Engineering Chemical Diversity: The Pivotal Role of 2,3-Oxidosqualene Cyclization in Triterpene Biosynthesis and Drug Discovery
This review explores the enzymatic conversion of 2,3-oxidosqualene into diverse triterpene scaffolds, a cornerstone of natural product biosynthesis.
Engineering Chemical Diversity: The Pivotal Role of 2,3-Oxidosqualene Cyclization in Triterpene Biosynthesis and Drug Discovery
Abstract
This review explores the enzymatic conversion of 2,3-oxidosqualene into diverse triterpene scaffolds, a cornerstone of natural product biosynthesis. We detail the foundational mechanisms of oxidosqualene cyclases (OSCs), the methodological approaches for studying and manipulating these pathways, common challenges in functional characterization and heterologous expression, and comparative analyses of OSC enzyme families across species. Targeting researchers and drug developers, this article synthesizes current knowledge to highlight how understanding and engineering this pivotal cyclization step can unlock novel bioactive compounds for therapeutic applications.
Unraveling the Cyclization Cascade: How 2,3-Oxidosqualene Fuels Triterpene Structural Diversity
This whitepaper details the central role of 2,3-oxidosqualene (OS) as the universal precursor in triterpene biosynthesis. Framed within a thesis on cyclization-driven diversity, we explore the enzymatic machinery that converts this single substrate into over 200,000 characterized triterpenoid scaffolds, underpinning drug discovery for cancer, infectious diseases, and metabolic disorders.
The thesis posits that the structural diversity of triterpenes is fundamentally dictated by the cyclization and subsequent rearrangement of the OS scaffold. This linear epoxy-terminated (C_{30}) isoprenoid is the obligate substrate for oxidosqualene cyclases (OSCs), a class of enzymes that catalyze the most complex cyclization reactions in nature, forming distinct tetra- or pentacyclic core structures.
The Cyclization Landscape: Pathways and Products
OSC-catalyzed reactions initiate with epoxide protonation, triggering a cascade of carbocationic cyclizations and rearrangements. The ultimate product is determined by the enzyme's active site topography, which governs the folding of the OS substrate and the sequence of Wagner-Meerwein shifts.
Table 1: Major OSC Product Classes and Their Fates
| Cyclization Product (Core) | Key Rearrangements | Representative End-Products | Biological Relevance |
|---|---|---|---|
| Protosteryl Cation | Deprotonation; Methyl/ Hydride Shifts | Lanosterol (animals, fungi); Cycloartenol (plants) | Essential membrane component (cholesterol precursor) |
| Dammarenyl Cation | Deprotonation; Backbone Rearrangements | Dammarene-diols, Ginsenosides | Adaptogens (phytomedicines) |
| β-Amyrin Cation | Limited Rearrangements | Oleanane-type Triterpenes (e.g., β-Amyrin) | Anti-inflammatory (e.g., glycyrrhizic acid) |
| α-Amyrin Cation | Limited Rearrangements | Ursane-type Triterpenes (e.g., α-Amyrin) | Anticancer, antimicrobial |
| Lupenyl Cation | Extensive Rearrangements | Lupane-type Triterpenes (e.g., Lupcol) | Anticancer, cholesterol-lowering |
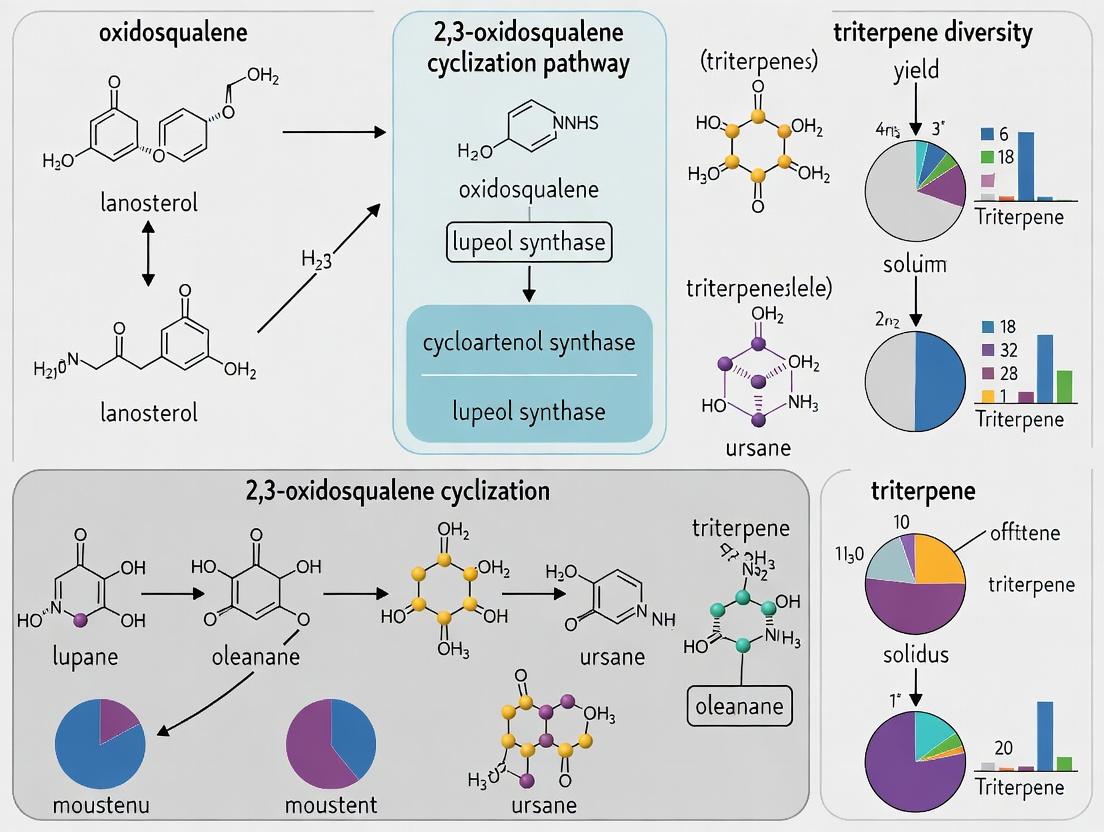
Diagram Title: Oxidosqualene Cyclization Pathways to Triterpene Scaffolds
Experimental Toolkit: Studying OSC Function
Table 2: Essential Research Reagent Solutions
| Reagent / Material | Function in OSC Research | Key Suppliers (Examples) |
|---|---|---|
| [3H]- or [14C]-Labeled 2,3-Oxidosqualene | Radiolabeled substrate for in vitro enzymatic assays to track cyclization products. | American Radiolabeled Chemicals, PerkinElmer |
| Recombinant OSC Enzymes (E. coli, Yeast, Insect Cell expressed) | Purified enzyme source for mechanistic and structural studies. | Custom cloning/expression; commercial cDNA from Addgene, ATCC. |
| Triterpene Standard Library (e.g., Lanosterol, β-Amyrin, Lupcol) | GC-MS/HPLC standards for product identification and quantification. | Sigma-Aldrich, Extrasynthese, ChromaDex |
