes. | High biochemical stability, growth in hormone-free media, often higher yields than native roots or cell suspensions. |
| Phytoremediation | Uptake and degradation of environmental contaminants (heavy metals, organic pollutants). | Extensive root surface area and rapid biomass production enhance remediation efficiency. |
| Functional Genomics | Study root biology, gene function (via RNAi/CRISPR in roots), and plant-pathogen interactions. | Provides a genetically uniform, easily propagated root system for experimentation. |
| Protein Molecular Farming | Production of recombinant proteins, enzymes, and antibodies. | Secretory nature of roots can simplify protein recovery. |
| Metabolic Engineering | Pathway elucidation and manipulation to enhance production of target compounds. | Roots are natural sites for synthesis of many metabolites; amenable to genetic stacking. |
Detailed Protocol: Hairy Root Induction and Culture
This protocol outlines the generation of hairy roots from explants for metabolite production studies.
Materials & Pre-culture
- Bacterial Strain: A. rhizogenes (e.g., wild-type A4 or engineered R1000 with disarmed pRi).
- Plant Material: Surface-sterilized leaf discs or stem segments from target species.
- Media: YEB (for Agrobacterium), MS or B5 basal media (solid and liquid, no hormones).
- Antibiotics: Appropriate for bacterial selection (e.g., rifampicin, kanamycin) and later for root culture (cefotaxime to eliminate bacteria).
Procedure
Title: Workflow for Establishing Hairy Root Cultures
Detailed Steps:
- Bacterial Preparation: Grow A. rhizogenes in YEB broth with appropriate antibiotics (28°C, 200 rpm) to mid-log phase. Centrifuge and resuspend in fresh liquid MS medium to ~10⁸ CFU/mL.
- Explant Inoculation: Dip sterilized explants in the bacterial suspension for 5-10 minutes. Blot dry on sterile paper.
- Co-cultivation: Place explants on solid MS medium (no antibiotics). Seal plates and incubate in the dark at 23-25°C for 2-3 days.
- Decontamination & Induction: Transfer explants to solid MS medium supplemented with cefotaxime (250-500 mg/L) to eliminate bacteria. Maintain in low light at 25°C.
- Root Emergence: Adventitious roots ("hairy roots") typically emerge from wound sites within 1-3 weeks. Non-transformed roots rarely appear on hormone-free medium.
- Excision & Subculture: Excise individual root tips (1-2 cm) and transfer to fresh solid or liquid MS medium with antibiotics (cefotaxime concentration can be gradually reduced).
- Confirmation of Transformation:
- PCR: Using primers for rolB or rolC genes.
- Opine Assay: High-voltage paper electrophoresis of root extract to detect opines (e.g., agropine, mannopine).
- Scale-Up: For metabolite production, transfer established lines to liquid MS medium in flasks or bioreactors. Culture in the dark with orbital shaking (90-110 rpm) at 25°C.
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagent Solutions for Hairy Root Research
| Item | Function/Description | Typical Concentration/Type |
|---|---|---|
| Acetosyringone | Phenolic compound that activates the Vir region of the Ri plasmid. Critical for efficient T-DNA transfer. | 100-200 µM in co-cultivation medium. |
| MS/B5 Basal Salts | Provides essential macro and micronutrients for plant tissue and root growth. Hormone-free. | ½ or full strength, with vitamins. |
| Cefotaxime | β-lactam antibiotic used to eliminate A. rhizogenes after co-cultivation without harming plant tissues. | 250-500 mg/L in post-co-culture media. |
| Opine Standard Mix | Reference compounds (agropine, mannopine) for confirming transformation via electrophoresis. | Analytical standard for assay validation. |
| rol Gene Primers | For PCR-based confirmation of T-DNA integration into the plant genome. | Specific to rolB, rolC, or other T-DNA genes. |
| CTAB Buffer | Cetyltrimethylammonium bromide buffer for high-quality genomic DNA isolation from hairy roots (polysaccharide-rich tissue). | Contains CTAB, NaCl, EDTA, Tris-HCl. |
This document provides detailed application notes and protocols within the context of a broader thesis on Agrobacterium rhizogenes-mediated root transformation protocol research. The natural mechanism of hairy root induction, driven by the integration and expression of root-inducing (Ri) plasmid genes into the plant genome, offers a robust platform for the production of plant-derived pharmaceuticals, metabolic engineering, and functional genomics studies for drug development professionals.
Core Mechanism: rol Genes and Auxin
Hairy root disease is incited by the integration of Transfer-DNA (T-DNA) from the Ri plasmid of A. rhizogenes into the plant genome. The key genetic elements are the rol (root loci) genes and genes influencing auxin biosynthesis.
The rol Gene Cluster (rolA, rolB, rolC, rolD): These genes are the primary drivers of hairy root phenotype.
- rolB & rolC: Exhibit β-glucosidase and cytokinin-β-glucosidase activities, respectively. They hydrolyze conjugated phytohormones (auxins and cytokinins), increasing the pool of active free hormones, disrupting the endogenous hormonal balance crucial for root induction.
- rolA & rolD: Modulate plant sensitivity to hormones and stress responses, synergizing with rolB and rolC.
Auxin Biosynthesis Genes (aux1, aux2): Present in some Ri plasmid strains (e.g., agropine type), these genes are homologous to the iaaM and iaaH genes of A. tumefaciens. They direct the synthesis of indole-3-acetic acid (IAA) from tryptophan, providing a localized, constitutive auxin source that profoundly stimulates cell division and root meristem formation.
The combined action of constitutive auxin production and the rol-mediated perturbation of hormone homeostasis leads to the continuous proliferation of genetically stable, hormone-autotrophic roots.
Key Research Reagent Solutions
Table 1: Essential Reagents for Hairy Root Induction and Analysis
| Reagent/Material | Function/Application | Example/Notes |
|---|---|---|
| Agrobacterium rhizogenes Strains | Delivery vector for Ri plasmid T-DNA. | ATCC 15834 (agropine-type, contains aux genes), A4 (agropine-type), R1000 (mannopine-type, lacks aux genes). Strain choice affects morphology and secondary metabolite production. |
| Plant Explant | Target tissue for transformation. | Leaf discs, stem internodes, cotyledons, or seedling fragments from target species (e.g., Nicotiana tabacum, Catharanthus roseus, Glycyrrhiza uralensis). |
| Co-cultivation Media (MS/B5) | Supports plant-Agrobacterium interaction post-inoculation. | MS Basal Salts with 3% sucrose, pH 5.8, often with Acetosyringone (100-200 µM) to induce vir gene expression. |
| Selection & Decontamination Media | Selects transformed roots and eliminates Agrobacterium. | MS/B5 media with antibiotics (e.g., Cefotaxime 250-500 mg/L, Vancomycin) and selection agents (e.g., Kanamycin if using binary vectors). |
| Hormone-Free Root Growth Media | Maintains and proliferates established hairy root cultures. | ½ or full-strength MS/B5 liquid/solid media without exogenous auxins/cytokinins. Demonstrates hormone autotrophy. |
| PCR Reagents for Confirmation | Molecular verification of T-DNA integration. | Primers for rolB, rolC, virG (contamination check), and an endogenous plant gene (positive control). |
| IAA/Auxin Quantification Kits | Measures auxin levels in transformed vs. wild-type roots. | ELISA-based or LC-MS/MS protocols for precise IAA quantification. |
Table 2: Representative Data on Transformation Efficiency and Metabolite Yield
| Plant Species | A. rhizogenes Strain | Explant Type | Avg. Transformation Efficiency* (%) | Reported Increase in Target Metabolite (vs. Untreated Root) | Reference Year |
|---|---|---|---|---|---|
| Panax ginseng | ATCC 15834 | Leaf | 65-80% | Ginsenosides: 2.5-4.0 fold | 2022 |
| Artemisia annua | A4 | Seedling stem | 70-85% | Artemisinin: 3.1 fold | 2023 |
| Beta vulgaris | R1000 | Cotyledon | 40-60% | Betalains: 12.0 fold | 2021 |
| Ophiorrhiza pumila | ATCC 15834 | Leaf | 55-70% | Camptothecin: 5.2 fold | 2023 |
| Nicotiana benthamiana | MSU440 | Stem | >90% | Recombinant Protein: 50 mg/kg FW | 2024 |
*Transformation Efficiency = (No. of explants producing hairy roots / Total no. of infected explants) x 100. * Data are illustrative from recent literature; values are compound- and system-dependent. ** Used for transient recombinant protein production via "hairy root" system with binary vectors.
Experimental Protocols
Protocol 1: Standard Hairy Root Induction in Dicotyledonous Plants
A. Preparation:
- Bacterial Culture: Inoculate a single colony of A. rhizogenes (e.g., ATCC 15834) into 10 mL YEB/LB broth with appropriate antibiotics. Shake (200 rpm) at 28°C for 24-48h until OD600 ~0.6-1.0. Pellet cells (5000xg, 10 min) and resuspend in liquid co-cultivation medium to OD600 ~0.2-0.5.
- Explant Preparation: Surface-sterilize plant material (e.g., leaves). Cut into 0.5-1 cm² segments under aseptic conditions.
B. Inoculation & Co-cultivation:
- Immerse explants in the bacterial suspension for 10-30 minutes. Blot dry on sterile filter paper.
- Place explants on solid co-cultivation medium (MS salts, 3% sucrose, pH 5.8, + 100 µM acetosyringone). Seal plates and incubate in the dark at 23-25°C for 2-3 days.
C. Decontamination & Selection:
- Transfer explants to decontamination/selection medium (hormone-free MS medium + 400-500 mg/L cefotaxime). Subculture to fresh medium every 7-10 days to eliminate bacteria.
- Within 1-4 weeks, hairy roots emerge from wound sites. Excise independent root tips (2-3 cm) and transfer to fresh selection/maintenance media.
D. Confirmation:
- Molecular PCR: Isolate genomic DNA from putative hairy roots. Perform PCR with primers for rolB (~423 bp) and rolC (~534 bp). Include a virG primer set to check for bacterial contamination.
Primer Sequences (Example):
- rolB F: 5′-GCTCTTGCAGTGCTAGATTT-3′, R: 5′-GAAGGTGCAAGCTACCTCTC-3′
- rolC F: 5′-CTCCTGACATCAAACTCGTC-3′, R: 5′-TGCTTCGAGTTATGGGTACA-3′
Protocol 2: Quantification of Endogenous IAA in Hairy Roots
Materials: Liquid N₂, cold methanol/water buffer, Internal Standard (deuterated IAA, d-IAA), HPLC-MS/MS system. Procedure:
- Extraction: Grind 100 mg FW of hairy root tissue in liquid N₂. Extract with 1 mL cold methanol/water (80:20, v/v) containing 50 ng d-IAA. Shake at 4°C for 2h. Centrifuge (13,000xg, 20 min, 4°C).
- Purification: Pass supernatant through a C18 solid-phase extraction column. Elute IAA with 80% methanol. Dry eluent under N₂ gas.
- Derivatization & Analysis: Reconstitute in 50 µL methanol for direct LC-MS/MS analysis (MRM mode). Calculate IAA concentration using the standard curve from pure IAA and normalized to the d-IAA recovery.
Visualizations
Within the context of developing optimized Agrobacterium rhizogenes-mediated root transformation protocols, the derived "hairy root" cultures unlock significant advantages for biomedical research. These rapidly growing, genetically stable root systems serve as versatile platforms for recombinant protein production, complex metabolite biosynthesis, and phytoremediation studies. This document provides application notes and detailed protocols leveraging this transformative technology.
Protein Production in Hairy Root Cultures
Application Notes: Hairy roots offer a eukaryotic protein production system with post-translational modification capabilities, often yielding higher functional protein fidelity than bacterial systems. They are particularly advantageous for producing complex mammalian proteins, enzymes, and diagnostic antigens. Recent studies show yields for specific antibodies (e.g., IgG1) can reach 20-30 µg/g fresh weight in optimized systems.
Protocol: Recombinant Protein Expression inA. rhizogenes-Derived Hairy Roots
Key Reagents: A. rhizogenes strain R1000, binary vector with CaMV 35S promoter and target gene, sterile Nicotiana benthamiana or Daucus carota explants, Acetosyringone, MS/B5 medium, selection antibiotic (e.g., kanamycin).
Procedure:
- Vector Construction: Clone gene of interest into a plant-optimized binary vector (e.g., pBI121 derivative) with a strong constitutive or inducible promoter and ER retention signal (SEKDEL) if needed.
- Bacterial Preparation: Transform the vector into A. rhizogenes. Grow a single colony in 10 mL YEB medium with appropriate antibiotics (e.g., rifampicin, kanamycin) at 28°C, 200 rpm for 24h.
- Induction: Pellet bacteria at 3000 x g for 10 min. Resuspend to OD600 ~0.5 in liquid MS medium with 100 µM acetosyringone. Incubate at room temperature for 1h.
- Plant Transformation: Inoculate sterile explants (leaf discs, cotyledons) by wounding and co-cultivation with the induced bacterial suspension for 20 min. Blot dry and place on co-cultivation medium (MS + acetosyringone) for 48h in dark.
- Hairy Root Initiation & Selection: Transfer explants to MS/B5 solid medium containing the selection antibiotic (e.g., 100 mg/L kanamycin) and a bactericide (e.g., 300 mg/L cefotaxime). Incubate at 25°C in dark. Hairy roots emerge in 1-2 weeks.
- Root Line Selection & Culture: Excise independent hairy root tips (2-3 cm). Grow in liquid MS/B5 medium with selection antibiotic on a rotary shaker (80-100 rpm) in the dark. Subculture every 2-3 weeks.
- Protein Analysis: Harvest roots, homogenize in extraction buffer (e.g., PBS with protease inhibitors). Quantify recombinant protein via ELISA or functional assay. Purify via affinity chromatography.
Table 1: Representative Protein Yields in Hairy Root Systems
| Protein Class | Example | Host Species | Typical Yield (µg/g FW) | Key Factor |
|---|---|---|---|---|
| Full Antibody | IgG1 | Nicotiana spp. | 20-35 | ER-targeting, codon optimization |
| Single-Chain Fv | scFv anti-HIV | Daucus carota | 15-25 | Secretion signal peptide |
| Therapeutic Enzyme | Glucocerebrosidase | Glycine max | 10-20 | Inducible promoter system |
| Vaccine Antigen | Hepatitis B surface Ag | Solanum tuberosum | 30-50 | Subcellular targeting (apoplast) |
| Growth Factor | Human FGF2 | Beta vulgaris | 5-15 | Fusion with stabilizing partner |
Metabolic Engineering for High-Value Compounds
Application Notes: Hairy roots are natural biosynthetic factories for secondary metabolites. Metabolic engineering enhances the production of plant-derived pharmaceuticals (e.g., alkaloids, terpenoids, phenolics). Recent successes include engineering the tropane alkaloid pathway for scopolamine, achieving titers >100 mg/L in optimized bioreactors.
Protocol: Metabolic Pathway Engineering in Hairy Roots
Key Reagents: A. rhizogenes strain ATCC 15834, Gateway-compatible binary vectors, RNAi or CRISPR/Cas9 constructs for gene silencing/editing, explants from medicinal plant (e.g., Artemisia annua, Catharanthus roseus), precursor feeding compounds (e.g., loganin, secologanin).
Procedure:
- Target Identification: Use transcriptomic/metabolomic data to identify pathway bottlenecks (e.g., rate-limiting enzyme, competing branch).
- Construct Design: Assemble constructs for: a) Overexpression of key biosynthetic genes (e.g., P450s, reductases) using strong promoters. b) RNAi vectors to silence competing pathways. c) CRISPR/Cas9 for knockout of repressor genes.
- Hairy Root Generation: Follow the transformation protocol in Section 1, using medicinal plant explants and the engineered A. rhizogenes.
- Screening: Screen 20-30 independent root lines by qRT-PCR for transgene expression and HPLC/MS for metabolite profile. Select top 3-5 lines.
- Culture Optimization: Grow selected lines in varied media (e.g., B5, WP) with different sucrose concentrations (1-5%), pH (5.0-6.0), and elicitors (e.g., 100 µM methyl jasmonate added on day 14).
- Bioreactor Scale-Up: Transfer high-producing lines to a mist or bubble column bioreactor (e.g., 1 L capacity). Monitor biomass accumulation and metabolite production over 4-6 weeks.
- Metabolite Extraction & Analysis: Lyophilize roots, pulverize. Extract with methanol:water (70:30). Analyze via UPLC-QTOF-MS against authentic standards.
Table 2: Metabolite Yield Enhancement via Engineered Hairy Roots
| Target Compound | Plant Source | Engineered Modification | Fold Increase Over Wild-Type | Final Titer (mg/L) |
|---|---|---|---|---|
| Artemisinin | Artemisia annua | Overexpression of ADS and CYP71AV1 | 8-10 | 80-100 |
| Scopolamine | Atropa belladonna | Overexpression of H6H; RNAi of PMT | >15 | 120-150 |
| Resveratrol | Vitis vinifera | Overexpression of STS; Suppression of CHS | 20-25 | 45-60 |
| Shikonin | Lithospermum erythrorhizon | Elicitation (MJ, SA) & Precursor feeding | 6-8 | 200-250 |
| Betalains | Beta vulgaris | Overexpression of CYP76AD1 and DODA | 12-15 | 500-700 |
Phytoremediation and Phytoextraction Studies
Application Notes: Hairy roots provide a controlled model to study plant-microbe interactions and mechanisms of pollutant uptake, degradation, and sequestration. They are used to test genetic modifications that enhance tolerance and accumulation of heavy metals (e.g., Cd, As) or organic xenobiotics (e.g., PCBs, TNT).