| OSC Inhibitors (e.g., Ro 48-8071, U18666A) | Pharmacological tools to probe OSC function in cellular systems. | Tocris Bioscience, Cayman Chemical |
| Squalene Epoxidase Inhibitor (NB-598) | Upstream inhibitor to modulate endogenous OS levels in cells. | Cayman Chemical |
| Silica Gel / Normal Phase HPLC Columns | Critical for separating non-polar cyclization products. | Waters, Agilent, Sigma-Aldrich |
| GC-MS with Quadrupole or TOF detector | Primary instrument for identifying and quantifying triterpene hydrocarbons. | Agilent, Shimadzu |
| Crystallization Kits for Membrane Proteins | For obtaining diffractable crystals of OSC enzymes (often in detergent). | Hampton Research, Molecular Dimensions |
| Site-Directed Mutagenesis Kit | For probing active site residues and altering product specificity. | Agilent (QuikChange), NEB |
Key Experimental Protocols
In VitroOSC Enzyme Assay & Product Analysis
Objective: To characterize the catalytic activity and product profile of a purified OSC. Detailed Protocol:
- Reaction Setup: In a glass vial, combine:
- 50 mM Potassium phosphate buffer (pH 7.0), 100 µL
- 0.1% (w/v) Triton X-100 or CHAPS (detergent), 10 µL
- Purified recombinant OSC enzyme, 10-50 µg
- Substrate: 50 µM unlabeled or 0.1 µCi of [³H]-2,3-oxidosqualene (in toluene, evaporated under N₂ and redissolved in detergent).
- Final reaction volume: 200 µL with H₂O.
- Incubation: Incubate at 30°C for 30-120 minutes.
- Termination & Extraction: Stop reaction by adding 200 µL of 20% KOH in 90% ethanol. Saponify at 85°C for 30 min. Cool, add 400 µL H₂O, and extract triterpene products with 3 x 500 µL n-hexane or n-pentane. Pool organic phases and dry under N₂.
- Analysis:
- TLC: Redissolve in hexane, spot on silica gel TLC plate, develop in toluene:ethyl acetate (9:1). Visualize with anisaldehyde stain or phosphomolybdic acid.
- GC-MS: Derivatize dried product with BSTFA (N,O-bis(trimethylsilyl)trifluoroacetamide) at 70°C for 30 min. Inject into GC-MS (e.g., DB-5MS column). Identify products by comparing retention times and mass spectra to authentic standards.
Heterologous Expression & Mutagenesis of OSC in Yeast
Objective: To produce mutant OSC enzymes and analyze altered product outcomes in vivo. Detailed Protocol:
- Strain & Vector: Use Saccharomyces cerevisiae erg7Δ (lanosterol synthase knockout) strain harboring a plasmid for ergosterol complementation. Clone wild-type or mutant OSC gene into a galactose-inducible yeast expression vector (e.g., pYES2/CT).
- Transformation & Culture: Transform yeast strain using lithium acetate method. Select on SC-Ura glucose plates. Inoculate single colony into SC-Ura raffinose medium, grow to mid-log phase, induce with 2% galactose for 24-48h.
- Sterol/Triterpene Extraction: Harvest cells, lyse with glass beads in methanol. Add internal standard (e.g., cholestanol). Saponify with 60% KOH at 80°C for 1h. Extract neutral lipids with hexane.
- Product Profiling: Derivatize extract (BSTFA) and analyze by GC-MS as in 4.1. Compare chromatograms to identify novel peaks resulting from mutant OSC activity.
Diagram Title: Core Experimental Workflow for OSC Characterization
Quantitative Data & Structural Insights
Table 3: Kinetic Parameters of Select Characterized OSCs
| OSC Enzyme (Source) | Primary Product | (K_m) for OS (µM) | (k_{cat}) (min⁻¹) | (k{cat}/Km) (µM⁻¹ min⁻¹) | PDB ID (Example) |
|---|---|---|---|---|---|
| Human Lanosterol Synthase | Lanosterol | 4.2 ± 0.5 | 0.21 ± 0.02 | 0.05 | 1W6K |
| Arabidopsis thaliana Lupcol Synthase | Lupcol | 16.3 ± 2.1 | 2.8 ± 0.3 | 0.17 | 6N4G |
| Pisum sativum β-Amyrin Synthase | β-Amyrin | 11.5 ± 1.8 | 1.05 ± 0.1 | 0.09 | - |
| Alicyclobacillus acidocaldarius SHC | Hopene | 8.7 ± 0.9 | 480 ± 30 | 55.2 | 3SQC |
The central dogma of OS as the universal precursor is the foundation for engineered biosynthesis and selective inhibition. Understanding precise cyclization mechanisms enables:
- Rational engineering of OSCs to produce high-value triterpene scaffolds.
- Selective targeting of pathogen OSCs (e.g., in fungi, protozoa) with minimal host toxicity.
- Modulation of endogenous triterpenes in plants for improved nutraceutical profiles.
This field remains driven by integrating structural biology, enzyme mechanics, and metabolic engineering to harness the vast chemical diversity encoded in the OS cyclization reaction.
The cyclization of 2,3-oxidosqualene (OS) into over 200 distinct triterpene scaffolds represents a paradigm of enzymatic catalysis and carbocation chemistry in generating chemical diversity. This whitepaper focuses on the precise stereochemical template—the Chair-Boat-Chair (CBC) conformation—that pre-organizes the linear OS substrate within the oxidosqualene cyclase (OSC) active site. The binding in this specific conformation is the non-negotiable prerequisite for the initiation of the complex cationic cascade that leads to the protosterol cation and, subsequently, to diverse downstream triterpenes. Understanding this conformational control is central to the broader thesis on triterpene diversity, as it defines the initial fold from which all structural permutations evolve.
The Chair-Boat-Chair Conformational Template
The OS substrate (C30) must adopt a specific folded geometry to enable the seamless, processive ring-forming and migration steps. X-ray crystallography and computational studies of OSCs (e.g., human lanosterol synthase) confirm this pre-folded state.
- Chair (C1-C6): The first six carbons (post-epoxide) adopt a cyclohexane-like chair conformation, positioning C1-C2 for initial epoxide opening and forming ring A.
- Boat (C7-C12): The next six carbons adopt a boat conformation, setting the trajectory for the formation of rings B and C.
- Chair (C13-C18): The final pre-folded segment adopts another chair conformation, guiding the formation of ring D and the establishment of the tetracyclic core.
This C-B-C folding aligns the reacting π-bonds and cationic centers with precise stereoelectronic control.
Table 1: Key Structural Determinants of the CBC Conformation in Model OSCs
| OSC Enzyme (Source) | PDB ID | Key Active Site Residues Stabilizing CBC | Substrate Conformation (RMSD from ideal CBC) | Reference (Year) |
|---|---|---|---|---|
| Homo sapiens Lanosterol Synthase | 1W6K | Tyr503, His232, Phe444 (hydrophobic contour) | 0.5 Å | Thoma et al. (2004) |
| Alicyclobacillus acidocaldarius SHC | 3SQC | Tyr420, Trp232, Arg415 (cation-π, electrostatic) | 0.7 Å | Hoshino & Sato (2002) |
| Trypanosoma brucei Lanosterol Synthase | 6W7K | Tyr509, His234, Phe450 (conserved motif) | 0.6 Å | Lee et al. (2020) |
| Arabidopsis thaliana Lupeol Synthase (Model) | N/A | Trp257, Phe443, Ile729 (product outcome specific) | N/A (Computational) | Ito et al. (2011) |
Initiation of the Cationic Cascade from the CBC Template
The CBC conformation directly enables the stepwise cationic cascade:
- Initiation: Acidic protonation (often via an aspartate residue, e.g., D455 in human LAS) of the 2,3-epoxide oxygen.
- Ring A Formation: Epoxide opening generates a tertiary C3 carbocation. Anti-Markovnikov attack of the C6-C7 π-bond on C2 forms the A-ring (6-membered) and a tertiary C7 cation.
- Processive Cyclization: The cascade proceeds through a series of cation-π additions, hydride shifts, and methyl migrations, all channeled by the enzyme's active site geometry. The initial CBC fold ensures the correct stereochemistry at each new chiral center.
Table 2: Quantitative Metrics of Key Cationic Cascade Intermediates
| Intermediate Name | Chemical Formula | Theoretical m/z ([M+H]+) | Relative Energy (DFT, kcal/mol)* | Lifetime (Estimated) | Detection Method |
|---|---|---|---|---|---|
| Protosteryl Cation | C30H51+ | 411.399 | 0.0 (reference) | Femtoseconds | Computational, Analogue Trapping |
| Dammarenyl Cation | C30H51+ | 411.399 | +5.2 | Picoseconds | Enzyme Mutagenesis Trapping |
| Baccharenyl Cation | C30H51+ | 411.399 | +8.7 | Picoseconds | Isotopic Labeling Studies |
| Lupanyl Cation | C30H51+ | 411.399 | +12.1 | Not Observed | Product Analysis Inference |
*Representative DFT values from studies on truncated models; absolute values vary with method/basis set.
Experimental Protocols for Studying the CBC Initiation
Protocol 4.1: Site-Directed Mutagenesis andin vitroEnzyme Assay for Cascade Interruption
Objective: To trap cationic intermediates by disrupting the active site contour that stabilizes the CBC fold or subsequent rearrangements. Methodology:
- Design primers to mutate conserved aromatic/acidic residues (e.g., Tyr503 → Phe, Asp455 → Ala) in the OSC gene clone.
- Express wild-type and mutant enzymes in a heterologous system (e.g., E. coli or yeast).
- Purify recombinant enzymes via affinity chromatography (His-tag).