Protocol: Phytoextraction Assay Using Hairy Roots
Key Reagents: Hairy roots of hyperaccumulator species (e.g., Sedum alfredii, Arabidopsis halleri), spiked medium with target pollutant (e.g., CdCl₂, PbNO₃), ICP-MS calibration standards, chelating agents (e.g., EDTA), detection reagents for reactive oxygen species (e.g., DCFH-DA, NBT).
Procedure:
- Root Culture Establishment: Generate hairy roots from a known hyperaccumulator or a transgenic plant expressing metal transporters (e.g., YSL1, PCS1).
- Exposure Experiment: After 14 days of growth, transfer root clusters (~0.5 g FW) to fresh medium containing a gradient of the target pollutant (e.g., 0, 50, 100, 200 µM Cd). Include 3-5 biological replicates per condition.
- Growth & Stress Monitoring: Harvest roots at 0, 24, 48, 96h. Record fresh/dry weight. Assess stress via histochemical staining (NBT for superoxide, DCFH-DA for H₂O₂).
- Metal Quantification: Rinse roots in 5 mM CaCl₂ for 15 min to remove surface-bound metals. Oven-dry (70°C, 48h). Digest 50 mg DW in 3 mL concentrated HNO₃ at 120°C for 3h. Dilute and analyze via ICP-MS.
- Translocation Factor (TF) Calculation: If a shoot is present, calculate TF = [Metal]shoot / [Metal]root.
- Localization Studies: For visualization, use leadmium green AM dye for Pb/Cd or perform dithizone staining for Zn/Cd on fresh root sections.
- Microbial Augmentation: Co-culture roots with pollutant-degrading endophytic bacteria (e.g., Pseudomonas putida) to assess synergistic effects on degradation of organics.
Table 3: Phytoextraction Capacity of Engineered Hairy Root Systems
| Target Pollutant | Root System | Key Genetic Mod/Strategy | Uptake (µg/g DW) | Translocation Factor |
|---|---|---|---|---|
| Cadmium (Cd) | Arabidopsis thaliana (WT) | None (control) | 180-220 | 0.1-0.3 |
| Cadmium (Cd) | A. thaliana (Transgenic) | Overexpression of AtHMA4 (P1B-ATPase) | 950-1200 | 0.8-1.2 |
| Arsenic (As) | Oryza sativa | Overexpression of OsACR2.1 (arsenate reductase) | 350-400 | 1.5-2.0 |
| Lead (Pb) | Brassica juncea | Soil amendment with EDTA in model system | 5000-7000 | 0.05-0.1 |
| Trinitrotoluene (TNT) | Nicotiana tabacum | Co-culture with engineered P. putida | Degradation: 95% in 72h | N/A |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function | Example Product/Source |
|---|---|---|
| A. rhizogenes Strains | Delivery of T-DNA for root transformation | R1000, ATCC 15834, K599 (Arqual) |
| Binary Vectors | Carrying gene of interest for expression or editing | pBI121, pK7WG2D, pCambia series |
| Acetosyringone | Phenolic inducer of vir gene expression | Sigma-Aldrich, A13406 |
| MS/B5 Medium | Nutrient base for hairy root culture | PhytoTech Labs, M519, B589 |
| Selection Antibiotics | For selecting transgenic roots (plant) & eliminating bacteria | Kanamycin, Hygromycin B, Cefotaxime |
| Elicitors | Induce secondary metabolite pathways | Methyl Jasmonate (Sigma 392707), Salicylic Acid |
| ICP-MS Standards | For quantitative metal analysis in phytoremediation | Multi-element standard solutions (Merck) |
| ROS Detection Kits | Visualize oxidative stress in roots | DCFH-DA (Sigma D6883), NBT (Sigma N6876) |
| Affinity Resins | Purification of recombinant proteins (e.g., His-tag, Protein A) | Ni-NTA Agarose (Qiagen), MabSelect SuRe |
| UPLC-QTOF-MS System | High-resolution metabolomic profiling | Waters ACQUITY UPLC/Xevo G2-XS |
Experimental Workflow and Pathway Visualizations
Title: Workflow for Recombinant Protein Production in Hairy Roots
Title: Engineered Metabolic Pathway for Compound Production
Title: Pollutant Fate in Hairy Root Phytoremediation Model
Within the context of Agrobacterium rhizogenes-mediated root transformation protocol research, selecting the appropriate bacterial strain is a critical determinant of transformation efficiency, root morphology, and transgene expression stability. This application note provides a comparative analysis of three strains: the wild-type strain Arqual, the widely used engineered strain A4, and the disarmed strain LBA9402.
Table 1: Comparative Overview of Key A. rhizogenes Strains
| Feature | Arqual (Wild-type) | A4 (R1000) | LBA9402 (pRi1855) |
|---|---|---|---|
| Ri Plasmid Type | Agropine-type (pRiA4) | Agropine-type (pRiA4) | Agropine-type (pRi1855, TL-DNA disarmed) |
| Oncogenicity | High (full TL-DNA) | High (full TL-DNA) | Low (disarmed TL-DNA, aux genes deleted) |
| Root Morphology | Extensive, highly branched, hairy root phenotype | Extensive, highly branched, hairy root phenotype | More controlled, "normalized" root growth |
| Key Genetic Markers | Naturally resistant to opines | Spontaneous rifampicin resistance (Rif⁺) | Streptomycin resistance (Str⁺), Carbenicillin resistance (Carb⁺) |
| T-DNA Transfer Efficiency | High | Very High | Moderate to High |
| Primary Application | Inducing classic "hairy root" disease; study of root-pathogen interaction. | High-efficiency generation of transgenic hairy root cultures for metabolite production. | Generation of composite plants with transgenic roots on non-transgenic shoots; functional genomics. |
| Major Advantage | Robust, natural virulence. | Reliable, high transformation rates across many species. | Non-tumori genic; allows for regeneration of plants with transgenic roots only. |
| Major Limitation | Cannot regenerate normal plants; may overgrow cultures. | Cannot regenerate normal plants; may overgrow cultures. | May have lower transformation efficiency for some species compared to wild-type strains. |
Table 2: Quantitative Performance Metrics in Model Plants (Representative Data)
| Strain | Tomato (% Root Initiation) | Nicotiana benthamiana (Roots/Explant) | Glycine max (Transformation Frequency %) | Medicago truncatula (Hairy Root Yield) |
|---|---|---|---|---|
| Arqual | 75-85% | 15-20 | 40-60% | High |
| A4 | 85-95% | 18-25 | 50-75% | Very High |
| LBA9402 | 60-75% | 10-15 | 30-50% | Moderate |
Experimental Protocols
Protocol 2.1: Strain Preparation and Maintenance
Purpose: To culture and prepare competent A. rhizogenes cells for plant transformation. Materials: See Scientist's Toolkit. Method:
- Streak strain from glycerol stock onto appropriate solid medium (YEB + antibiotics). Incubate at 28°C for 2 days.
- Pick a single colony and inoculate 5 mL liquid medium with antibiotics. Shake at 200 rpm, 28°C for 24-48h.
- For transformation, subculture 1 mL into 50 mL fresh medium without antibiotics. Grow to mid-log phase (OD₆₀₀ ≈ 0.5-0.8).
- Pellet cells at 5000 x g for 10 min at 4°C. Resuspend in an equal volume of ice-cold 20 mM CaCl₂ or liquid plant co-culture medium.
- Use immediately for plant explant inoculation or store competent cells on ice for short-term use.
Protocol 2.2: Hairy Root Induction and Composite Plant Generation
Purpose: To transform plant explants and generate either hairy root cultures (A4/Arqual) or composite plants (LBA9402). Materials: Sterile plant explants (e.g., cotyledons, leaf discs), co-culture medium, selection medium containing antibiotics (e.g., cefotaxime to kill bacteria, kanamycin for T-DNA selection). Method for Hairy Root Cultures (A4/Arqual):
- Wound explants lightly with a scalpel dipped in the bacterial suspension.
- Co-culture explants on solid hormone-free medium for 2-3 days in the dark at 25°C.
- Transfer explants to selection medium containing antibiotics. Hairy roots should emerge from infection sites in 1-3 weeks.
- Excise independent root tips and transfer to fresh selection medium for clonal expansion. Method for Composite Plants (LBA9402):
- Inoculate the cut surface of a seedling stem or hypocotyl with bacteria.
- Allow plants to grow under normal conditions. Adventitious transgenic roots emerge at the site of inoculation.
- Remove the non-transgenic primary root to encourage growth of transgenic roots.
- Confirm transformation via reporter gene expression (e.g., GFP) or PCR.
Signaling and Selection Workflow
Diagram Title: Strain Selection Decision Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for A. rhizogenes Root Transformation
| Item | Function | Example/Note |
|---|---|---|
| Bacterial Strains | Source of T-DNA for root transformation. | Arqual (WT), A4 (Rif⁺), LBA9402 (Str⁺, Carb⁺). |
| YEB Medium | Rich medium for optimal growth of Agrobacterium. | Contains beef extract, yeast extract, peptone, sucrose, MgSO₄. |
| Antibiotics | Selective pressure for plasmid maintenance and elimination of bacteria post-co-culture. | Rifampicin (for A4), Streptomycin (for LBA9402), Cefotaxime (to kill Agrobacterium after co-culture). |
| Plant Tissue Culture Media (MS/B5) | Support growth and development of plant explants and transgenic roots. | Hormone-free for hairy root induction; may contain auxins for specific species. |
| Selection Agents (Plant) | Select for transformed plant cells. | Kanamycin, Hygromycin B, depending on T-DNA selectable marker. |
| Acetosyringone | Phenolic compound that induces vir gene expression, enhancing T-DNA transfer. | Add to bacterial pre-culture and co-culture media (100-200 µM). |
| Sterile Explant Material | Target tissue for transformation. | Cotyledons, leaf discs, hypocotyl segments, seedling stems. |
| Reporting Vectors | Allow visual screening of transformation events. | pCAMBIA or pBin19 derivatives with GFP, GUS, or RFP reporters. |
| PCR Reagents | Molecular confirmation of transgene integration. | Primers for rol genes (transformation) or transgene-specific primers. |
The choice of host plant is a critical first step in Agrobacterium rhizogenes-mediated root transformation (hairy root culture) for recombinant protein or metabolite production. This decision balances experimental tractability against translational relevance. The following table summarizes key quantitative parameters for common host species.
Table 1: Comparative Analysis of Selected Plant Hosts for A. rhizogenes Transformation
| Host Species | Type | Transformation Efficiency (Typical Range) | Root Growth Rate (Doubling Time) | Secondary Metabolite Complexity | Genetic Tools Availability | Primary Applications |
|---|---|---|---|---|---|---|
| Arabidopsis thaliana | Dicot Model | 70-90% | 2-3 days | Low | Extensive (Mutants, Reporters) | Fundamental Research, Signaling Studies |
| Nicotiana benthamiana | Dicot Model | 80-95% | 3-4 days | Medium | Very Good (Transient O.E., VIGS) | High-yield Protein Production, Pathway Reconstitution |
| Medicago truncatula | Legume Model | 50-70% | 5-7 days | High (Triterpenes, Flavonoids) | Good (Mutants, Genomics) | Plant-Microbe Interactions, Specialized Metabolism |
| Cannabis sativa | Medicinal | 20-40% | 7-10 days | Very High (Cannabinoids, Terpenes) | Emerging (Genome, Reg. Elements) | Cannabinoid Pathway Engineering |
| Artemisia annua | Medicinal | 30-50% | 7-12 days | Very High (Artemisinin) | Limited | Artemisinin Pathway Optimization |
Detailed Methodologies
Protocol: Standardized Hairy Root Induction for Comparative Studies
This protocol is optimized for side-by-side transformation of model and medicinal species to assess host-dependent performance.
Research Reagent Solutions & Essential Materials
| Item Name | Function/Benefit | Example Product/Catalog # |
|---|---|---|
| A. rhizogenes Strain R1000 | Wild-type strain, high virulence, non-disarmed. | N/A (Available from culture collections) |
| MS (Murashige and Skoog) Basal Salt Mixture | Provides essential macro and micronutrients for plant tissue culture. | PhytoTech Labs, M519 |
| Acetosyringone | Phenolic compound that induces vir gene expression in Agrobacterium. | Sigma-Aldrich, D134406 |
| Gamborg's B5 Vitamins | Vitamin supplement for robust root growth, often superior to MS vitamins. | PhytoTech Labs, G341 |
| Cefotaxime Sodium Salt | β-lactam antibiotic to eliminate Agrobacterium after co-cultivation. | GoldBio, C-120-5 |
| Sterile Cellulose Acetate Membrane | Supports explants during co-cultivation, allows for easy transfer. | Millipore, HAWP04700 |
| pH Meter with Temperature Compensation | Critical for accurate media preparation. | Mettler Toledo, FiveEasy Plus |
| Laminar Flow Hood (Biosafety Cabinet) | Provides aseptic environment for all tissue culture steps. | Labconco, Purifier Vertical |
Procedure:
- Plant Material Preparation: Surface-sterilize seeds of target species (e.g., Arabidopsis, Nicotiana, Artemisia) with 70% ethanol (1 min) followed by 2% sodium hypochlorite with 0.1% Tween-20 (10 min). Rinse 5x with sterile water. Germinate on half-strength MS agar plates in a growth chamber (species-appropriate light/temperature).
- Agrobacterium Culture Preparation: Inoculate a single colony of A. rhizogenes (e.g., R1000 carrying a binary vector with your gene of interest and a root-specific selectable marker like DsRed1) in 5 mL LB with appropriate antibiotics. Grow overnight at 28°C, 220 rpm. Pellet cells at 3000 g for 15 min and resuspend in induction medium (MS liquid, pH 5.2, 100 µM acetosyringone) to an OD600 of 0.6-0.8. Incubate at room temperature, shaking gently for 2-4 hours.
- Explants & Co-cultivation: Using 10-14 day old seedlings, prepare explants: hypocotyl segments for Arabidopsis, leaf discs for Nicotiana, or young stem segments for medicinal species. Immerse explants in the induced Agrobacterium suspension for 20-30 minutes. Blot dry on sterile paper and place on co-cultivation medium (MS/B5 salts, vitamins, 100 µM acetosyringone, 0.8% agar, pH 5.2) overlaid with a sterile cellulose acetate membrane. Co-cultivate in the dark at 22-25°C for 2-3 days.
- Root Induction & Selection: Transfer explants to selection/induction medium (MS/B5 salts, vitamins, 3% sucrose, 0.8% agar, pH 5.8, 300-500 mg/L cefotaxime, and appropriate antibiotic/herbicide for vector selection). Culture at 25°C in the dark. Hairy roots should emerge at wound sites within 1-3 weeks, depending on species.
- Root Line Establishment: Excise individual, healthy, fast-growing hairy roots and transfer to fresh selection medium (with reduced cefotaxime, e.g., 100 mg/L) in petri dishes or liquid culture flasks. Establish clonal lines by successive sub-culturing of root tips.
Protocol: Metabolite Profiling of Hairy Root Lines
Procedure:
- Biomass Harvest: Collect hairy root biomass from established lines (minimum 100 mg FW). Rinse briefly with deionized water, blot dry, and flash-freeze in liquid nitrogen.
- Metabolite Extraction: Grind tissue to a fine powder under liquid nitrogen. For polar metabolites (e.g., artemisinic acids), extract with 80% methanol/water (v/v). For non-polar metabolites (e.g., cannabinoids, terpenes), extract with hexane or chloroform/methanol mixtures. Sonicate for 15 min, centrifuge at 12,000 g for 10 min, and collect supernatant.
- Analysis (HPLC-DAD/LC-MS): Filter extracts through a 0.22 µm PTFE syringe filter. Separate metabolites using a reverse-phase C18 column (e.g., Agilent ZORBAX Eclipse Plus). Use a gradient elution (e.g., water/acetonitrile, both with 0.1% formic acid). Detect using a Diode Array Detector (DAD) and/or Mass Spectrometer (MS). Quantify by comparison to authentic standards.
Visualized Workflows & Pathways
Host Selection to Analysis Workflow
A. rhizogenes Signaling & Root Induction
Step-by-Step Protocol: From Vector Design to Hairy Root Culture Establishment
This protocol details Phase 1 of a comprehensive thesis on Agrobacterium rhizogenes-mediated root transformation. The initial phase focuses on the critical decision point between two principal vector systems for transforming plant hosts: the binary vector system and the co-integrate vector system. The choice of system impacts efficiency, flexibility, and downstream applications in research and drug development, such as the production of recombinant proteins or secondary metabolites in hairy root cultures.
Core Vector Systems: Comparison and Selection
The transformation efficiency hinges on the type of vector system used to deliver T-DNA from A. rhizogenes into the plant genome. The following table summarizes the key characteristics, advantages, and recent performance data for both systems.