- Perform in vitro cyclase assay: Incubate purified enzyme (10-100 µg) with 50 µM [³H]- or [¹⁴C]-labeled 2,3-oxidosqualene substrate in assay buffer (pH 7.4, 25 mM Tris-HCl, 1 mM DTT, 0.1% CHAPS) at 30°C for 30 min.
- Extract reaction products with hexane/ethyl acetate (3:1).
- Analyze extracts by radio-HPLC or GC-MS. Mutants often produce aborted cyclization products (e.g., dammarenediol-II from a LAS mutant), which can be characterized by NMR.
Protocol 4.2: Computational Molecular Dynamics (MD) Simulation of Substrate Docking
Objective: To visualize the stabilization energy and dynamics of the CBC conformation within the active site. Methodology:
- Obtain OSC crystal structure (e.g., PDB: 1W6K). Prepare the protein (add hydrogens, assign charges using AMBER ff14SB force field).
- Generate a 3D structure of 2,3-oxidosqualene. Minimize its geometry using Gaussian (HF/6-31G*).
- Dock the substrate into the active site using induced-fit docking protocols (Schrödinger Suite or AutoDock).
- Solvate the protein-ligand complex in a TIP3P water box with 10 Å padding. Add ions to neutralize.
- Run equilibrium MD simulation (NAMD/AMBER) for 100-200 ns at 300K, 1 atm.
- Analyze trajectories for root-mean-square deviation (RMSD) of the substrate, key residue distances (e.g., epoxide O to catalytic Asp), and conformational stability of the CBC fold.
Visualization: The Conformational-Cascade Pathway
Diagram Title: OSC Catalytic Pathway from CBC Fold to Diversity
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function & Rationale | Example Product / Supplier |
|---|---|---|
| Radiolabeled [³H]-2,3-Oxidosqualene | High-sensitivity tracer for in vitro enzyme kinetics and product detection; enables quantification of picomolar product formation. | American Radiolabeled Chemicals, Inc. (ART-0112) |
| Recombinant OSC Expression System | Heterologous protein production; often E. coli with pET vectors or yeast (Pichia pastoris) for eukaryotic post-translational modifications. | Thermo Fisher Scientific (Champion pET vectors) |
| Affinity Chromatography Resin | Purification of His-tagged recombinant OSCs; critical for obtaining contaminant-free enzyme for mechanistic studies. | Cytiva (HisTrap HP nickel columns) |
| Mechanistic Probes (e.g., 2,3;22,23-Dioxidosqualene) | Substrate analogues designed to trap specific cationic intermediates or alter cascade progression. | Custom synthesis (e.g., Sigma-Aldrich Custom Synthesis) |
| Deuterated Quenching Reagents (D₂O, NaBD₄) | Used to trap cations as deuterated products, revealing hydride shift and termination steps via MS/NMR analysis. | Cambridge Isotope Laboratories (DLM-4-99) |
| Molecular Dynamics Software Suite | For simulating substrate binding, CBC stability, and cation migration pathways (e.g., AMBER, GROMACS, Schrödinger). | Schrödinger (Desmond), AMBER22 |
| Stable Isotope-Labeled Mevalonate ([¹³C₆]-MVA) | Metabolic precursor fed to living systems to produce isotopically labeled OS, enabling detailed NMR mapping of carbons through the cascade. | Isotec/Sigma-Aldrich (491638-1G) |
This technical guide examines the role of oxidosqualene cyclase (OSC) active site architecture as the primary determinant in triterpene scaffold diversification. Within the broader thesis of 2,3-oxidosqualene cyclization research, we detail how precise active site contours, amino acid positioning, and conformational dynamics direct cationic cyclization cascades to yield distinct tetra- and pentacyclic products. This document provides updated structural data, experimental protocols for functional analysis, and a toolkit for researchers probing this foundational biosynthetic node.
The cyclization of 2,3-oxidosqualene (OS) represents a critical branch point in isoprenoid biosynthesis, leading to over 100 distinct carbon skeletons. This reaction is catalyzed by OSCs, which function as precise architects, orchestrating stereoselective ring formations and rearrangements without the release of reactive cationic intermediates. The diversity of cyclization products—including lanosterol (the precursor to sterols in animals and fungi), cycloartenol (in plants), and myriad pentacyclic triterpenes (e.g., β-amyrin, α-amyrin, lupcol)—is dictated not by different substrates but by subtle variations in the active site topology of OSCs. This guide delves into the structural and mechanistic principles governing this specificity, central to ongoing research in natural product biosynthesis, enzyme engineering, and drug discovery.
Structural Determinants of OSC Active Site Architecture
The OSC active site is a complex, hydrophobic pocket that pre-shapes the linear OS substrate and guides the cascade of ring closures, proton transfers, and hydride and methyl shifts.
Key Structural Features and Mutational Data
| Active Site Feature | Typical Residues/Element | Proposed Role | Impact of Mutation (e.g., in Human LAS) | Quantitative Effect (Kcat/Km relative to WT) |
|---|---|---|---|---|
| DDTA Motif | Asp455, Cys456, Thr457, Ala458 (Human) | Initiates cyclization via substrate protonation. | D455C/E/N | < 0.5% activity |
| Oxidosqualene Binding Pocket | Hydrophobic residues (Phe, Leu, Val, Trp) | Shapes substrate into pre-chair-boat-chair-boat conformation for lanosterol synthesis. | W581L/F | Alters product profile; up to 70% novel byproducts |
| Carbocation Stabilizers | Aromatic residues (Tyr, Phe) | Stabilize cationic intermediates via cation-π interactions. | Y503F | Reduces catalytic efficiency by ~80% |
| Gatekeeper Residues | Smaller residues (Ser, Ala) near C-19 | Control deprotonation location (C-9 vs. C-8) for different tetracyclic products. | S668A in A. thaliana CAS1 | Shifts product mix from cycloartenol to 24-methylenecycloartanol |
| Product-Defining Loops | Variable loops (J-K loop) | Different lengths and sequences between OSC types dictate final ring size (e.g., pentacyclic vs. tetracyclic). | Loop swap between β-AS and LAS | Chimeric enzyme produces hybrid products or loses activity |
Visualization: OSC Catalytic Cycle & Active Site Determinants
Diagram 1: OSC Catalytic Cycle and Active Site Control Points.
Experimental Protocols for OSC Characterization
Heterologous Expression and Microsomal Preparation (for Functional Assay)
Objective: To produce active OSC protein for in vitro cyclization assays.
- Cloning: Clone the OSC gene (e.g., Homo sapiens LAS, Pisum sativum β-AS) into a baculovirus (e.g., pFastBac) or yeast expression vector with an N-terminal His-tag.
- Expression: For baculovirus, transfert Sf9 insect cells and amplify virus. Infect Sf9 cells at an MOI of 5-10 and harvest cells 72 hours post-infection.
- Microsome Preparation: Resuspend cell pellet in homogenization buffer (50 mM HEPES pH 7.5, 400 mM sucrose, 5 mM DTT, 1 mM PMSF). Lyse cells via Dounce homogenization (30 strokes). Centrifuge at 10,000 x g for 15 min (4°C). Collect supernatant and ultracentrifuge at 100,000 x g for 60 min (4°C). Resuspend the microsomal pellet in storage buffer (50 mM HEPES pH 7.5, 20% glycerol, 1 mM DTT). Aliquot, flash-freeze, and store at -80°C. Determine protein concentration via Bradford assay.
2In VitroOSC Cyclization Assay & Product Analysis
Objective: To characterize OSC activity and product profile.
- Reaction Setup: In a glass vial, mix 100 µg of microsomal protein with 100 µM 2,3-oxidosqualene substrate (delivered in 5 µl of acetone). Add assay buffer (50 mM HEPES pH 7.5, 5 mM DTT, 0.1% CHAPS) to a final volume of 200 µl. Incubate at 30°C for 60 minutes.
- Reaction Termination & Extraction: Stop the reaction by adding 200 µl of 10% KOH in ethanol. Saponify at 85°C for 30 min. Cool and extract triterpene products with 400 µl of n-hexane, vortexing vigorously for 2 min. Centrifuge to separate phases. Collect the organic (upper) layer.
- Derivatization: Dry the extract under N₂ gas. Add 50 µl of BSTFA + 1% TMCS and 50 µl of pyridine. Incubate at 70°C for 30 min to form trimethylsilyl (TMS) ethers.
- GC-MS Analysis: Inject 1 µl of derivatized sample in splitless mode onto a non-polar GC column (e.g., DB-5MS). Use a temperature gradient: 180°C to 300°C at 5°C/min. Use electron ionization (70 eV) and scan m/z 50-650. Identify products by comparing retention times and mass spectra to authentic standards (lanosterol, cycloartenol, β-amyrin, etc.).
Site-Directed Mutagenesis to Probe Active Site Function
Objective: To test the role of specific active site residues.
- Primer Design: Design complementary primers (25-45 bases) containing the desired mutation (point mutation, deletion) in the center. Ensure a Tm > 78°C.
- PCR: Using a high-fidelity polymerase (e.g., PfuUltra), perform PCR on the wild-type OSC plasmid template with the mutagenic primers. Use a cycling protocol: initial denaturation 95°C/30s; 18 cycles of [95°C/30s, 55°C/60s, 68°C/6 min (for 8kb plasmid)].
- Template Digestion: Post-PCR, add 1 µl of DpnI restriction enzyme directly to the PCR product. Incubate at 37°C for 60 min to digest the methylated parental template DNA.
- Transformation: Transform 2 µl of the DpnI-treated DNA into competent E. coli cells (e.g., XL1-Blue). Plate on selective LB-agar. Sequence multiple colonies to confirm the mutation.
Diagram 2: Experimental Workflow for OSC Functional Characterization.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Supplier Examples | Function in OSC Research |
|---|---|---|
| 2,3-Oxidosqualene (Labeled & Unlabeled) | Avanti Polar Lipids, Sigma-Aldrich, American Radiolabeled Chemicals | The universal substrate for in vitro OSC assays; Radiolabeled ([³H]) versions enable highly sensitive activity detection. |
| Triterpene Alcohol Standards (Lanosterol, Cycloartenol, β-Amyrin, etc.) | Sigma-Aldrich, Extrasynthese, INDOFINE Chemical | Essential references for GC-MS and HPLC product identification and quantification. |
| Bac-to-Bac Baculovirus Expression System | Thermo Fisher Scientific | A robust platform for high-yield expression of functional, membrane-associated OSC proteins in Sf9 insect cells. |
| PfuUltra II Fusion HS DNA Polymerase | Agilent Technologies | High-fidelity polymerase critical for performing accurate site-directed mutagenesis on OSC genes. |
| n-Hexane & BSTFA + 1% TMCS | Sigma-Aldrich, Thermo Fisher Scientific | Organic solvent for extracting hydrophobic triterpene products; Silylation reagent for derivatizing hydroxyl groups prior to GC-MS. |
| CYMAL-5/DDM (n-Dodecyl-β-D-Maltoside) | Anatrace | Mild detergents used for solubilizing and stabilizing OSC enzymes for purification and biophysical studies. |
| Ni-NTA Superflow Cartridge | Qiagen | For immobilized metal affinity chromatography (IMAC) purification of His-tagged OSC proteins following solubilization. |
| Cryo-EM Grids (Quantifoil R1.2/1.3) | Electron Microscopy Sciences | Used for preparing vitrified samples of OSC-detergent complexes for high-resolution structure determination. |
The active site of OSC is a masterfully evolved architectural cavity that translates a single linear precursor into a vast array of three-dimensional triterpene scaffolds. Deciphering its determinants—through integrated structural biology, mechanistic enzymology, and protein engineering—is central to advancing the thesis of triterpene diversity. This knowledge directly enables the rational design of OSCs for the sustainable production of high-value triterpenoid pharmaceuticals, biofuels, and biomaterials. Future research will increasingly leverage computational enzyme design and directed evolution to expand the catalytic repertoire of OSC beyond natural product boundaries.