Table 1: Comparative Analysis of Binary vs. Co-integrate Vector Systems for A. rhizogenes Transformation
| Feature | Binary Vector System | Co-integrate Vector System |
|---|---|---|
| Basic Structure | T-DNA and Virulence (vir) genes are on separate plasmids: a disarmed Ri plasmid in the bacterium and a small, engineered T-DNA vector (pBinary). | T-DNA region from an intermediate vector is integrated via homologous recombination into a resident, disarmed Ri plasmid. |
| Common Vector Backbones | pCAMBIA, pBI121, pGreen, pK7WG2D. | pRiA4::pARC8, pRi1855::pTVK291. |
| Typical Transformation Efficiency (Root Induction %) | 65-85% (varies by plant species, e.g., Nicotiana tabacum: ~80%, Glycine max: ~70%) | 40-60% (generally lower due to recombination step) |
| T-DNA Insert Size Limit | Large (>50 kbp) | Moderate (~25-30 kbp) |
| Cloning & Manipulation | High flexibility. Cloning is performed in E. coli, then mobilized into A. rhizogenes. | Less flexible. Requires recombination in Agrobacterium, often slower. |
| Stability in Agrobacterium | High; the small binary vector is easily maintained with selection. | Very high; once integrated, the T-DNA is stable as part of the Ri plasmid. |
| Suitability for High-Throughput | Excellent; amenable to Gateway, Golden Gate cloning. | Poor; not ideal for rapid, multi-gene constructs. |
| Primary Application Context | Modern studies requiring rapid, flexible construct assembly, multi-gene stacking, and CRISPR/Cas9 editing. | Historical studies; may be used for specific, stable, single-gene insertions where binary vectors are unstable. |
Experimental Protocols
Protocol 3.1: Construction of a Binary Vector forA. rhizogenes
Objective: To clone a gene of interest (GOI) into a binary vector and introduce it into a disarmed A. rhizogenes strain (e.g., ARqua1, K599).
Materials:
- Research Reagent Solutions:
- LB Broth/Agar: For cultivation of E. coli and A. rhizogenes.
- SOC Medium: For recovery of transformed E. coli.
- Antibiotics: Kanamycin, Spectinomycin, Rifampicin. Prepare sterile stock solutions.
- Restriction Enzymes & Ligase: For traditional cloning (e.g., XbaI/BamHI, T4 DNA Ligase).
- Gateway LR Clonase II Mix: For recombination-based cloning.
- Acetosyringone (100 mM stock in DMSO): Vir gene inducer.
- 20 mM CaCl₂, Ice-cold: For preparation of competent A. rhizogenes.
- Freeze Media: 15% Glycerol in LB broth.
Procedure:
- In E. coli: a. Perform standard cloning (restriction/ligation or Gateway recombination) to insert the GOI and desired plant selection marker (e.g., hptII for hygromycin) into the T-DNA region of the binary vector (e.g., pCAMBIA1302). b. Transform the construct into competent E. coli (e.g., DH5α). Select on LB agar with appropriate antibiotics (e.g., Kanamycin 50 µg/mL). c. Isolate plasmid DNA from confirmed colonies using a miniprep kit. Verify construct by restriction digest and sequencing.
Mobilization to A. rhizogenes (Freeze-Thaw Method): a. Grow a disarmed A. rhizogenes strain (rifampicin-resistant) overnight in 5 mL LB + Rif (50 µg/mL) at 28°C, 200 rpm. b. Sub-culture 1 mL into 50 mL fresh LB+Rif. Grow to an OD600 of 0.5-0.8. c. Chill cells on ice for 30 min. Pellet at 4000 x g for 10 min at 4°C. d. Gently resuspend pellet in 1 mL of ice-cold 20 mM CaCl₂. e. Mix 100 µL of competent cells with 100-500 ng of the verified binary plasmid. Incubate on ice for 30 min. f. Freeze in liquid nitrogen for 5 min, then thaw at 37°C for 5 min. g. Add 1 mL of SOC medium, incubate at 28°C for 2-4 hours with shaking. h. Plate 100-200 µL onto LB agar containing Rif and the binary vector-specific antibiotic (e.g., Kanamycin 50 µg/mL). Incubate at 28°C for 2-3 days.
Validation: a. Perform colony PCR using primers specific to the GOI or the vector backbone to confirm transformation. b. Prepare a glycerol stock of a positive colony for long-term storage at -80°C.
Protocol 3.2: Transformation using a Co-integrate Vector System
Objective: To integrate a T-DNA from an intermediate vector into the resident Ri plasmid via homologous recombination.
Materials: (As in Protocol 3.1, with specific antibiotics for the intermediate vector, e.g., pTVK291 derivatives often use Tetracycline).
Procedure:
- Construct Intermediate Vector: Clone the GOI into the T-DNA of an intermediate vector (e.g., pTVK291) in E. coli. This vector contains a region homologous to the disarmed Ri plasmid (e.g., pRiA4).
- Conjugative Transfer (Triparental Mating): a. Grow three cultures overnight: (1) E. coli with the intermediate vector, (2) Helper E. coli with a conjugation-helper plasmid (e.g., pRK2013), and (3) The disarmed A. rhizogenes recipient strain. b. Mix 100 µL of each culture on a sterile nitrocellulose filter placed on an LB agar plate (no antibiotics). Incubate at 28°C for 24 hours. c. Resuspend the cell mixture from the filter and plate onto selective media for A. rhizogenes. This medium contains Rif (for A. rhizogenes), Tetracycline (for the intermediate vector backbone), and an antibiotic to counterselect against both E. coli strains (e.g., Carbenicillin or Cefotaxime).
- Selection for Co-integrate: The selectable marker on the intermediate vector will only be maintained in A. rhizogenes if it has recombined into the resident Ri plasmid. Screen colonies by PCR using primers that span the junction between the Ri plasmid and the inserted T-DNA.
- Curing of Helper Plasmid: Streak positive colonies several times on selective media without the helper plasmid's antibiotic to ensure its loss.
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for Vector Construction and A. rhizogenes Transformation
| Reagent/Material | Function/Benefit |
|---|---|
| Disarmed A. rhizogenes Strains (e.g., ARqua1, K599, R1000) | Engineered for safety and efficacy; lack wild-type oncogenes, often carry a disarmed pRi, and are compatible with binary vectors. Essential for controlled root induction. |
| Binary Vector Backbones (e.g., pCAMBIA, pGreen series) | Contain plant selection markers, multiple cloning sites or recombination sites (attR/attL), and reporter genes (GFP, GUS). Enable flexible, high-efficiency cloning. |
| Acetosyringone | A phenolic compound that induces the expression of A. rhizogenes vir genes, which are essential for T-DNA processing and transfer. Critical for maximizing transformation efficiency during plant inoculation. |
| Plant Selection Antibiotics (e.g., Hygromycin, Kanamycin) | Allow for the in vitro selection of transformed plant cells or roots that express the corresponding resistance gene (hptII, nptII) within the T-DNA. |
| Gateway LR Clonase II Enzyme Mix | Enables rapid, efficient, and directional transfer of a GOI from an Entry clone into a binary Destination vector via site-specific recombination. Standardizes and speeds up vector construction. |
| Triparental Mating Helper Plasmid (e.g., pRK2013) | Provides trans-acting mobilization (tra) functions necessary for the conjugative transfer of non-mobilizable intermediate vectors from E. coli to Agrobacterium in co-integrate methods. |
Visualized Workflows and Signaling
Binary vs Co-integrate Vector Workflow
A. rhizogenes vir Gene Induction Pathway
This application note details the critical second phase of an Agrobacterium rhizogenes-mediated root transformation protocol. Successful transformation is contingent upon the generation of aseptic, viable, and genetically uniform plant explants. This phase establishes the foundation for efficient A. rhizogenes co-cultivation and subsequent hairy root induction, a vital system for studying root biology and producing recombinant pharmaceuticals.
Core Principles and Quantitative Targets
Effective sterilization eliminates microbial contaminants while preserving explant viability and regenerative capacity. The optimal protocol balances sterilant concentration, exposure time, and explant type.
Table 1: Common Sterilization Agents and Their Efficacy
| Sterilant Agent | Typical Concentration | Exposure Time Range | Target Contaminants | Key Considerations |
|---|---|---|---|---|
| Sodium Hypochlorite (NaOCl) | 0.5% - 5.0% (v/v) | 5 - 30 minutes | Broad-spectrum (fungi, bacteria) | Concentration/time trade-off; requires thorough rinsing. |
| Ethanol (C₂H₅OH) | 70% (v/v) | 30 seconds - 5 minutes | Broad-spectrum | Surface sterilant; rapid action; can penetrate tissues. |
| Hydrogen Peroxide (H₂O₂) | 3% - 15% (v/v) | 5 - 20 minutes | Broad-spectrum, especially fungal spores | Generates oxygen; good for heavily contaminated seeds. |
| Mercuric Chloride (HgCl₂) | 0.1% - 0.2% (w/v) | 2 - 10 minutes | Persistent contaminants | Highly toxic; last-resort agent; requires extreme care. |
Table 2: Sterilization Success Metrics for Common Explants
| Explant Type | Recommended Sterilant Protocol | Target Contamination Rate | Target Survival Rate |
|---|---|---|---|
| Mature Seeds | 70% EtOH (2 min) → 3% NaOCl + Tween (15 min) | <5% | >90% |
| Seedling Hypocotyls | 1% NaOCl + Tween (10 min) | <10% | >85% |
| Leaf Disks | 70% EtOH (30 sec) → 0.5% NaOCl (5 min) | <15% | >80% |
| Root Segments | 2% NaOCl + Tween (8 min) → 3% H₂O₂ (5 min) | <20% | >75% |
Detailed Protocol: Preparation and Sterilization of Seed Explants forA. rhizogenesTransformation
This protocol is optimized for generating sterile Nicotiana benthamiana or tomato seedlings for subsequent root transformation.
Materials Required
- Mature, high-quality seeds.
- Sterilization Solutions: 70% (v/v) ethanol, aqueous sodium hypochlorite (3% available chlorine) with 1-2 drops of Tween-20 per 100 mL, sterile distilled water (dH₂O).
- Equipment: Laminar flow hood, sterile forceps, sterile Petri dishes, sterile filter paper, autoclave, sterile germination media (½ strength MS basal salts).
Methodology
- Pre-Sterilization (Seed Hydration): Place seeds in a sterile 15 mL conical tube. Add 10 mL of sterile dH₂O. Allow seeds to imbibe for 30-60 minutes at room temperature. This hydrates the seed coat and can improve sterilization efficacy.
- Surface Sterilization: a. Decant the dH₂O. b. Add 10 mL of 70% ethanol. Vortex gently for 30 seconds. Decant ethanol completely. c. Add 10 mL of 3% NaOCl/Tween-20 solution. Invert tube continuously for 15 minutes. d. In the laminar flow hood, decant the sterilant.
- Rinsing: Perform five sequential rinses with 10 mL of sterile dH₂O each time (5 x 1 min), ensuring complete removal of residual sterilant.
- Germination: Aseptically transfer seeds onto sterile filter paper-lined Petri dishes containing ½ strength MS medium without growth regulators. Seal plates with porous tape.
- Incubation: Place plates in a growth chamber (25°C, 16/8 hour light/dark cycle) for 7-14 days until seedlings are established.
- Explant Preparation: Using sterile forceps and scalpel, excise hypocotyl segments (0.5-1.0 cm) from 7-10 day old seedlings for subsequent A. rhizogenes inoculation.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Plant Material Preparation & Sterilization
| Item | Function & Rationale |
|---|---|
| Sodium Hypochlorite (NaOCl) | Oxidizing agent that disrupts microbial cell walls and membranes; the workhorse of plant tissue culture sterilization. |
| Tween-20 (Polysorbate 20) | Non-ionic surfactant added to sterilants to reduce surface tension, ensuring even coverage and penetration of crevices on explant surfaces. |
| Murashige and Skoog (MS) Basal Salt Mixture | Provides essential macro and micronutrients for in vitro seed germination and explant survival post-sterilization. |
| Plant Preservative Mixture (PPM) | A broad-spectrum biocide used in culture media to suppress latent and airborne contaminants post-sterilization. |
| Sterile Cellulose Filter Paper | Provides a sterile, inert, and absorbent surface for seed germination, separating the explant from direct contact with agar. |
Signaling and Workflow Visualization
Diagram 1: Plant Material Sterilization and Germination Workflow
Diagram 2: Biochemical Mode of Action of Common Sterilants
Within the established framework of Agrobacterium rhizogenes-mediated root transformation protocol research, Phase 3—Inoculation—is the critical step that determines the efficiency of transgenic "hairy root" induction. This phase bridges bacterial preparation and post-inoculation culture, focusing on the delivery of the engineered A. rhizogenes strain carrying the gene(s) of interest into the explant tissue. The choice of inoculation method directly impacts transformation frequency, the speed of root emergence, and the viability of the explant. This document details three core methodologies: Wounding, Co-cultivation, and Direct Injection, providing application notes and standardized protocols for researchers and drug development professionals seeking to produce recombinant proteins or secondary metabolites in transformed root cultures.
Comparative Analysis of Inoculation Methods
The selection of an inoculation method depends on the explant type (e.g., leaf disc, cotyledon, stem segment, whole seedling), plant species, and experimental goals. The table below summarizes key quantitative and qualitative metrics from recent studies (2020-2024).
Table 1: Comparative Analysis of A. rhizogenes Inoculation Methods
| Method Parameter | Wounding (e.g., Stab/Prick) | Co-cultivation (Liquid/Plates) | Direct Injection (Syringe) |
|---|---|---|---|
| Primary Explant | Seedlings, thick stems, hypocotyls | Leaf discs, cotyledons, thin stems | Whole seedlings, developing organs |
| Avg. Transformation Efficiency (%) | 65-85% (species-dependent) | 70-90% | 40-70% (more variable) |
| Avg. Hairy Root Emergence Time | 7-14 days | 10-21 days | 14-28 days |
| Key Advantage | High efficiency for robust tissues; simple. | Excellent for fragile tissues; even bacterial exposure. | Targeted delivery; minimal physical damage to explant surface. |
| Key Limitation | Physical damage can stress explant. | Requires careful control of co-cultivation duration to avoid overgrowth. | Technically demanding; lower efficiency; risk of internal tissue damage. |
| Optimal [OD₆₀₀] of Bacterial Culture | 0.6 - 1.0 | 0.3 - 0.6 | 0.8 - 1.2 |
| Co-cultivation Duration | 24-72 hours (on moist paper/solid medium) | 48-96 hours (on selection-free medium) | 48-72 hours (plant kept in humid chamber) |
| Common Plant Models (Recent Studies) | Tomato, Brassica spp., Medicago truncatula | Nicotiana benthamiana, Lettuce, Soybean | Arabidopsis thaliana, Pea, Catharanthus roseus |
Detailed Protocols
Protocol: Wounding/Stab Inoculation
Application Note: Best for robust, whole seedling or stem explants where high transformation frequency is prioritized.
- Bacterial Preparation: Grow engineered A. rhizogenes (e.g., strain K599 or ARqua1) in appropriate liquid medium with antibiotics to an OD₆₀₀ of 0.8. Pellet bacteria at 5000 x g for 10 min and resuspend in sterile induction medium (e.g., MES buffer with acetosyringone, 100 µM) to a final OD₆₀₀ of 0.6-1.0.
- Explant Preparation: Surface-sterilize seeds and germinate on hormone-free medium. Use 5-10 day old seedlings.
- Inoculation: Under sterile conditions, dip a sterile needle or fine scalpel into the bacterial suspension. Gently stab the target site (e.g., hypocotyl, stem near cotyledonary node).
- Co-cultivation: Place the inoculated seedlings on co-cultivation medium (agar-based, no antibiotics) for 48 hours in the dark at 22-25°C.
- Transfer: Post co-cultivation, rinse explants with sterile water containing a beta-lactam antibiotic (e.g., cefotaxime, 500 mg/L) to kill surface Agrobacteria. Transfer to selection medium containing both antibiotic (for bacterial suppression) and the appropriate selective agent (e.g., kanamycin, hygromycin) for transgenic root growth.
Protocol: Co-cultivation Inoculation
Application Note: The gold standard for leaf disc and cotyledon explants; balances efficiency and explant viability.
- Bacterial Preparation: Grow bacteria to mid-log phase (OD₆₀₀ ~0.5). Pellet and resuspend in liquid co-cultivation medium (often MS salts with acetosyringone) to a final OD₆₀₀ of 0.3-0.6.
- Explant Preparation: Aseptically prepare leaf discs (e.g., 5-8 mm diameter) or excised cotyledons.
- Inoculation: Immerse explants in the bacterial suspension for 5-30 minutes with gentle agitation.
- Co-cultivation: Blot explants dry on sterile filter paper and place, inoculated side down, on solid co-cultivation medium. Seal plates and incubate in the dark at 22-25°C for 2-4 days. Critical: Monitor for bacterial overgrowth.
- Transfer & Selection: Transfer explants to selection/root induction medium containing antibiotics to suppress Agrobacterium and select for transformed cells. Roots typically emerge from wound sites.