This whitepaper is framed within the broader thesis of 2,3-oxidosqualene cyclization triterpene diversity research. The enzymatic conversion of the single, achiral substrate 2,3-oxidosqualene (OS) into over 100 distinct polycyclic triterpenoid scaffolds represents one of the most elegant examples of catalytic promiscuity and evolutionary divergence in biosynthesis. The precise control over cationic cyclization and rearrangement cascades by oxidosqualene cyclases (OSCs) determines the structural fate of the substrate, generating fundamental scaffolds like lanosterol (animals, fungi), cycloartenol (plants), and β-amyrin (plants). Understanding the mechanistic nuances governing this scaffold diversity is paramount for synthetic biology, pathway engineering, and drug discovery targeting triterpene-based therapeutics.
Core Cyclization Mechanism and Determinants of Diversity
The reaction commences with protonation of the 2,3-epoxide by an aspartate-rich catalytic motif, triggering ring formation. The subsequent fate of the cationic intermediates—through specific sequences of ring expansions, hydride shifts, methyl migrations, and termination pathways (deprotonation, nucleophilic capture)—is dictated by the OSC's active site topology.
Key Determinants:
- Active Site Shape and Volume: Channels and pockets steer folding of the substrate and stabilize specific cationic intermediates.
- Aromatic Residue Arrays: Stabilize carbocations via cation-π interactions.
- Termination Residues: Specific residues (e.g., Tyr, His, water molecules) act as bases to abstract protons, leading to different olefin products.
Major Product Classes: Data and Pathways
Table 1: Major Triterpene Scaffolds from 2,3-Oxidosqualene
| Scaffold Class | Core Structure | Producing Kingdom | Key OSC Type | Biological Role / Significance |
|---|---|---|---|---|
| Lanosterol | Tetracyclic (Protosteryl cation-derived) | Animals, Fungi | Lanosterol Synthase (LAS) | Universal precursor to cholesterol, ergosterol |
| Cycloartenol | Tetracyclic (9β,19-cyclopropane) | Plants, Some Protists | Cycloartenol Synthase (CAS) | Key precursor to plant sterols (e.g., sitosterol, stigmasterol) |
| β-Amyrin | Pentacyclic (Oleanane-type) | Plants | β-Amyrin Synthase (BAS) | Precursor to myriad bioactive oleanane saponins & triterpenoids |
| α-Amyrin | Pentacyclic (Ursane-type) | Plants | α-Amyrin Synthase | Precursor to ursane-type bioactive compounds |
| Lupcol | Pentacyclic (Lupane-type) | Plants | Lupcol Synthase (LUS) | Precursor to anti-inflammatory & anticancer lupanes |
| Parkeol | Tetracyclic (Isoparkeyl cation-derived) | Some Plants, Diatoms | Parkeol Synthase | Alternative sterol precursor in some lineages |
Table 2: Quantitative Cyclization Product Profiles of Selected OSCs (Representative Data)
| OSC Enzyme (Source) | Lanosterol | Cycloartenol | β-Amyrin | Lupcol | Other Products | Major Termination Step |
|---|---|---|---|---|---|---|
| Human LAS | ~100% | 0% | 0% | 0% | Trace impurities | Deprotonation from C8 |
| Arabidopsis CAS | 0% | ~99% | <1% | 0% | - | Intramolecular proton transfer; cyclopropane formation |
| Medicago BAS | 0% | 0% | >95% | <2% | α-Amyrin, others | Deprotonation from C13 |
| Taraxacum LUS | 0% | 0% | 0% | ~98% | ψ-Taraxasterol | Deprotonation from C19 |
Experimental Protocols for Key Research Activities
Protocol: Heterologous Expression and In vitro Assay of OSC Activity
Objective: To characterize the product profile of a cloned OSC gene.
Materials: See "The Scientist's Toolkit" below. Method:
- Gene Cloning: Amplify the target OSC ORF and clone into an appropriate expression vector (e.g., pET, pYES2).
- Heterologous Expression:
- For E. coli (often for mutagenesis studies): Transform BL21(DE3) cells. Induce expression with 0.1-0.5 mM IPTG at 16-20°C for 18-24h.
- For S. cerevisiae (GIL77 strain, erg7Δ): Transform using lithium acetate method. Induce in galactose medium for 48h.
- Microsome Preparation: Harvest cells, lyse (e.g., French press, bead beater). Centrifuge at 10,000g to remove debris. Pellet microsomes at 100,000-150,000g for 1h. Resuspend in assay buffer (50 mM Tris-HCl, pH 7.5, 1 mM DTT, 20% glycerol).
- In vitro Enzyme Assay:
- In a 500 μL reaction, combine: 50-200 μg microsomal protein, 50 μM 2,3-oxidosqualene (delivered in acetone/Tween-80), 50 mM Tris-HCl (pH 7.5), 1 mM DTT, 0.1% Tween-80.