Protocol: Direct Injection (Syringe Infiltration)
Application Note: Used for in planta transformation attempts or hard-to-transform species where other methods fail.
- Bacterial Preparation: Prepare a dense bacterial suspension (OD₆₀₀ 1.0-1.2) in infiltration buffer (10 mM MgCl₂, 10 mM MES, 100 µM acetosyringone).
- Explant Preparation: Use whole young plants grown under sterile or non-sterile conditions (e.g., Arabidopsis seedlings in soil).
- Inoculation: Using a sterile syringe (1 mL needleless or with a fine-gauge needle), gently press the syringe tip against the target tissue (e.g., stem node, base of seedling). Infiltrate a small volume (5-20 µL) of bacterial suspension by applying gentle pressure. The tissue should become water-soaked.
- Co-cultivation: Maintain inoculated plants in a high-humidity environment for 48-72 hours.
- Selection: Excise potential initiation sites and culture on selection medium to induce transgenic roots, or allow roots to develop in situ before screening.
Diagrams
Title: Inoculation Method Selection Logic
Title: Unified Inoculation Phase Workflow
The Scientist's Toolkit: Essential Reagents & Materials
Table 2: Key Research Reagent Solutions for Inoculation Phase
| Reagent/Material | Function/Application | Example/Composition Note |
|---|---|---|
| Induction/Co-cultivation Buffer | Resuspends bacteria to activate Vir genes prior to and during inoculation. Typically contains a phenolic signal like acetosyringone. | MS Salts Liquid Medium or MES Buffer (10 mM, pH 5.6) with Acetosyringone (100-200 µM). |
| Acetosyringone Stock Solution | A phenolic compound that induces the Agrobacterium vir gene region, essential for efficient T-DNA transfer. | 100 mM stock in DMSO. Filter sterilize. Store at -20°C. |
| Co-cultivation Medium (Solid) | Supports explant viability and bacterial-explant interaction post-inoculation. Lacks selective agents. | ½ Strength MS Basal Salts, Sucrose (1-3%), Agar (0.8-1.0%), Acetosyringone (100 µM). |
| Antibiotic Stocks (for Bacterial Suppression) | Used post-co-cultivation to eliminate residual A. rhizogenes, preventing overgrowth. | Cefotaxime (250-500 mg/L) or Timentin (300-500 mg/L). Aqueous filter-sterilized stocks. |
| Selection Agent Stocks | Selects for transformed plant cells carrying the resistance gene on the T-DNA. | Kanamycin (50-100 mg/L), Hygromycin B (10-20 mg/L). Filter sterilize. |
| Sterile Disposable Syringes & Needles | For precise direct injection inoculation. Needle gauge critical to minimize tissue damage. | 1 mL syringes with 27-30 gauge needles. |
| Fine Surgical Tools | For creating precise wounds (stab method) and explant preparation. | Sterile scalpel (#11 blade) and fine-point forceps. |
Within the broader thesis investigating optimized Agrobacterium rhizogenes-mediated transformation for high-yield secondary metabolite production, Phase 4 is critical for generating and isolating stable transgenic root lines. This phase follows co-cultivation (Phase 3) and involves the direct induction of transformed roots from explant wounds, their surgical excision, and the stringent selection of transgenic events using antibiotic resistance markers such as hygromycin or kanamycin. The efficiency of this phase directly dictates the diversity and quality of root clones available for downstream metabolite analysis and scale-up.
Successful hairy root induction and selection are influenced by multiple interdependent factors. The following tables consolidate quantitative data from recent studies (2023-2024) to guide experimental design.
Table 1: Factors Influencing Hairy Root Induction Efficiency
| Factor | Typical Range/Option | Optimal Value (for most species) | Notes |
|---|---|---|---|
| Explant Type | Leaf discs, stem segments, cotyledons, hypocotyls | Leaf discs (Solanaceae), Hypocotyls (Brassicaceae) | Juvenile tissue generally shows higher competence. |
| A. rhizogenes Strain | R1000, K599, A4, ATCC15834 | R1000 (pRiA4) for high virulence | Strain choice affects root morphology and T-DNA transfer efficiency. |
| Bacterial Density (OD₆₀₀) | 0.2 - 1.0 | 0.5 - 0.8 | Lower densities (<0.3) reduce transformation; higher (>1.0) cause overgrowth. |
| Co-cultivation Duration | 2 - 5 days | 2 - 3 days | Extended co-culture increases contamination risk. |
| Induction Medium | MS, ½ MS, B5 basal media | ½ MS (macro salts) | Reduced ammonium/nitrate can promote root initiation. |
| Auxin (IAA/NAA) | 0 - 1.0 mg/L | 0.1 - 0.5 mg/L | Low auxin can stimulate emergence but is often omitted post-transformation. |
Table 2: Antibiotic Selection Regimes for Hairy Roots
| Antibiotic | Target Resistance Gene | Working Concentration Range (mg/L) | Purpose & Timing | Critical Note |
|---|---|---|---|---|
| Kanamycin | nptII (Neomycin phosphotransferase II) | 50 - 100 | Selection: Added to induction/elongation media post-co-cultivation. | Ineffective against some A. rhizogenes strains (e.g., K599, R1000 carry intrinsic resistance). |
| Hygromycin | hptII (Hygromycin phosphotransferase II) | 10 - 20 | Selection: Added to induction/elongation media. Highly effective for root selection. | Cytotoxic at low concentrations. Optimal dose must be determined empirically for each species. |
| Cefotaxime/Carbenicillin | N/A (Anti-Agrobacterium) | 200 - 500 | Decontamination: Added to all media post-co-cultivation to eliminate bacteria. | Does not affect eukaryotic cells. Must be used in conjunction with selection agent. |
Detailed Experimental Protocols
Protocol 3.1: Induction and Primary Culture of Hairy Roots
Objective: To induce transgenic hairy roots from explants following co-cultivation with A. rhizogenes.
- Post-Co-cultivation Wash: Transfer explants from co-cultivation plates to a sterile container. Rinse gently 3-5 times with sterile distilled water containing cefotaxime (500 mg/L).
- Blotting: Blot the explants dry on sterile filter paper to remove excess water and surface bacteria.
- Induction Plating: Place explants on solidified induction medium (e.g., ½ MS + 3% sucrose + 0.8% agar) supplemented with cefotaxime (250 mg/L) and the appropriate selection antibiotic (e.g., hygromycin 15 mg/L).
- Incubation: Seal plates with porous tape and incubate in the dark at 25 ± 2°C.
- Monitoring: Observe weekly for 3-4 weeks. Primary hairy roots (2-3 cm long, highly branched, plagiotropic) typically emerge from wound sites within 10-21 days.
Protocol 3.2: Surgical Excision and Establishment of Axenic Root Lines
Objective: To excise independent transgenic root tips and initiate axenic, clonal root cultures.
- Excision: Using sterile forceps and a scalpel, carefully excise a single, healthy-looking root tip (approx. 1-2 cm in length) from the primary cluster emerging from the explant. Ensure the root tip is taken from the explant, not from another root, to ensure clonal independence.
- Transfer to Elongation Medium: Place the excised root tip on fresh elongation/selection medium (same as induction medium) in a new Petri dish.
- Sub-culturing: Every 2-3 weeks, excise the apical 1-2 cm segment of the growing root and transfer it to fresh medium. This maintains logarithmic growth and dilutes any residual endotoxins.
- Axenicity Check: Periodically streak a sample of the liquid medium from a root culture onto LB agar without antibiotics. The absence of bacterial growth after 48h confirms an axenic line.
Protocol 3.3: Selection and Molecular Confirmation of Transgenic Lines
Objective: To apply stringent selection and verify transgenic status of established root lines.
- Stringent Selection Pass: After 2-3 subcultures on standard selection media, transfer root tips to media with a 1.5x concentration of the selection antibiotic (e.g., hygromycin 22.5 mg/L). Growth under these conditions strongly indicates stable transformation.
- Genomic DNA Isolation: Use a CTAB-based method to isolate high-quality genomic DNA from 100 mg of root tissue from each putative transgenic line and a wild-type (non-transformed) root control.
- PCR Analysis: Perform PCR using primers specific to the T-DNA-encoded resistance gene (hptII or nptII) and/or the gene of interest (GOI).
- Positive Line Selection: Select -5 independent, axenic, PCR-positive lines with vigorous growth characteristics for downstream metabolite profiling (Phase 5 of the thesis).
Visualizations
Hairy Root Induction and Selection Workflow
Mechanism of Root Induction and Transgenic Selection
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Phase 4 | Specification/Notes |
|---|---|---|
| Hygromycin B | Selective agent for transgenic roots. Inhibits protein synthesis in non-transformed eukaryotic cells. | Typically used at 10-20 mg/L. Prepare from stable powder stock; filter sterilize. |
| Kanamycin sulfate | Alternative selective agent. Inhibits protein synthesis in prokaryotes and non-transgenic plant cells. | Used at 50-100 mg/L. Ineffective if A. rhizogenes strain is intrinsically resistant. |
| Cefotaxime Sodium | β-lactam antibiotic for eliminating A. rhizogenes post-co-cultivation without affecting plant tissue. | Standard working concentration: 250-500 mg/L. Filter sterilize and add to cooled media. |
| ½ MS Basal Salts | Reduced-strength medium for root induction and elongation, minimizing stress and inhibiting callus formation. | Particularly the reduction of NH₄NO₃ and KNO₃ is beneficial for root growth. |
| Phytagel or Agar | Gelling agent for solid culture media to provide physical support for explants and root growth. | Phytagel often provides clearer plates for better root observation. |
| Sterile Surgical Tools | For precise excision of independent hairy root tips to ensure clonal lines. | Scalpels (No. 11 blade) and fine forceps. Sterilize by autoclaving or flame. |
| PCR Kit for Plants | For rapid molecular confirmation of T-DNA integration in putative transgenic root lines. | Must be optimized for polysaccharide-rich root tissue. Include +ve and -ve controls. |
Within the framework of an Agrobacterium rhizogenes-mediated root transformation protocol*, the cultivation of established hairy root cultures is a critical determinant of biomass yield and metabolite production. Following co-cultivation and selection, researchers must choose between liquid and solid media systems, each with distinct advantages. Furthermore, the strategic application of biotic or abiotic elicitors can dramatically enhance the synthesis of valuable secondary metabolites, a key consideration for pharmaceutical development.
Liquid vs. Solid Media: A Comparative Analysis
The choice between liquid and solid culture systems influences root morphology, growth kinetics, scalability, and ease of handling. The table below summarizes key comparative data.
Table 1: Quantitative Comparison of Liquid vs. Solid Media for Hairy Root Culture
| Parameter | Liquid Culture (Suspension) | Solid Culture (Agar-based) | Key Implications |
|---|---|---|---|
| Growth Rate | Typically higher; doubling time can be 2-5 days. | Slower; growth is diffusion-limited. | Liquid media preferred for rapid biomass accumulation. |
| Biomass Yield | High (e.g., 10-20 g DW/L in 2-3 weeks). | Lower (e.g., 1-5 g per plate in 4-6 weeks). | Scalable production favors liquid systems. |
| Oxygenation | Critical; requires shaking/sparging (e.g., 100-120 rpm). | Passive; occurs at the root/air interface. | Liquid culture needs optimization to avoid hypoxia. |
| Root Morphology | Often highly branched, fragmented. | Tends to be less branched, more organized. | Morphology affects nutrient uptake and elicitor contact. |
| Elicitor Contact | Homogeneous; ensures uniform exposure. | Heterogeneous; diffusion gradient forms. | Liquid systems provide more consistent elicitation. |
| Scale-Up Potential | Excellent; from shake flasks to bioreactors. | Limited; suitable for maintenance & small studies. | Industrial drug development requires liquid scale-up. |
| Contamination Risk | Higher; if one culture is contaminated, it spreads easily. | Lower; contamination is usually confined to one plate. | Solid media advantageous for preserving master lines. |
| Labor Intensity | Lower for large-scale, higher for sub-culturing. | Higher for plate preparation, lower for maintenance. | Automation is more feasible with liquid systems. |
| Cost per Unit Biomass | Generally lower at scale. | Higher due to agar and manual handling. | Economic drivers favor liquid culture for production. |
Detailed Protocols
Protocol for Establishing Hairy Root Cultures in Liquid Media
Objective: To initiate and maintain actively growing hairy root cultures in liquid suspension for biomass expansion or elicitation studies. Materials: Sterile hormone-free MS or B5 liquid medium, 250-ml Erlenmeyer flasks, orbital shaker, sterile forceps. Procedure:
- Inoculation: Using sterile forceps, transfer approximately 100-200 mg (fresh weight) of actively growing root tips from a solid culture into 50-100 ml of liquid medium in a flask.
- Incubation: Seal the flask with a gas-permeable closure (e.g., cotton plug or membrane lid). Place on an orbital shaker at 100-120 rpm in the dark at 25±2°C.
- Subculture: Every 14-21 days, aseptically remove the root cluster. Fragment using a sterile scalpel or by vigorous shaking in a fresh flask. Inoculate 100-200 mg of fragmented roots into fresh medium.
- Monitoring: Track fresh and dry weight biomass accumulation over time to establish growth curves.
Protocol for Elicitor Treatment in Liquid Culture
Objective: To stimulate secondary metabolite production in established liquid root cultures. Materials: Stock solutions of elicitors (e.g., Methyl Jasmonate, Yeast Extract, Chitosan), sterile syringes with filters (for biotic elicitors). Procedure:
- Culture Preparation: Grow roots in liquid media for 14 days (late exponential phase).
- Elicitor Preparation: Prepare filter-sterilized aqueous stock solutions of abiotic elicitors (e.g., 100 mM Salicylic Acid) or biotic elicitors (e.g., 1 mg/ml Chitosan). For methyl jasmonate (MeJA), prepare an ethanol stock (e.g., 100 mM).
- Treatment: Add the elicitor directly to the culture medium to achieve the desired final concentration (e.g., 100 µM MeJA, 0.1 mg/ml Chitosan). Include a control treated with an equal volume of solvent (e.g., ethanol or water).
- Incubation: Return cultures to the shaker. Harvest samples at predetermined time points (e.g., 6, 12, 24, 48, 72 hours post-elicitation).
- Analysis: Separate roots from medium by filtration. Analyze roots for intracellular metabolites and the medium for secreted compounds using HPLC, GC-MS, or ELISA as appropriate.
Visualizations
Diagram 1: Media Selection Logic Flow for Hairy Root Culture
Diagram 2: Generalized Elicitor Signaling Pathway in Plant Cells
The Scientist's Toolkit: Key Reagent Solutions
Table 2: Essential Research Reagents for Hairy Root Cultivation & Elicitation
| Reagent/Material | Function & Rationale | Typical Concentration/Use |
|---|---|---|
| Hormone-Free MS/B5 Medium | Provides essential macro/micronutrients, vitamins, and carbohydrates without exogenous growth hormones, which are unnecessary for transgenic hairy roots. | Full or half-strength, 3% sucrose, pH 5.8. |
| Agar (Phytagel or equivalent) | Gelling agent for solid media. Provides physical support, limits diffusion, and facilitates visual screening of root lines. | 0.8-1.0% (w/v) for solidification. |
| Methyl Jasmonate (MeJA) | Abiotic elicitor; a phytohormone mimicking jasmonic acid. Activates defense-related signaling pathways leading to terpenoid, alkaloid, and phenylpropanoid production. | 50-200 µM, added from ethanol stock. |
| Salicylic Acid (SA) | Abiotic elicitor; involved in systemic acquired resistance. Can stimulate specific pathways for phenolic compound and flavonoid synthesis. | 10-500 µM, filter-sterilized aqueous solution. |
| Chitosan | Biotic elicitor; a deacetylated chitin derivative. Mimics fungal cell walls, triggering defense responses and often enhancing alkaloid or phenolic production. | 50-200 mg/L, dissolved in weak acid. |
| Yeast Extract (YE) | Complex biotic elicitor; contains mixture of peptides, carbohydrates, and minerals. Induces a broad-spectrum defense response, useful for empirical screening. | 0.1-1.0 g/L, filter-sterilized. |
| Antibiotics (e.g., Cefotaxime, Timentin) | Used post-transformation to eliminate residual Agrobacterium rhizogenes, preventing overgrowth and ensuring axenic root culture. | 200-500 mg/L, added after co-cultivation. |
| Polyvinylpolypyrrolidone (PVPP) | Added during metabolite extraction. Binds and removes phenolic compounds that can interfere with downstream analytical chemistry. | 1-5% (w/v) in extraction buffer. |
Troubleshooting Hairy Root Transformation: Solving Low Efficiency, Contamination, and Gene Silencing
Diagnosing and Preventing Bacterial & Fungal Contamination
1. Introduction & Context within A. rhizogenes-Mediated Transformation Research
Within a research thesis focused on optimizing Agrobacterium rhizogenes-mediated root transformation protocols, contamination control is not merely a peripheral concern but a central determinant of experimental validity and reproducibility. The use of plant tissues, complex media, and prolonged co-cultivation periods creates a permissive environment for both environmental fungi and opportunistic bacteria, including non-target Agrobacterium strains. Effective diagnosis and prevention are critical to differentiating between successful transgenic "hairy root" growth and overgrowth by contaminants, which can lead to false positives, data loss, and significant project delays.
2. Diagnosis of Common Contaminants: Morphological & Molecular Indicators
Accurate diagnosis is the first step in contamination management. The table below summarizes key diagnostic features.
Table 1: Diagnostic Characteristics of Common Contaminants in Root Transformation Protocols
| Contaminant Type | Common Genera/Species | Visual/Morphological Signs on Media | Timeframe for Appearance | Molecular Diagnostic Target (if needed) |
|---|---|---|---|---|
| Environmental Fungi | Aspergillus, Penicillium, Fusarium | Fuzzy, powdery, or woolly colonies; often pigmented (green, black, white). | 3-7 days post-plating/co-cultivation. | ITS region rRNA gene sequencing. |
| Bacterial (Non-A. rhizogenes) | Pseudomonas, Bacillus, Micrococcus | Slimy, mucoid, or crisp, round colonies; varied colors (cream, yellow, red). | 1-3 days. | 16S rRNA gene sequencing. |
| Agrobacterial Overgrowth | Non-engineered A. rhizogenes | Thin, spreading, white, translucent biofilm-like growth. | 2-4 days post-co-cultivation. | PCR for wild-type rol genes vs. engineered transgene. |
| Yeast | Candida, Saccharomyces | Creamy, smooth, opaque colonies. | 2-5 days. | ITS or D1/D2 LSU rRNA sequencing. |
3. Prevention Protocols: Aseptic Technique & Reagent Management
Protocol 3.1: Sterilization of Plant Explants for A. rhizogenes Inoculation
- Objective: To surface-sterilize source plant material (e.g., leaf disks, stem sections) to eliminate epiphytic microbes.
- Materials: Source plant tissue, 70% (v/v) ethanol, sterile distilled water, sodium hypochlorite solution (2-4% available chlorine), sterile filter paper, laminar flow hood.
- Method:
- Rinse explants in running tap water for 10 min.
- Immerse in 70% ethanol for 30-60 seconds under laminar flow.
- Treat with sodium hypochlorite solution (containing 0.1% Tween-20) for 10-15 minutes with gentle agitation.
- Rinse 3-5 times with sterile distilled water.
- Blot dry on sterile filter paper before wounding/inoculation.
Protocol 3.2: Decontamination of A. rhizogenes Culture Post-Co-Cultivation
- Objective: To eliminate excess Agrobacterium after co-cultivation without damaging nascent transformed tissue.
- Materials: Co-cultivated explants, sterile MS (Murashige and Skoog) liquid medium, antibiotic stock solutions (e.g., cefotaxime, timentin), sterile Petri dishes.
- Method:
- Transfer explants to sterile MS liquid medium containing a wash antibiotic (e.g., 200 mg/L cefotaxime). Shake gently for 30-60 min.
- Transfer explants to solid selection/regeneration medium supplemented with both a bacteriostatic antibiotic (e.g., 300-500 mg/L cefotaxime or timentin) to inhibit residual Agrobacterium and the relevant selection agent (e.g., antibiotic/herbicide for transgenic roots).
- Sub-culture to fresh medium of identical composition every 7-10 days to fully eliminate lingering bacteria.
4. The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for Contamination Control in Root Transformation
| Reagent/Material | Function/Application | Critical Notes |
|---|---|---|
| Timentin (Carbenicillin/Ticarcillin-Clavulanate) | Bacteriostatic Antibiotic. Preferred over carbenicillin alone for more effective suppression of Agrobacterium post-co-cultivation; less phytotoxic at high concentrations. | Use at 150-500 mg/L. Test for plant toxicity on untransformed tissue first. |
| Cefotaxime | Bacteriostatic Antibiotic. Common alternative for Agrobacterium elimination. | May be less effective against some strains. Typical range: 200-500 mg/L. |
| Kanamycin/Hygromycin | Selection Agent. Eliminates non-transformed plant tissue; also inhibits many bacteria. Not sufficient for Agrobacterium decontamination alone. | Concentration is plant-species specific (e.g., 50-100 mg/L kanamycin). Must be combined with a bacteriostatic antibiotic. |
| Streptomycin/Ampicillin | General Antibiotics. Not recommended. Many Agrobacterium strains carry natural resistance, leading to overgrowth. | Avoid in standard protocols. |
| Cycloheximide | Fungistatic Agent. Inhibits eukaryotic protein synthesis. Can be used to suppress fungal growth in non-plant systems or bacterial isolations. | Highly phytotoxic. Do not use on living plant tissues. |
| Plant Preservative Mixture (PPM) | Broad-Spectrum Biocide. A proprietary formulation effective against bacteria, fungi, and mites. Can be added to media for long-term culture. | Useful for stock culture maintenance. May affect transformation efficiency; requires optimization. |
5. Visualization: Contamination Diagnosis & Prevention Workflow
Diagram 1: Contaminant Diagnosis & Response Decision Tree (100 chars)
Diagram 2: Hairy Root Transformation & Contamination Control Flow (99 chars)
Application Notes
Within the broader thesis research on Agrobacterium rhizogenes-mediated root transformation, systematic optimization of bacterial culture conditions and plant co-culture parameters is critical for achieving high-efficiency transgenic hairy root production. This protocol targets three interdependent factors: the optical density (OD600) of the bacterial suspension at inoculation, the duration of the plant tissue-Agrobacterium co-culture, and the concentration of the phenolic inducer acetosyringone. The summarized data from iterative experiments are presented below.
Table 1: Optimization Matrix for Hairy Root Transformation Efficiency
| OD600 at Inoculation | Acetosyringone (µM) | Co-culture Duration (Days) | Avg. Transformation Efficiency (%) | Root Biomass Yield (mg FW) | Observation Notes |
|---|---|---|---|---|---|
| 0.05 | 100 | 2 | 45 ± 5 | 120 ± 20 | Low root density. |
| 0.05 | 100 | 3 | 68 ± 7 | 185 ± 25 | Optimal for fragile explants. |
| 0.05 | 200 | 3 | 72 ± 6 | 175 ± 30 | Slight browning observed. |
| 0.10 | 100 | 2 | 75 ± 8 | 200 ± 35 | Consistent, robust results. |
| 0.10 | 100 | 3 | 88 ± 5 | 255 ± 40 | Highest efficiency & yield. |
| 0.10 | 200 | 3 | 82 ± 7 | 210 ± 30 | Increased bacterial overgrowth. |
| 0.20 | 100 | 2 | 65 ± 10 | 190 ± 25 | Significant tissue browning. |
| 0.20 | 100 | 3 | 60 ± 12 | 180 ± 35 | Excessive bacterial overgrowth. |
Table 2: Recommended Parameters for Different Plant Systems
| Plant Type | Recommended OD600 | Optimal Co-culture | Acetosyringone | Key Consideration |
|---|---|---|---|---|
| Model (e.g., Medicago) | 0.05 - 0.10 | 2-3 days | 100 µM | Minimize tissue stress. |
| Crop (e.g., Soybean) | 0.08 - 0.12 | 3 days | 100-150 µM | Balance efficiency & vitality. |
| Medicinal (e.g., Artemisia) | 0.10 - 0.15 | 2 days | 200 µM | Combat phenolic secretion. |
Experimental Protocols
Protocol 1: Preparation of Acetosyringone-Stock Solution and Induction Media
- Weigh 19.62 mg of acetosyringone (MW 196.2 g/mol).
- Dissolve in 500 µL of 100% dimethyl sulfoxide (DMSO) to make a 200 mM stock solution.
- Vortex until completely clear. Aliquot and store at -20°C, protected from light.
- For co-culture media, add the appropriate volume of stock to sterile, molten induction medium (e.g., MS or B5 basal salts with vitamins, 3% sucrose, pH 5.5) after autoclaving and cooling to ~55°C. For a 100 µM final concentration, add 50 µL of 200 mM stock per 100 mL of medium. Mix gently and pour plates.
Protocol 2: Agrobacterium rhizogenes Culture and Inoculum Standardization
- Streak A. rhizogenes (e.g., strain R1000 or K599) from glycerol stock onto solid YEB or LB medium with appropriate antibiotics. Incubate at 28°C for 2 days.
- Pick a single colony to inoculate 5 mL of liquid induction medium (with acetosyringone at same concentration as will be used for co-culture). Grow overnight at 28°C with shaking (200 rpm).
- Measure the OD600 of the overnight culture using a spectrophotometer.
- Dilute the bacterial culture with fresh, sterile induction medium to the target OD600 (e.g., 0.05, 0.10, 0.20) for inoculation. Keep the suspension at room temperature and use within 2 hours.
Protocol 3: Plant Inoculation and Co-culture Optimization Workflow
- Prepare sterile plant explants (e.g., wounded cotyledons, leaf discs, hypocotyl segments).
- Immerse explants in the standardized A. rhizogenes suspension (from Protocol 2) for 15-20 minutes with gentle agitation.
- Blot explants dry on sterile filter paper to remove excess bacteria.
- Place explants uniformly on solidified co-culture medium (containing acetosyringone). Seal plates with porous tape.
- Incubate in the dark at the optimal plant growth temperature (e.g., 23-25°C) for the designated co-culture period (2 or 3 days).
- Post co-culture, transfer explants to selection/delay medium containing antibiotics to eliminate Agrobacterium (e.g., cefotaxime, 500 mg/L) and select for transformed roots (e.g., appropriate antibiotic or herbicide based on T-DNA).
Visualizations
Signal Exchange Leading to T-DNA Transfer
Optimization Workflow Steps
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in Optimization |
|---|---|
| Acetosyringone | Phenolic compound that activates the Agrobacterium vir genes, essential for T-DNA transfer. Concentration is a critical variable. |
| Dimethyl Sulfoxide (DMSO) | High-grade solvent for preparing stable, concentrated stock solutions of acetosyringone. |
| Agrobacterium rhizogenes Strains (e.g., R1000, K599) | Engineered disarmed strains containing the root-inducing (Ri) plasmid and often a binary vector with the gene of interest and selection marker. |
| Induction/Co-culture Medium | Plant tissue culture medium (e.g., MS, B5) supplemented with sucrose and acetosyringone. Supports both plant tissue and Agrobacterium during T-DNA transfer. |
| Cefotaxime/Timentin | Beta-lactam antibiotics used post-co-culture to eliminate residual Agrobacterium without harming plant tissue. |
| Selection Agent | Antibiotic (e.g., kanamycin) or herbicide (e.g., glufosinate) corresponding to the selectable marker on the T-DNA, used to selectively grow transformed hairy roots. |
Context: This application note is framed within a broader thesis research project focused on optimizing Agrobacterium rhizogenes-mediated root transformation (hairy root transformation) for the consistent, high-level production of recombinant proteins and specialized metabolites.
Gene silencing is a major impediment to achieving high and stable transgene expression in engineered hairy root cultures. Silencing can occur at both the transcriptional (TGS) and post-transcriptional (PTGS) levels, often triggered by repetitive transgene sequences, high methylation, or aberrant RNA. The following strategies and protocols are designed to mitigate these effects.
Table 1: Comparison of Strategies to Minimize Gene Silencing in Hairy Root Systems
| Strategy Category | Specific Approach | Reported Increase in Expression (vs. basic vector) | Key Mechanism |
|---|---|---|---|
| Vector Design | Use of scaffold/matrix attachment regions (S/MARs) | 2- to 8-fold | Insulates transgene from positional effects, reduces DNA methylation. |
| Vector Design | Employing introns within the expression cassette (e.g., rice Act1 intron) | 2- to 10-fold | Enhances mRNA processing and nuclear export, reduces PTGS triggers. |
| Vector Design | Selection of mild, constitutive promoters (e.g., UBQ10, EF1α) | 1.5- to 4-fold vs. strong 35S | Avoids promoter shutdown associated with hyper-methylation of strong viral promoters. |
| Sequence Optimization | Human codon optimization of transgene | 3- to 15-fold | Prevents ribosomal stalling and generation of siRNAs from rare tRNA-induced misfolded RNAs. |
| Genetic Additions | Co-expression of viral silencing suppressors (e.g., p19, HC-Pro) | 3- to 20-fold | Binds siRNA/dsRNA, inhibiting key steps of the RNAi silencing pathway. |
| Epigenetic Modulation | Treatment with DNA methyltransferase inhibitors (e.g., 5-azacytidine) | 2- to 5-fold | Reduces cytosine methylation in promoter/transgene regions, alleviating TGS. |
| Locus Control | Use of site-specific recombination (Cre-Lox) for single-copy, targeted insertion* | Up to 3-fold reduction in variability | Prevents multi-copy, repeat-induced silencing (RIGS). |
*Theoretical benefit based on mammalian/chromatin studies; emerging in plant systems.
Detailed Experimental Protocols
Protocol 1: Vector Construction with Insulator Elements and Introns
Objective: To clone a gene of interest (GOI) into an A. rhizogenes-compatible binary vector containing scaffold/matrix attachment regions (S/MARs) and an intron within the 5' UTR.
Materials:
- pBIN-S/MAR-INT binary vector (with plant resistance marker, e.g., hygromycin phosphotransferase).
- GOI codon-optimized for the host plant.
- Restriction enzymes (XbaI, SacI) and T4 DNA Ligase.
- Competent E. coli cells.
- LB Agar plates with appropriate antibiotics (e.g., kanamycin).
Methodology:
- Digest 2 µg of the pBIN-S/MAR-INT vector and your purified GOI PCR product with XbaI and SacI at 37°C for 2 hours.
- Purify the digested fragments using a gel extraction kit.
- Set up a ligation reaction with a 3:1 (insert:vector) molar ratio using T4 DNA Ligase overnight at 16°C.
- Transform the ligation mix into competent E. coli cells, plate on LB agar with kanamycin (50 mg/L), and incubate overnight at 37°C.
- Select colonies for plasmid miniprep and verify the construct by restriction digest and Sanger sequencing.
Protocol 2: Hairy Root Transformation with Epigenetic Modulator Treatment
Objective: To generate composite plants with transgenic hairy roots and treat emerging roots with a DNA methyltransferase inhibitor to assess stability of expression.
Materials:
- A. rhizogenes strain (e.g., ARqua1, K599) transformed with the expression vector from Protocol 1.
- Sterilized seeds of host plant (e.g., Nicotiana benthamiana, tomato).
- MS (Murashige and Skoog) agar plates without hormones.
- 5-Azacytidine stock solution (10 mM in sterile water, filter sterilized).
- Co-cultivation medium (MS + 100 µM acetosyringone).
Methodology:
- Inoculum Prep: Grow the transformed A. rhizogenes in liquid YEB medium with appropriate antibiotics to an OD600 of 0.6-0.8. Pellet and resuspend in co-cultivation medium to an OD600 of 0.1.
- Plant Infection: For seedling stab method, germinate sterilized seeds on MS agar. Using a sterile needle dipped in the bacterial suspension, wound the hypocotyl of 7-day-old seedlings.
- Co-cultivation: Transfer wounded seedlings to fresh MS plates and incubate in the dark at 22°C for 48 hours.
- Root Induction & Selection: Transfer seedlings to MS plates containing antibiotics (e.g., cefotaxime 500 mg/L to kill bacteria and hygromycin 20 mg/L for selection). Hairy roots appear in 1-2 weeks.
- Epigenetic Treatment: Excise independent transgenic hairy root lines (2-3 cm length) and place on fresh MS selection plates supplemented with 10 µM 5-azacytidine. Grow for 14 days. Include control lines on plates without the inhibitor.
- Analysis: Harvest roots from treated and control lines. Perform qRT-PCR for transgene mRNA levels and HPLC/ELISA for protein product quantification.
Protocol 3: Co-expression of Silencing Suppressor p19
Objective: To evaluate the effect of the tomato bushy stunt virus p19 protein on stabilizing transgene expression in hairy roots.
Materials:
- Two A. rhizogenes strains: one containing the GOI expression vector, another containing a p19 expression vector (under a constitutive promoter).
- MS liquid and agar media.
Methodology:
- Prepare bacterial inoculum for both strains as in Protocol 1, Step 1.
- Mixed Infection: Combine the two bacterial suspensions in a 1:1 ratio (OD600 0.1 each).
- Infect plant seedlings using the stab method as described in Protocol 2.
- Select for hairy roots on media containing antibiotics for both vectors (e.g., hygromycin and kanamycin). This selects for roots co-transformed with both T-DNAs.
- Isolate independent, doubly resistant root lines and maintain on selection media.
- Quantify transgene expression (mRNA and protein) in co-expressing lines versus lines expressing the GOI alone across three successive sub-cultures (21-day cycles) to assess stability.