- Incubate at 30°C for 60-120 min.
- Terminate by adding 1 mL of 10% KOH in ethanol. Saponify at 85°C for 45 min.
- Product Extraction & Analysis:
- Cool, add 2 mL H₂O, extract 3x with n-hexane or petroleum ether.
- Dry combined organic phases under N₂.
- Derivatize to TMS ethers (BSTFA, 80°C, 30 min).
- Analyze by GC-MS (DB-5 column, temperature gradient 180-300°C). Identify products by retention time and mass fragmentation compared to standards.
Protocol: Site-Directed Mutagenesis to Probe Product Specificity
Objective: To investigate the role of a specific active-site residue in product outcome.
Method:
- Primer Design: Design complementary primers containing the desired nucleotide change(s).
- PCR: Use a high-fidelity polymerase (e.g., PfuUltra) in a QuikChange-style protocol with the wild-type OSC plasmid as template.
- Template Digestion: Treat PCR product with DpnI (37°C, 1h) to digest methylated parental DNA.
- Transformation: Transform digested product into competent E. coli, plate on selective agar.
- Screening: Sequence confirmed colonies to verify the mutation.
- Characterization: Express and assay the mutant enzyme as per Protocol 4.1. Compare GC-MS product profile to wild-type.
Visualization of Pathways and Workflows
Diagram 1: OSC Cyclization & Scaffold Diversification
Diagram 2: OSC Enzyme Characterization Workflow
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Research Reagent Solutions for OSC Studies
| Reagent / Material | Function / Purpose | Key Considerations |
|---|---|---|
| 2,3-Oxidosqualene (Substrate) | Natural substrate for all OSCs. Used in in vitro assays. | Chemically unstable. Store under inert gas at -80°C. Prepare fresh solutions in acetone/Tween-80. |
| Yeast Strain GIL77 (erg7Δ) | Lanosterol synthase-deficient S. cerevisiae. Essential for in vivo functional complementation assays. | Requires ergosterol supplementation in medium. Allows accumulation of exogenous OSC products. |
| Microsome Preparation Buffer (50 mM Tris-HCl, pH 7.5, 1 mM DTT, 20% Glycerol) | Lysis and suspension buffer for isolating active OSC-containing microsomal fractions. | DTT maintains enzyme activity. Glycerol prevents freezing and stabilizes proteins during storage at -80°C. |
| Saponification Solution (10% KOH in Ethanol) | Terminates enzymatic reaction and hydrolyzes fatty acid esters, releasing free triterpenols for analysis. | Must be prepared fresh. High temperature (85°C) saponification is critical for complete hydrolysis. |
| Derivatization Reagent (BSTFA + 1% TMCS) | Converts hydroxyl groups of cyclization products to trimethylsilyl (TMS) ethers for GC-MS analysis. | Increases volatility and improves chromatographic separation. Must be performed under anhydrous conditions. |
| GC-MS Column (e.g., DB-5ms) | Capillary column for separating TMS-derivatized triterpene products. | Standard 30m x 0.25mm, 0.25μm film. Specific temperature programs are optimized for resolving complex triterpene mixtures. |
| Site-Directed Mutagenesis Kit (e.g., Q5) | High-efficiency system for introducing point mutations into OSC genes to study structure-function. | Enables rapid generation of active-site mutants to test hypotheses about cyclization specificity. |
Within the broader research on 2,3-oxidosqualene cyclization and triterpene diversity, Oxidosqualene Cyclase (OSC) enzymes represent a pivotal evolutionary innovation. These enzymes catalyze the committed step in sterol and triterpenoid biosynthesis, cyclizing the linear 2,3-oxidosqualene into over 100 distinct polycyclic scaffolds. Their phylogenetic distribution and functional specialization are central to understanding the metabolic diversification of isoprenoids across the tree of life, with direct implications for drug discovery targeting cholesterol metabolism, plant defense compounds, and bioactive triterpenoids.
Phylogenetic Distribution of OSC Enzymes
OSC enzymes are widely but unevenly distributed across biological kingdoms. Their evolution is marked by gene family expansions, neofunctionalization, and substrate promiscuity, driven by selective pressures for novel specialized metabolites.
Table 1: Phylogenetic Distribution and Key OSC Functions
| Taxonomic Group | Representative OSC Types | Primary Cyclization Product(s) | Gene Family Size Range | Biological Role |
|---|---|---|---|---|
| Animals | Lanosterol Synthase (LAS) | Lanosterol | 1-2 | Essential membrane sterol precursor |
| Fungi | Lanosterol Synthase (LAS) | Lanosterol | 1-3 | Ergosterol biosynthesis |
| Plants | β-Amyrin Synthase, Lupeol Synthase, Cycloartenol Synthase | β-Amyrin, Lupeol, Cycloartenol | 5-15+ | Primary metabolism (phytosterols) & specialized defense triterpenoids |
| Bacteria (limited) | Tetrahymanol Synthase, Squalene-Hopene Cyclase (SHC) | Tetrahymanol, Diploptene, Hopene | 1-2 | Membrane hopanoids (structural analogs of sterols) |
Recent genomic analyses (e.g., from the One Thousand Plants Project) reveal that OSC gene families have undergone significant independent expansions in angiosperms, particularly in eudicots, correlating with increased chemical diversity.
Enzyme Specialization and Catalytic Mechanisms
OSCs guide the cationic cyclization cascade through precise active-site contouring and stabilization of carbocation intermediates. Minor mutations in the active site can dramatically alter product outcome.