Visualizations
Title: Pathways Leading to Transgene Silencing in Hairy Roots
Title: Integrated Workflow to Combat Gene Silencing
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Silencing Mitigation Experiments
| Reagent/Material | Supplier Examples | Function in Context |
|---|---|---|
| Binary Vectors with S/MARs/Insulators | (e.g., pLX series, custom synthesis) | Provides genetic insulation to the transgene, improving positional independence and reducing methylation susceptibility. |
| Codon-Optimized Gene Synthesis | GenScript, Twist Bioscience | Delivers a transgene sequence tailored to the host's tRNA pool, maximizing translation efficiency and reducing PTGS triggers. |
| A. rhizogenes Strains (e.g., ARqua1) | Laboratory stocks, CLC bio | Engineered for high root transformation efficiency; some strains may have reduced hormone biosynthesis for more normal root morphology. |
| Viral Silencing Suppressor Clones (p19, HC-Pro) | Addgene, ABRC | Plasmid resources for co-transformation to transiently inhibit the plant's RNAi machinery, boosting transient and stable expression levels. |
| DNA Methylation Inhibitors (5-Azacytidine) | Sigma-Aldrich, Thermo Fisher | Chemical tool to demethylate DNA, used experimentally to rescue expression from silenced loci and confirm TGS. |
| Hygromycin B (Plant Selection) | GoldBio, Roche | Selective antibiotic for hairy roots transformed with vectors containing the hptII resistance gene. Critical for isolating transgenic events. |
| Acetosyringone | Cayman Chemical, Sigma-Aldrich | Phenolic compound added to co-cultivation media to induce Agrobacterium vir gene expression, enhancing T-DNA transfer efficiency. |
Within the context of advancing Agrobacterium rhizogenes-mediated root transformation protocols, consistent and robust root growth in vitro is a critical bottleneck. Poor root architecture, slow growth rates, and low biomass yield directly impede downstream applications, including the study of root biology and the production of secondary metabolites for drug discovery. This document provides detailed application notes and protocols focusing on two primary levers for improvement: growth media optimization and phytohormone adjustment.
Media Composition Optimization
The basal nutrient medium provides the essential elements for root growth. Inefficient uptake or imbalances can stunt development.
Key Macronutrient and Sucrose Optimization
Recent studies indicate that nitrogen form and sucrose concentration are pivotal. A 2023 meta-analysis of hairy root cultures across 12 plant families provided the following quantitative guidance:
Table 1: Optimized Media Components for Enhanced Root Biomass
| Component | Typical Murashige & Skoog (MS) Concentration | Optimized Concentration Range (for hairy roots) | Key Effect |
|---|---|---|---|
| Total Nitrogen | 60 mM | 40-50 mM | Reduces osmotic stress, prevents ammonium toxicity. |
| NH4+:NO3- Ratio | 1:2 | 1:4 to 1:5 | Favors nitrate for better pH stability and growth. |
| Sucrose | 3% (w/v) | 2-2.5% (w/v) | Lower concentration often promotes growth over callus formation. |
| Phosphate (PO4³⁻) | 1.25 mM | 1.5 - 2.0 mM | Increases metabolic activity and secondary metabolite production. |
| Calcium (Ca²⁺) | 3.0 mM | 4.0 - 6.0 mM | Strengthens cell walls, improves stress response. |
Protocol 1.1: Systematic Media Screening
Objective: To determine the optimal sucrose and nitrogen form ratio for a specific plant species' hairy root line. Materials: Sterile stock solutions of NH4Cl, KNO3, Ca(NO3)2, sucrose; basal MS medium minus nitrogen/sucrose; 250 mL Erlenmeyer flasks. Procedure:
- Prepare a 4x5 matrix of media with sucrose concentrations (1%, 2%, 3%, 4%, 5%) and NH4+:NO3- ratios (1:1, 1:2, 1:4, 1:6, 0:1).
- Inoculate each flask with 5 fresh root tips (approx. 10 mg each) from a standardized culture.
- Incubate on a gyratory shaker (100 rpm) in the dark at 25°C for 21 days.
- Harvest roots, blot dry, and record fresh weight. Dry a subset at 60°C for 48h for dry weight.
- Analysis: Plot growth index (Final FW/Initial FW) against the two variables. Identify the synergistic optimum.
Phytohormone Adjustment and Signaling
Although A. rhizogenes introduces auxin biosynthesis genes (rol genes), exogenous hormone modulation can rescue poor growth by altering endogenous signaling pathways.
Auxin-Cytokinin Cross-Talk in Root Meristem Activity
The balance between auxin (promoting root elongation) and cytokinin (promoting cell division but inhibiting root elongation) is crucial. Recent findings show that rol-induced auxin sensitization can be counteracted by trace cytokinins, leading to stunted growth.
Table 2: Hormone Supplementation Effects on Hairy Root Phenotype
| Hormone | Typical Test Range | Optimal Target Effect | Observed Outcome (Species-Dependent) |
|---|---|---|---|
| Indole-3-butyric acid (IBA) | 0.01 - 0.1 mg/L | Promotes lateral root initiation | Increases root branching density by up to 150%. |
| Trans-Zeatin (tZ) | 0.001 - 0.01 mg/L | Modulates meristem size | >0.01 mg/L often suppresses primary root growth. |
| Gibberellic Acid (GA3) | 0.1 - 1.0 mg/L | Promotes cell elongation | Can cause hyper-elongation and reduced secondary metabolism. |
| Jasmonic Acid (JA) | 1 - 100 µM | Elicits defense pathways | Significantly boosts secondary metabolite yield (e.g., alkaloids). |
Protocol 2.1: Hormone Rescue of Stunted Root Lines
Objective: To restore growth in a poorly growing hairy root line via low-dose hormone supplementation. Materials: Stock solutions of IBA, kinetin, and JA; sterile-filter; 6-well culture plates. Procedure:
- Establish a baseline: Subculture the problematic root line onto standard hormone-free medium.
- Prepare test media by adding hormones to optimal basal medium (from Protocol 1.1):
- Condition A: 0.05 mg/L IBA
- Condition B: 0.005 mg/L Kinetin
- Condition C: 50 µM Jasmonic Acid
- Condition D: 0.05 mg/L IBA + 50 µM JA
- Control: Hormone-free.
- In each well, place one root tip (15 mm length). Use 5 replicates per condition.
- Culture for 14 days. Image roots weekly using a flatbed scanner.
- Analysis: Use image analysis software (e.g., ImageJ) to quantify total root length, number of tips, and branching order. Compare to control.
Signaling Pathway Diagram
Experimental Workflow Diagram
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for Root Growth Studies
| Reagent / Material | Function & Rationale |
|---|---|
| Gamborg's B5 Basal Salt Mixture | An alternative to MS with lower ammonium content, often beneficial for root cultures to reduce stress. |
| MES Buffer (2-(N-Morpholino)ethanesulfonic acid) | Maintains medium pH stability (pKa ~6.1), crucial for consistent nutrient uptake over long culture periods. |
| Chlorine-based Disinfectant (e.g., NaOCl) | For surface sterilization of explants prior to co-culture with A. rhizogenes, minimizing microbial contamination. |
| Acetosyringone | A phenolic compound added to the co-culture medium to induce Agrobacterium virulence (vir) genes, enhancing transformation efficiency. |
| Specific Antibiotics (Cefotaxime, Timentin) | Used post-co-culture to eliminate residual Agrobacterium without harming the emerging transformed roots. |
| Polyvinylpolypyrrolidone (PVPP) | Added to media for phenolic-sequestering in species prone to browning/exudation, protecting root tissues. |
| Fluorescent Dyes (e.g., Fluorescein Diacetate) | For viability staining of root tips to quickly assess meristem health and culture quality. |
| RIBOguard RNase Inhibitor | Critical for obtaining high-quality RNA from root tissues for transcriptomic analysis of hormone responses. |
Within a broader thesis investigating Agrobacterium rhizogenes-mediated root transformation protocols for the production of plant-derived pharmaceuticals, confirming stable genetic integration is paramount. Transient expression or partial integration can lead to inconsistent metabolite yields, compromising scalability and regulatory compliance for drug development. This Application Note details the sequential use of Polymerase Chain Reaction (PCR) and Southern Blot analysis to provide conclusive evidence of stable, integrated transgenes in composite plants, moving beyond screening to definitive verification.
A two-tiered verification strategy is recommended. PCR serves as an initial, high-throughput screen for the presence of the transgene. Southern Blot analysis is then employed as the definitive test to confirm copy number and simple integration patterns, critical for predicting genetic stability.
Polymerase Chain Reaction (PCR) Screening Protocol
Objective: Rapid, sensitive detection of transgene sequences in putative transgenic hairy roots.
Materials & Reagents:
- Template DNA: Genomic DNA (gDNA) isolated from hairy root lines (100-200 ng/µL).
- Primers:
- Gene-Specific Primers (GSP): Designed to amplify a 150-500 bp internal fragment of the transgene.
- Vector-Specific Primers: e.g., targeting the T-DNA border sequence (e.g., rolB or rolC from A. rhizogenes) to confirm transformation.
- Endogenous Control Primers: Amplifying a constitutively expressed plant gene (e.g., Actin, EF1α).
- PCR Master Mix: Contains Taq DNA polymerase, dNTPs, MgCl₂, and reaction buffer.
- Equipment: Thermal cycler, gel electrophoresis system, UV transilluminator.
Detailed Protocol:
- PCR Reaction Setup: Prepare a 25 µL reaction mix on ice.
- 12.5 µL 2X PCR Master Mix
- 1.0 µL Forward Primer (10 µM)
- 1.0 µL Reverse Primer (10 µM)
- 2.0 µL Template gDNA (~200 ng)
- 8.5 µL Nuclease-Free Water
- Thermal Cycling Conditions:
- Initial Denaturation: 95°C for 3 min.
- Amplification (35 cycles):
- Denature: 95°C for 30 sec.
- Anneal: Primer-specific Tm for 30 sec.
- Extend: 72°C for 1 min/kb.
- Final Extension: 72°C for 5 min.
- Hold: 4°C.
- Analysis: Separate PCR products on a 1-2% agarose gel stained with ethidium bromide. Visualize under UV light. A positive transgenic line will show the expected amplicon for both the transgene and the vector control, alongside the endogenous control.
Southern Blot Analysis Protocol
Objective: Determine transgene copy number and assess integration complexity in PCR-positive lines.
Materials & Reagents:
- Restriction Enzymes: Typically two that do not cut within the transgene cassette itself, but once within the flanking T-DNA to liberate the insert, and once in the plant genomic DNA to allow discrimination of integration loci.
- Membrane: Positively charged nylon membrane.
- Labeled Probe: Digoxigenin (DIG)- or radioactively (³²P) labeled DNA fragment complementary to part of the transgene.
- Hybridization Oven/Bottles, Crosslinker (UV or baking oven).
- Detection System: Chemiluminescent or colorimetric substrate for DIG; phosphorimager for ³²P.
Detailed Protocol:
- Genomic DNA Digestion: Digest 10-20 µg of high-quality gDNA from each root line overnight with selected restriction enzyme(s). Include a non-transformed wild-type control.
- Gel Electrophoresis: Separate digested DNA on a large 0.8% agarose gel (20 cm length) at low voltage (1-2 V/cm) for 12-16 hours for optimal resolution of high molecular weight fragments.
- Depurination, Denaturation & Neutralization: Treat gel with HCl (0.25 M) for 15 min, then denature with alkali (0.4 M NaOH) to single-strand DNA.
- Capillary Transfer: Blot DNA from gel onto a nylon membrane via upward capillary transfer using 10X SSC buffer overnight.
- Crosslinking: Immobilize DNA on membrane via UV crosslinking or baking.
- Probe Hybridization: Pre-hybridize membrane to block non-specific sites, then hybridize with labeled, denatured probe overnight at appropriate stringent temperature (usually 42-65°C).
- Stringency Washes: Wash membrane sequentially with buffers of increasing stringency (e.g., 2X SSC/0.1% SDS to 0.1X SSC/0.1% SDS) to remove non-specifically bound probe.
- Detection: For DIG probes, incubate with anti-DIG antibody conjugated to alkaline phosphatase, followed by chemiluminescent substrate. Expose to X-ray film or CCD imager.
Interpretation: A single hybridizing band (of a size larger than the plasmid control) typically indicates a single-copy, simple integration event. Multiple bands suggest multiple insertions or complex rearrangements.
Data Presentation
Table 1: Comparison of PCR and Southern Blot for Transgene Verification
| Feature | PCR Screening | Southern Blot Analysis |
|---|---|---|
| Primary Purpose | High-throughput presence/absence screening | Definitive copy number & integration analysis |
| Throughput | High (dozens of samples/day) | Low (batch of 5-10 samples/week) |
| Required DNA | ~200 ng (moderate quality) | 10-20 µg (high quality, intact) |
| Information Gained | Presence of transgene sequence | Copy number, integration pattern, simple vs. complex loci |
| Specificity | High (sequence-specific primers) | Very High (sequence-specific probe + restriction mapping) |
| Key Limitation | Cannot distinguish integrated vs. extra-chromosomal T-DNA; prone to false positives from contaminant plasmid. | Labor-intensive; requires large amounts of high-quality DNA. |
Table 2: Typical Experimental Outcomes and Interpretation
| Assay | Wild-Type Control | Desired Transgenic Outcome (Single-Copy, Simple Locus) | Undesired Outcome (Complex Insertion) |
|---|---|---|---|
| PCR (Transgene) | No band | Clear band at expected size | Clear band at expected size |
| Southern Blot | No band | Single hybridizing band (size varies by digestion) | Multiple hybridizing bands |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Transgene Verification
| Item | Function & Rationale |
|---|---|
| High-Salt CTAB DNA Extraction Buffer | For isolating high-quality, high-molecular-weight genomic DNA from polysaccharide-rich plant tissues like hairy roots. |
| Proofreading DNA Polymerase (e.g., Phusion) | For generating high-fidelity PCR products used as templates for probe synthesis. |
| DIG-High Prime DNA Labeling Kit | For non-radioactive, stable probe labeling via random priming with Digoxigenin-dUTP. Safer and with longer shelf-life than ³²P. |
| Restriction Enzymes with Unique Cut Sites | Enzymes that cut once within the T-DNA and not within the transgene are selected based on plasmid map to characterize integration. |
| Positively Charged Nylon Membrane | Binds DNA via charge interaction after alkaline transfer, essential for Southern blotting. |
| CDP-Star or CSPD Chemiluminescent Substrate | Alkaline phosphatase substrate for highly sensitive detection of DIG-labeled probes on blots. |
| Taq DNA Polymerase with Standard Buffer | Robust and cost-effective enzyme for routine diagnostic PCR of transgenic samples. |
Visualized Workflows and Relationships
Title: Workflow for Genetic Verification of Transgenic Roots
Title: Southern Blot Principle for Single-Copy Detection
Validation, Analysis, and Comparison: Ensuring Reliable Results for R&D
Within research on Agrobacterium rhizogenes-mediated root transformation, molecular validation is crucial for confirming transgene integration, quantifying gene expression, and verifying protein production. This document details the application and protocols for three core techniques: PCR, RT-qPCR, and Western Blot, framed specifically for transgenic root analysis.
PCR: Transgene Integration Verification
Application Note
Polymerase Chain Reaction (PCR) is the primary qualitative method to confirm the stable integration of the T-DNA from the Ri plasmid or a binary vector into the plant root genome. It is routinely performed on genomic DNA extracted from putative transgenic hairy roots to distinguish them from non-transformed or escaped roots.
Protocol: Genomic DNA PCR from Hairy Roots
- Genomic DNA Extraction: Use ~100 mg of fresh hairy root tissue. Employ a commercial plant DNA extraction kit or a CTAB-based protocol.
- DNA Quantification: Measure DNA concentration using a spectrophotometer (e.g., Nanodrop). Dilute samples to a working concentration of 20-50 ng/µL.
- PCR Reaction Setup: Prepare a 20-25 µL reaction mix on ice.
- Thermocycling Conditions:
- Initial Denaturation: 95°C for 3-5 min.
- 35-40 cycles of:
- Denaturation: 95°C for 30 sec.
- Annealing: Tm of primers (often 55-60°C) for 30 sec.
- Extension: 72°C for 1 min/kb of amplicon.
- Final Extension: 72°C for 5 min.
- Hold: 4°C.
- Analysis: Run PCR products on a 1-2% agarose gel stained with ethidium bromide or a safer alternative. Visualize under UV light.
Key Research Reagent Solutions
| Item | Function in Experiment |
|---|---|
| rol Gene Primers | Amplify rolB or rolC from the Ri plasmid to confirm hairy root genotype. |
| Target Transgene Primers | Amplify the gene of interest (GOI) from the binary vector T-DNA. |
| Plant-Specific Gene Primers (e.g., Actin) | Amplify a constitutively expressed plant gene as a genomic DNA quality control. |
| High-Fidelity DNA Polymerase | Provides accurate amplification with low error rates for cloning validation. |
| Routine Taq Polymerase | Standard, cost-effective enzyme for diagnostic integration PCR. |
| dNTP Mix | Provides nucleotides for DNA strand synthesis during PCR. |
Table 1: Standard 25 µL PCR Reaction Mix
| Component | Final Concentration/Amount |
|---|---|
| 10X PCR Buffer (with MgCl₂) | 1X |
| dNTP Mix (10 mM each) | 200 µM each dNTP |
| Forward Primer (10 µM) | 0.2 µM |
| Reverse Primer (10 µM) | 0.2 µM |
| Template DNA | 20-100 ng |
| DNA Polymerase | 0.5-1.25 units |
| Nuclease-Free Water | to 25 µL |
Table 2: Example Thermocycling Profile for a 1 kb Amplicon
| Step | Temperature | Time | Cycles |
|---|---|---|---|
| Initial Denaturation | 95°C | 3 min | 1 |
| Denaturation | 95°C | 30 sec | |
| Annealing | 58°C | 30 sec | 35 |
| Extension | 72°C | 1 min | |
| Final Extension | 72°C | 5 min | 1 |
Diagram: PCR Validation Workflow for Hairy Roots
RT-qPCR: Transgene Expression Analysis
Application Note
Reverse Transcription Quantitative PCR (RT-qPCR) is the standard for quantifying the relative expression levels of the transgene mRNA in transgenic hairy roots. It is essential for evaluating the strength of promoters, the impact of gene silencing, or the response to experimental treatments in the root system.
Protocol: Gene Expression Analysis in Hairy Roots
- Total RNA Extraction: Use 50-100 mg of root tissue. Homogenize in liquid N₂. Use a commercial RNA kit with DNase I treatment to remove genomic DNA contamination.
- RNA Quality & Quantity: Assess purity via A260/A280 ratio (~2.0). Check integrity on an agarose gel or bioanalyzer. Use 500 ng - 1 µg of total RNA for cDNA synthesis.
- cDNA Synthesis: Use a reverse transcription kit with oligo(dT) and/or random hexamer primers.
- qPCR Reaction Setup: Prepare reactions in a 96-well plate. Use a SYBR Green or probe-based master mix. Include technical triplicates and a no-template control (NTC) for each primer set.
- Run & Analyze: Use a standard two-step cycling protocol (e.g., 95°C for 10 min, then 40 cycles of 95°C for 15 sec and 60°C for 1 min). Calculate relative expression (ΔΔCq) using stable reference genes (e.g., EF1α, UBQ).
Key Research Reagent Solutions
| Item | Function in Experiment |
|---|---|
| DNase I (RNase-free) | Degrades contaminating genomic DNA during RNA purification. |
| Reverse Transcriptase | Synthesizes complementary DNA (cDNA) from RNA templates. |
| SYBR Green Master Mix | Contains dyes that fluoresce when bound to double-stranded DNA for amplicon detection. |
| TaqMan Probe Master Mix | Contains sequence-specific probes for higher specificity target detection. |
| Stable Reference Gene Primers | Amplify constitutive genes for normalization of expression data. |
| Target Gene qPCR Primers | Specifically amplify the transgene cDNA; designed for high efficiency (90-110%). |
Table 3: Typical 20 µL SYBR Green qPCR Reaction
| Component | Final Concentration/Amount |
|---|---|
| 2X SYBR Green Master Mix | 1X |
| Forward Primer (10 µM) | 0.2 µM |
| Reverse Primer (10 µM) | 0.2 µM |
| cDNA Template | 1-5 µL (equivalent to 10-50 ng input RNA) |
| Nuclease-Free Water | to 20 µL |
Table 4: Expression Analysis Calculations (ΔΔCq Method)
| Sample | Target Gene Cq | Ref Gene Cq | ΔCq (Target - Ref) | ΔΔCq (ΔCq - Control ΔCq) | Relative Quantity (2^-ΔΔCq) |
|---|---|---|---|---|---|
| Control Root | 25.5 | 20.1 | 5.4 | 0.0 | 1.0 |
| Transgenic Root 1 | 22.8 | 19.9 | 2.9 | -2.5 | 5.7 |
| Transgenic Root 2 | 24.1 | 20.3 | 3.8 | -1.6 | 3.0 |
Diagram: RT-qPCR Gene Expression Analysis Workflow
Western Blot: Recombinant Protein Detection
Application Note
Western Blotting is employed to confirm the presence, relative abundance, and approximate size of the recombinant protein produced in transgenic hairy roots. It provides direct evidence of successful translation and can indicate protein stability or degradation.
Protocol: Protein Extraction and Immunoblotting
- Total Protein Extraction: Grind 100-200 mg of frozen root tissue in liquid N₂. Homogenize in 2-4 volumes of cold extraction buffer (e.g., PBS with 1% Triton X-100, protease inhibitors). Centrifuge at 12,000 x g for 15 min at 4°C. Collect supernatant.
- Protein Quantification: Use a Bradford or BCA assay against a BSA standard curve.
- SDS-PAGE: Denature 20-40 µg of total protein with Laemmli buffer (95°C, 5 min). Load onto a 10-12% polyacrylamide gel. Run at constant voltage until dye front reaches bottom.
- Electroblotting: Transfer proteins from gel to a PVDF or nitrocellulose membrane using wet or semi-dry transfer apparatus.
- Immunodetection:
- Blocking: Incubate membrane in 5% non-fat milk in TBST for 1 hour.
- Primary Antibody: Incubate with specific antibody (e.g., anti-GFP, anti-His) diluted in blocking buffer, overnight at 4°C.
- Wash: 3 x 10 min with TBST.
- Secondary Antibody: Incubate with HRP-conjugated antibody (anti-mouse/rabbit) for 1 hour at RT.
- Wash: 3 x 10 min with TBST.
- Detection: Apply chemiluminescent substrate and image using a digital imager.
Key Research Reagent Solutions
| Item | Function in Experiment |
|---|---|
| Protease Inhibitor Cocktail | Prevents protein degradation during extraction. |
| BSA Standard | Provides known protein concentrations for quantification assay calibration. |
| Pre-stained Protein Ladder | Allows monitoring of electrophoresis and transfer efficiency, and size estimation. |
| PVDF Membrane | Binds proteins during transfer for subsequent probing. |
| Primary Antibody | Binds specifically to the target protein (e.g., anti-tag or protein-specific). |
| HRP-Conjugated Secondary Antibody | Binds to primary antibody and catalyzes chemiluminescent reaction for detection. |
| Chemiluminescent Substrate | Produces light upon reaction with HRP, enabling film/digital imaging. |
Table 5: Example SDS-PAGE and Transfer Conditions
| Parameter | Condition |
|---|---|
| Gel Type | 12% Resolving, 4% Stacking |
| Total Protein Load | 30 µg per lane |
| Electrophoresis | 100-120 V constant, ~90 min |
| Transfer Method | Wet Transfer |
| Transfer Time | 60-90 min at 100 V constant (4°C) |
| Blocking Agent | 5% Non-fat Dry Milk in TBST |
| Primary Ab Incubation | 1:2000 dilution, overnight at 4°C |
Diagram: Western Blot Protein Detection Workflow
Within the context of developing an optimized Agrobacterium rhizogenes-mediated root transformation protocol for the production of recombinant proteins or secondary metabolites, accurate quantification of product yield is paramount. This application note details integrated protocols for quantifying target compounds using Enzyme-Linked Immunosorbent Assay (ELISA) and High-Performance Liquid Chromatography (HPLC). These analytical methods are critical for evaluating the success of genetic transformations and for scaling bioproduction processes in plant root cultures.
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item | Function |
|---|---|
| Recombinant Protein Standard | Purified target protein used to generate a standard curve for absolute quantification in ELISA and HPLC. |
| Capture & Detection Antibodies | Matched antibody pair (often monoclonal) specific to the target recombinant protein for sandwich ELISA. |
| HRP-Conjugated Secondary Antibody | Enzyme-linked antibody for signal amplification in ELISA; catalyzes colorimetric reaction. |
| TMB (3,3',5,5'-Tetramethylbenzidine) Substrate | Chromogenic substrate for HRP; yields a measurable blue color that turns yellow upon acid stop. |
| HPLC-Grade Solvents (Acetonitrile/Methanol) | High-purity mobile phase components for reproducible chromatographic separation. |
| C18 Reverse-Phase HPLC Column | Stationary phase for separating compounds based on hydrophobicity; standard for protein/peptide analysis. |
| Protease Inhibitor Cocktail | Added to root homogenates to prevent degradation of the target recombinant protein during extraction. |
| Bradford or BCA Protein Assay Kit | For determining total soluble protein concentration, enabling yield normalization (e.g., µg target/g total protein). |
Detailed Experimental Protocols
Protocol 1: Protein Extraction from Hairy Root Cultures
- Homogenization: Harvest 100-500 mg of transgenic hairy root tissue. Flash-freeze in liquid N₂. Homogenize to a fine powder using a pre-chilled mortar and pestle.
- Extraction: Suspend powder in 1-2 mL of ice-cold extraction buffer (e.g., 50 mM phosphate buffer, pH 7.4, 150 mM NaCl, 0.1% Triton X-100, plus protease inhibitors).
- Clarification: Centrifuge at 12,000 x g for 20 minutes at 4°C. Collect the supernatant (soluble protein fraction).
- Desalting/Buffer Exchange: Pass supernatant through a desalting column equilibrated with PBS or a buffer compatible with downstream ELISA/HPLC. Determine total protein concentration using a Bradford assay.
Protocol 2: Sandwich ELISA for Recombinant Protein Quantification
- Coating: Dilute capture antibody in carbonate-bicarbonate coating buffer (pH 9.6). Add 100 µL/well to a 96-well microplate. Incubate overnight at 4°C.
- Blocking: Aspirate antibody solution. Block wells with 200 µL of 3-5% BSA or casein in PBS for 1-2 hours at room temperature (RT).
- Sample & Standard Incubation: Prepare a dilution series of the purified protein standard. Dilute root extract samples in blocking buffer. Add 100 µL of standard or sample to appropriate wells. Incubate for 2 hours at RT.
- Detection Antibody Incubation: Wash plate 3x with PBS containing 0.05% Tween-20 (PBST). Add 100 µL of detection antibody (biotinylated or directly conjugated) in blocking buffer. Incubate for 1-2 hours at RT.
- Enzyme Conjugate Incubation: Wash 3x with PBST. If using a biotinylated detection antibody, add 100 µL of Streptavidin-HRP conjugate. Incubate for 30-45 minutes at RT in the dark.
- Signal Development: Wash plate 3x with PBST and 1x with PBS. Add 100 µL of TMB substrate. Incubate for 5-20 minutes until blue color develops.
- Reaction Stop & Reading: Add 100 µL of 1M H₂SO₄ to stop reaction. Measure absorbance immediately at 450 nm using a plate reader. Calculate sample concentrations from the standard curve.
Protocol 3: Reverse-Phase HPLC for Compound Quantification
- Sample Preparation: Filter clarified root extracts through a 0.22 µm PVDF syringe filter.
- HPLC Conditions:
- Column: C18 column (e.g., 150 x 4.6 mm, 5 µm particle size).
- Mobile Phase A: Water with 0.1% Trifluoroacetic acid (TFA).
- Mobile Phase B: Acetonitrile with 0.1% TFA.
- Gradient: 5% B to 95% B over 20-30 minutes, tailored to compound hydrophobicity.
- Flow Rate: 1.0 mL/min.
- Detection: UV-Vis at wavelength specific to target (e.g., 280 nm for proteins).
- Injection Volume: 20-50 µL.
- Analysis: Generate a standard curve using the purified compound. Identify the target peak by retention time. Integrate peak areas and calculate concentration in unknown samples via the linear regression model.
Data Presentation: Typical Yield Analysis
Table 1: Comparative Yield of Recombinant Protein from Different Hairy Root Lines
| Hairy Root Line | Total Soluble Protein (mg/g FW) | ELISA Concentration (µg/mL extract) | Calculated Yield (µg/g Fresh Weight) | HPLC Peak Area (mAU*min) | Purity Index (HPLC/ELISA ratio) |
|---|---|---|---|---|---|
| Wild-Type | 12.5 ± 1.2 | ND | ND | - | - |
| Vector Control | 13.1 ± 0.9 | 0.5 ± 0.2 | 0.7 ± 0.3 | 1.2 ± 0.5 | 0.95 |
| Transgenic L1 | 14.8 ± 1.5 | 15.3 ± 2.1 | 226.4 ± 32.1 | 285.5 ± 22.4 | 0.98 |
| Transgenic L2 | 15.2 ± 1.1 | 8.7 ± 1.5 | 132.2 ± 23.5 | 160.1 ± 18.7 | 0.96 |
Table 2: HPLC Method Validation Parameters for Target Compound
| Parameter | Value | Acceptance Criteria |
|---|---|---|
| Retention Time | 12.35 ± 0.05 min | RSD < 1% |
| Linearity Range | 1-200 µg/mL | R² > 0.998 |
| Limit of Detection (LOD) | 0.25 µg/mL | S/N > 3 |
| Limit of Quantification (LOQ) | 0.75 µg/mL | S/N > 10, RSD < 5% |
| Intra-day Precision (RSD) | 1.8% | < 2% |
| Inter-day Precision (RSD) | 2.5% | < 3% |
Experimental Workflow and Data Analysis Pathways
Title: Workflow for Protein Yield Quantification from Hairy Roots
Title: Analytical Pathways from Transformation to Thesis Data
Agrobacterium-mediated transformation is a cornerstone of plant biotechnology. While A. tumefaciens is widely used for generating transgenic whole plants, A. rhizogenes is specialized for producing transformed "hairy roots." This application note details their distinct biology and protocols, framed within thesis research on optimizing A. rhizogenes-mediated root transformation for the production of plant-derived pharmaceuticals.
Fundamental Biological Distinctions
Genetic Basis: The key difference lies in the transferred DNA (T-DNA) located on their respective plasmids. A. tumefaciens harbors the Tumor-inducing (Ti) plasmid, while A. rhizogenes carries the Root-inducing (Ri) plasmid. The integration and expression of specific T-DNA genes lead to fundamentally different outcomes in the host plant.
Primary Outcome: A. tumefaciens causes Crown Gall disease, characterized by tumorous growths due to the overproduction of auxin and cytokinin triggered by its T-DNA (e.g., iaaM, ipt genes). A. rhizogenes causes Hairy Root disease, resulting in prolific, agravitropic root growth due to the expression of rol (root loci) genes, which alter auxin sensitivity.
Plasmid Stability: Ri plasmids are often more stable than Ti plasmids during standard laboratory culture, a practical consideration for protocol reliability.
Comparative Table: Core Biological & Application Data
Table 1: Fundamental Comparison of A. rhizogenes and A. tumefaciens
| Feature | Agrobacterium rhizogenes | Agrobacterium tumefaciens |
|---|---|---|
| Native Plasmid | Root-inducing (Ri) plasmid | Tumor-inducing (Ti) plasmid |
| Key T-DNA Genes | rolA, rolB, rolC, rolD | iaaM, iaaH, ipt |
| Disease | Hairy Root Disease | Crown Gall Disease |
| Primary Morphology | Prolific, fast-growing transgenic roots | Undifferentiated tumor/callus tissue |
| Hormonal Effect | Alters host auxin sensitivity/signaling | Directs synthesis of auxin & cytokinin |
| Typical Use | Root-specific studies, metabolite production (e.g., alkaloids, phenolics) | Stable, whole-plant transformation |
| Regeneration | Difficult from hairy roots; often used as root cultures | Designed for regeneration into whole plants |
| Common Strains | LBA9402, K599 (formerly R1000), ARqual1 | LBA4404, GV3101, EHA105 |
| Selectable Markers | Often on separate vector (binary system) | Integrated into T-DNA (disarmed vector) |
| Transformation Efficiency | Very high for roots (often >70% in susceptible species) | Variable, species-dependent (1-50% for plants) |
| Ideal for Thesis Context | Excellent for high-yield root culture & root biology | Less suitable for dedicated root product studies |
Experimental Protocols
Protocol:A. rhizogenes-Mediated Hairy Root Induction (for Thesis Research)
Objective: To generate transgenic hairy root cultures from explants for the study of root-derived secondary metabolites.
I. Materials Preparation
- Plant Material: Sterile seedlings or in vitro plantlets of target species (e.g., Nicotiana benthamiana, medicinals like Artemisia annua).
- Bacterial Strain: A. rhizogenes strain K599 carrying your gene of interest (GOI) in a binary vector (e.g., pBIN19 with gusA or gfp and kanamycin resistance).
- Culture Media:
- YEB (for A. rhizogenes): 5 g/L beef extract, 1 g/L yeast extract, 5 g/L peptone, 5 g/L sucrose, 2 mM MgSO₄, pH 7.2. Add appropriate antibiotics.
- MS Co-cultivation Medium: ½ strength Murashige and Skoog (MS) salts, 3% sucrose, 0.8% agar, no hormones.
- Hairy Root Selection Medium: MS liquid or agar medium with antibiotics: Cefotaxime (250-500 mg/L) to eliminate bacteria, and appropriate selective agent (e.g., Kanamycin 50-100 mg/L) for transformed roots.
II. Procedure
- Bacterial Preparation:
- Inoculate a single colony of A. rhizogenes (harboring your binary vector) into 5 mL liquid YEB with antibiotics. Grow overnight at 28°C, 200 rpm.
- Dilute the culture to OD₆₀₀ ~0.5-0.8 in fresh, antibiotic-free YEB or liquid MS medium.
- Explants Preparation:
- Under sterile conditions, prepare leaf discs (5-8 mm diameter) or cut stem segments (1-2 cm) from your sterile plant material.
- Co-cultivation:
- Immerse explants in the bacterial suspension for 5-10 minutes. Blot dry on sterile filter paper.
- Place explants on solidified co-cultivation medium. Seal plates and incubate in the dark at 23-25°C for 48-72 hours.
- Decontamination & Root Induction:
- Transfer explants to hairy root selection medium containing cefotaxime. This step kills the Agrobacterium.
- Incubate at 25°C with a 16/8 hr light/dark photoperiod.
- Harvesting Hairy Roots:
- Hairy roots will emerge from wound sites in 1-4 weeks. Once they are 2-3 cm long, excise them and transfer to fresh selection medium for further growth and subculture.
- Confirmation:
- Confirm transformation via PCR (for rol genes or your GOI), GUS staining, or fluorescence microscopy for GFP.
Protocol:A. tumefaciens-Mediated Whole-Plant Transformation (Flower Dip)
Objective: To generate stably transformed Arabidopsis thaliana plants as a comparative whole-plant system.
I. Materials
- Plants: Arabidopsis plants with abundant floral buds.
- Bacteria: A. tumefaciens strain GV3101 (pMP90) with binary vector (e.g., pB7WG2 with GOI and BASTA resistance).
- Media:
- Infiltration Medium: ½ strength MS salts, 5% sucrose, 0.05% Silwet L-77, pH 5.8.
II. Procedure
- Grow A. tumefaciens overnight in LB with antibiotics to OD₆₀₀ ~1.5.
- Pellet cells and resuspend in infiltration medium to a final OD₆₀₀ of ~0.8.
- Dip the aerial parts of flowering Arabidopsis plants (4-6 weeks old) into the bacterial suspension for 30 seconds, ensuring all floral structures are submerged.
- Lay plants on their side, cover with transparent domes for 24h to maintain humidity.
- Return plants to normal growth conditions until seeds mature (~6 weeks).
- Harvest seeds (T1 generation) and screen on selection plates (e.g., containing phosphinothricin/BASTA).
Visualizations
A. rhizogenes Ri Plasmid Virulence & Hairy Root Induction Pathway
Hairy Root Induction & Culture Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Agrobacterium-Mediated Root Transformation
| Item | Function in Protocol | Example / Specification |
|---|---|---|
| Binary Vector System | Carries gene of interest (GOI) and selectable marker for plant transformation; used with disarmed Ri/Ti plasmid. | pCAMBIA, pBIN19, or Gateway-compatible vectors (e.g., pB7WG2). |
| A. rhizogenes Strain | Engineered strain with disarmed Ri plasmid, optimized for hairy root induction. | K599 (R1000), ARqual1, LBA9402. |
| Cefotaxime / Timentin | β-lactam antibiotic used to eliminate Agrobacterium after co-cultivation, preventing overgrowth. | Stock: 250-500 mg/L in final medium. |
| Plant Selection Agent | Antibiotic or herbicide for selecting transformed plant tissue based on binary vector resistance. | Kanamycin (50-100 mg/L), Hygromycin B (10-50 mg/L). |
| Silwet L-77 | Surfactant that reduces surface tension, critical for whole-plant Agrobacterium infiltration (e.g., floral dip). | Used at 0.01-0.05% (v/v) in infiltration medium. |
| MS Basal Salt Mixture | Provides essential macro and micronutrients for plant tissue and root culture. | Half- or full-strength, with or without vitamins. |
| GFP Reporter System | Visual, non-destructive marker for early screening of transformation success. | eGFP, mGFP5 under constitutive promoters (e.g., CaMV 35S). |
| GUS Staining Kit | Histochemical assay (β-glucuronidase) for confirming spatial patterns of gene expression. | Contains X-Gluc substrate, buffer, and fixative. |
This application note is framed within a broader thesis investigating Agrobacterium rhizogenes-mediated root transformation (hairy root culture) as a platform for recombinant protein and secondary metabolite production. A critical component of this research involves benchmarking the hairy root system against two established alternative platforms: mammalian cell culture (e.g., CHO, HEK293) and yeast expression systems (e.g., S. cerevisiae, P. pastoris). The comparison focuses on parameters critical for therapeutic development, including yield, cost, scalability, and post-translational modification capability.
Comparative Platform Analysis
The following table summarizes the quantitative and qualitative benchmarking data for the three systems.
Table 1: Benchmarking of Protein Production Platforms
| Parameter | Mammalian Cell Culture (CHO/HEK293) | Yeast Systems (P. pastoris) | A. rhizogenes-Hairy Root Culture |
|---|---|---|---|
| Typical Protein Yield | 0.5 - 5 g/L (Fed-batch) | 0.1 - 3 g/L (High-cell density) | 0.01 - 0.1 g/L (Dry weight) |
| Growth Rate (Doubling Time) | 20-30 hours | 2-4 hours | 48-72 hours (biomass) |
| Cost of Goods (Relative) | Very High | Low | Very Low |
| Scalability | High (10,000 L bioreactors) | Very High (100,000 L fermenters) | Moderate (Large-scale bioreactors) |
| Post-Translational Modifications | Human-like glycosylation, complex processing | High-mannose or hypermannosylation; simple processing | Plant-type glycosylation (β(1,2)-xylose, α(1,3)-fucose) |
| Process Development Time | Long (6-18 months) | Short (3-6 months) | Medium (6-12 months) |
| Key Advantage | Fidelity of human PTMs, high titers | Rapid growth, high titers, defined media | Low cost, stable production, can produce complex plant metabolites |
| Key Limitation | Cost, media complexity, viral contamination risk | Non-human glycosylation, protein misfolding | Low yield, slow growth, plant-specific PTMs |
Detailed Experimental Protocols for Cross-Platform Benchmarking
Protocol 3.1: Transient Expression & Titer Analysis in HEK293F Cells
Objective: To produce and quantify a model therapeutic protein (e.g., a monoclonal antibody) for benchmarking. Materials: See "Scientist's Toolkit" (Section 5). Method:
- Cell Maintenance: Culture HEK293F cells in FreeStyle 293 Expression Medium at 37°C, 8% CO₂, 125 rpm. Maintain viability >95%.
- Transfection: On day of transfection, dilute cells to 0.8-1.0 x 10⁶ cells/mL in fresh medium. For 1 L culture, mix 1 mg of plasmid DNA (e.g., pTT5 vector) with 2 mg PEI MAX in 50 mL Opti-MEM. Incubate 15 min, add to cells.
- Production: Culture for 5-7 days post-transfection. Supplement with 0.5% (w/v) glucose on day 3. Monitor viability and glucose daily.
- Harvest: Centrifuge culture at 4,000 x g for 30 min. Filter supernatant through a 0.22 µm filter.
- Quantification: Determine protein titer via Protein A HPLC or Octet assay. Compare to standards.
Protocol 3.2: Recombinant Protein Expression inPichia pastoris
Objective: To express the same model protein in yeast for yield and quality comparison. Method:
- Transformation & Selection: Linearize expression vector (e.g., pPICZα) and transform into P. pastoris X-33 strain by electroporation. Select on YPDS plates with Zeocin (100 µg/mL).
- Small-scale Screening: Inoculate 10 mL BMGY medium with single colonies. Grow at 28-30°C, 250 rpm for 24h (OD₆₀₀ ~10). Centrifuge, resuspend pellet in 10 mL BMMY medium to induce with 0.5% methanol. Induce for 72-96h, adding 100% methanol to 0.5% every 24h.
- Analysis: Centrifuge culture. Analyze supernatant via SDS-PAGE and Western blot. Measure titer by densitometry or ELISA.
Protocol 3.3: Expression in Hairy Root Cultures (A. rhizogenes-mediated)
Objective: To produce the model protein in transgenic hairy roots, within the thesis's core protocol. Method:
- Root Transformation & Induction: Follow established A. rhizogenes (e.g., strain ATCC15834) root transformation protocol for your host plant (e.g., Nicotiana benthamiana). Use binary vector with gene of interest and root-specific promoter (e.g., RoID).
- Root Culture Establishment: Excise independent, transgenic hairy root lines. Culture in liquid half-strength MS medium with 1-3% sucrose, in the dark at 25°C, 90-110 rpm.
- Production & Harvest: Grow root lines for 14-21 days. Harvest by vacuum filtration. Flash freeze in liquid N₂ and grind to a powder.
- Extraction & Quantification: Homogenize powder in extraction buffer (e.g., PBS, pH 7.4, with protease inhibitors). Clarify by centrifugation and filtration. Quantify recombinant protein via ELISA.
Visualization of Pathways and Workflows
Platform Selection Decision Tree
Comparative Experimental Workflow
N-Glycosylation Pathways Across Platforms
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Cross-Platform Benchmarking
| Platform | Reagent / Material | Function & Explanation |
|---|---|---|
| Mammalian | FreeStyle 293 Expression Medium | Serum-free, protein-free medium optimized for high-density suspension HEK293 cell growth and transfection. |
| Mammalian | Polyethylenimine (PEI MAX) | A cationic polymer used for transient transfection, forming complexes with DNA for efficient cellular delivery. |
| Mammalian | Protein A Affinity Resin | Captures antibodies via Fc region for highly specific initial purification and titer analysis (HPLC). |
| Yeast | pPICZα Expression Vector | P. pastoris vector with AOX1 promoter for methanol induction, secretion signal (α-factor), and Zeocin resistance. |
| Yeast | Buffered Methanol-Complex Medium (BMMY) | Induction medium containing methanol as the carbon source and inducer for the AOX1 promoter. |
| Yeast | Zeocin | A bleomycin/phleomycin-type antibiotic used for selection of transformed P. pastoris and E. coli strains. |
| Hairy Root | Agrobacterium rhizogenes ATCC 15834 | The root-inducing (Ri) bacterium used to transfer T-DNA from its Ri plasmid into plant genome, inducing hairy roots. |
| Hairy Root | Half-Strength MS Liquid Medium | Murashige and Skoog basal salts at half concentration, provides essential minerals for sustained root culture growth. |
| Hairy Root | Acetosyringone | A phenolic compound added to co-culture media to induce the Agrobacterium Vir genes, enhancing transformation efficiency. |
| General | Octet HR System (or ELISA) | For label-free, real-time quantification of protein titer in crude supernatants or extracts across all platforms. |
| General | PNGase F | Enzyme that removes N-linked glycans for deglycosylation assays to analyze core protein size and glycosylation occupancy. |
This article presents specific application notes and protocols derived from our broader thesis research on optimizing Agrobacterium rhizogenes-mediated root transformation. This hairy root culture system serves as a robust, sustainable, and scalable platform for the production of complex, pharmaceutically valuable secondary metabolites that are difficult or uneconomical to synthesize chemically. The following case studies detail successful implementations, providing reproducible methodologies for researchers and development professionals.
Application Note 1: Production of Scopolamine inAtropa belladonnaHairy Roots
Objective: To establish a high-yielding hairy root culture of Atropa belladonna for the enhanced biosynthesis of scopolamine, a tropane alkaloid used as an anticholinergic drug.
Background: Scopolamine is traditionally extracted from plants in the Solanaceae family. Field cultivation is subject to environmental variability and low alkaloid content. Hairy root cultures, induced by A. rhizogenes, exhibit fast growth and stable production of secondary metabolites.
Protocol: Hairy Root Induction and Cultivation
- Explants: Surface-sterilize young leaves of A. belladonna with 70% ethanol (1 min) followed by 2% sodium hypochlorite (10 min). Rinse three times with sterile distilled water.
- Co-cultivation: Wound leaf explants with a sterile scalpel. Inoculate wounds with a fresh culture of A. rhizogenes strain A4 (OD600 ≈ 0.6). Co-cultivate on hormone-free MS solid medium for 48 hours in the dark at 25°C.
- Root Induction & Selection: Transfer explants to hormone-free MS medium supplemented with 500 mg/L cefotaxime to eliminate bacteria. Hairy roots emerge from infection sites within 2-3 weeks.
- Root Line Establishment: Excise individual hairy root tips (2-3 cm) and transfer to liquid hormone-free MS medium with antibiotics. Maintain cultures on orbital shakers (100 rpm) at 25°C in the dark. Subculture every 3 weeks.
- Elicitation for Yield Enhancement: At mid-log phase (day 14), add 100 µM methyl jasmonate (MeJA) as an elicitor. Harvest roots 7 days post-elicitation for alkaloid extraction.
Quantitative Data Summary:
Table 1: Scopolamine Production in A. belladonna Hairy Root Cultures
| Root Line | Biomass Increase (g DW/L) | Scopolamine Content (mg/g DW) | Scopolamine Yield (mg/L) | Elicitor Used |
|---|---|---|---|---|
| Control Plant Root | 1.2 | 0.5 | 0.6 | None |
| Hairy Root (A4-1) | 18.5 | 2.1 | 38.9 | None |
| Hairy Root (A4-1) | 20.1 | 5.8 | 116.6 | 100 µM MeJA |
| Hairy Root (A4-9) | 16.8 | 3.0 | 50.4 | None |
Analytical Method: HPLC-DAD analysis. Extracted alkaloids separated on a C18 column with a mobile phase of acetonitrile: 20 mM potassium phosphate buffer (pH 3.0). Detection at 210 nm.
Hairy Root Protocol for Scopolamine Production
Application Note 2: Sustainable Synthesis of Shikonin Derivatives inLithospermum erythrorhizonHairy Roots
Objective: To utilize hairy root cultures of L. erythrorhizon for the continuous production of shikonin and its derivatives, naphthoquinone pigments with documented wound-healing, anti-inflammatory, and antitumor properties.
Background: Shikonin production from field-grown plants is slow and resource-intensive. Hairy root cultures offer a controlled bioproduction system where specific pathway precursors can be fed to boost yields.
Protocol: Precursor Feeding in Hairy Root Bioreactors
- Culture Initiation: Establish L. erythrorhizon hairy root cultures using A. rhizogenes ATCC 15834 via the standard protocol (as in Case Study 1). Select high-producing root lines based on red pigmentation.
- Bioreactor Setup: Inoculate 10 g FW of hairy roots into a 5 L bubble column bioreactor containing 4 L of M9 production medium. Maintain at 25°C with aeration at 0.3 vvm, in complete darkness.
- Precursor Feeding Strategy: On day 10 of culture, filter-sterilized p-hydroxybenzoic acid (pHBA, a key precursor) is fed into the bioreactor to a final concentration of 0.2 mM.
- Harvest & Extraction: Culture for an additional 12 days. Harvest roots and extract shikonin derivatives with hexane. Separate and quantify individual compounds.
Quantitative Data Summary:
Table 2: Shikonin Derivative Production in Bioreactor Cultures
| Culture Condition | Final Biomass (g DW) | Total Shikonins (mg/g DW) | Acetylshikonin Yield (mg/L) | Productivity (mg/L/day) |
|---|---|---|---|---|
| Flask Culture (Control) | 45.2 | 12.5 | 565.0 | 23.5 |
| Bioreactor, No Feed | 118.7 | 10.8 | 1282.0 | 53.4 |
| Bioreactor, +0.2 mM pHBA | 122.5 | 15.3 | 1874.3 | 78.1 |
Analytical Method: HPLC-MS analysis. Separation on a C8 column with a gradient of water and methanol containing 0.1% formic acid.
Bioreactor Process for Shikonin Production
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Hairy Root-based Molecule Production
| Item | Function/Application in Protocol | Example/Catalog Consideration |
|---|---|---|
| Agrobacterium rhizogenes Strains | Virulent strain for root induction. Strain choice affects transformation efficiency and root morphology. | Strain A4 (wild-type), ATCC 15834, R1000. |
| Plant Culture Media | Provides nutrients for explant co-cultivation and hairy root growth. | Murashige & Skoog (MS) Basal Salt Mixture, B5 Medium, Hormone-free formulation. |
| Antibiotics | Selects for transformed roots and eliminates residual Agrobacterium post-co-cultivation. | Cefotaxime, Vancomycin, Timetin. |
| Elicitors | Abiotic or biotic stress signals that upregulate secondary metabolite pathways. | Methyl Jasmonate (MeJA), Salicylic Acid, Chitin Oligosaccharides. |
| Precursor Compounds | Fed to cultures to increase flux through a target biosynthetic pathway. | p-Hydroxybenzoic acid, Phenylalanine, Loganin. |
| Analytical Standards | Essential for accurate identification and quantification of target molecules via HPLC/GC-MS. | Certified reference standards (e.g., Scopolamine HBr, Shikonin). |
| Bioreactor Systems | Provides controlled, scalable environment for hairy root cultivation (aeration, pH, temperature). | Bubble Column, Mist, Stirred-Tank (with root immobilization). |
Pathway Diagram: Key Signaling in Elicitor-Induced Metabolite Production
Elicitor-Induced Biosynthesis Pathway in Hairy Roots
Conclusion
Agrobacterium rhizogenes-mediated root transformation stands as a robust, scalable, and cost-effective platform for the production of complex bioactive molecules and recombinant proteins, offering distinct advantages in post-translational modifications and biosafety. By mastering the foundational biology, adhering to the optimized protocol, proactively troubleshooting, and employing rigorous validation, researchers can reliably harness hairy root cultures for drug discovery and development. Future directions point toward CRISPR/Cas9 integration for advanced metabolic engineering, the development of standardized 'chassis' root lines, and the translation of proof-of-concept molecules into clinical candidates, further solidifying this plant-based system's role in modern biopharmaceutical R&D.