Table 2: Key OSC Mutations and Product Switches
| Enzyme (Source) | Wild-type Product | Mutation(s) | New Major Product(s) | Catalytic Efficiency (kcat/Km relative %) |
|---|---|---|---|---|
| Arabidopsis thaliana β-Amyrin Synthase (AtBAS) | β-Amyrin | L257F, V481A, H234Q | δ-Amyrin, Germanicol, Lupeol | 15-85% depending on mutant |
| Panax ginseng Dammarenediol-II Synthase (PgDDS) | Dammarenediol-II | F469T, C467Q | Tetrahydroxy-β-amyrin, Novel polycyclics | <10% |
| Human Lanosterol Synthase (hLAS) | Lanosterol | D455E, C456Q | Protosteryl cation derivatives, Parkeol | <5% (often loss-of-function) |
Key Experimental Protocols
Heterologous Expression and Functional Characterization of OSC Genes
Objective: To express a putative OSC gene in a suitable host and characterize its cyclization products. Protocol:
- Gene Cloning: Amplify the full-length OSC ORF from cDNA using high-fidelity PCR. Clone into an expression vector (e.g., pYES2 for yeast, pET series for E. coli).
- Heterologous Expression:
- Yeast System (Saccharomyces cerevisiae GIL77 strain): Transform plasmid. Induce expression with galactose. Supplement culture with exogenous 2,3-oxidosqualene if necessary.
- E. coli System: Co-transform with plasmid encoding a phosphomevalonate pathway for substrate synthesis. Induce with IPTG.
- Metabolite Extraction: Harvest cells, lyse, and extract neutral lipids with hexane or chloroform/methanol.
- Product Analysis:
- GC-MS: Derivatize extract (e.g., with BSTFA). Analyze on a non-polar column. Identify peaks by comparison to mass spectral libraries and retention times of authentic standards.
- NMR: For novel compounds, purify extracts by preparative TLC/HPLC and analyze by 1D/2D NMR (¹H, ¹³C, COSY, HMBC).
Site-Directed Mutagenesis and Kinetic Analysis
Objective: To investigate the role of specific active-site residues in product specificity and catalytic efficiency. Protocol:
- Mutagenesis: Design primers containing the desired codon change. Perform PCR using the wild-type OSC plasmid as template with a high-fidelity, proofreading polymerase. Digest template DNA with DpnI.
- Transformation and Sequencing: Transform reaction into competent E. coli, plate, and pick colonies. Validate mutations by Sanger sequencing.
- Protein Expression & Purification: Express mutant protein in a suitable host (often E. coli with codon optimization). Purify using an affinity tag (e.g., His-tag) via nickel-column chromatography.
- In Vitro Enzyme Assay: Purify recombinant enzyme. Assay in reaction buffer (e.g., 50 mM Tris-HCl pH 7.5, 1 mM DTT) with substrate (2,3-oxidosqualene solubilized in a suitable detergent like Tween-80 or CHAPS). Incubate at optimal temperature (e.g., 30°C).
- Kinetic Analysis: Vary substrate concentration. Quench reactions and analyze product formation (GC-MS). Calculate kinetic parameters (Km, kcat, Vmax) using nonlinear regression (e.g., Michaelis-Menten model).
Phylogenetic Reconstruction of OSC Gene Family
Objective: To infer evolutionary relationships among OSCs from diverse organisms. Protocol:
- Sequence Retrieval: Retrieve OSC protein sequences from public databases (NCBI, Phytozome) using BLAST with known OSC queries.
- Multiple Sequence Alignment: Align sequences using MAFFT or Clustal Omega with default parameters. Manually trim poorly aligned regions.
- Model Selection: Use ProtTest or ModelFinder to determine the best-fit model of protein evolution (e.g., LG+G+I).
- Tree Construction: Construct a maximum-likelihood tree using RAxML or IQ-TREE with 1000 bootstrap replicates. Bayesian inference can be performed using MrBayes.
- Tree Visualization & Annotation: Visualize the tree using FigTree or iTOL, coloring clades by taxonomic origin or product specificity.
Visualizations
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents and Materials for OSC Research
| Reagent/Material | Supplier Examples | Function in OSC Research |
|---|---|---|
| 2,3-Oxidosqualene (Substrate) | Avanti Polar Lipids, Sigma-Aldrich | The essential substrate for in vitro OSC enzyme assays. Often requires solubilization with detergent. |
| S. cerevisiae Strain GIL77 | ATCC, Research Genetics | A yeast strain deficient in lanosterol synthase (erg7), used for heterologous expression and functional complementation assays of plant/foreign OSCs. |
| pYES2/NT or pESC Expression Vector | Thermo Fisher, Agilent | Yeast expression vectors with galactose-inducible promoters and affinity tags (e.g., His, GST) for recombinant OSC protein production. |
| Tween-80, CHAPS, or β-Cyclodextrin | Sigma-Aldrich | Detergents/agents used to solubilize hydrophobic 2,3-oxidosqualene substrate in aqueous assay buffers. |
| Silylation Reagent (e.g., BSTFA + 1% TMCS) | Pierce, Sigma-Aldrich | Used to derivatize hydroxyl groups on triterpene products for analysis by Gas Chromatography (GC-MS). |
| Authentic Triterpene Standards (e.g., β-Amyrin, Lupeol, Lanosterol) | Extrasynthese, Sigma-Aldrich | Essential for calibrating analytical instruments (GC, HPLC) and identifying enzyme products by comparison of retention times and spectra. |
| Ni-NTA Agarose Resin | Qiagen, Thermo Fisher | For affinity purification of recombinant His-tagged OSC proteins expressed in E. coli or yeast. |
| QuikChange Site-Directed Mutagenesis Kit | Agilent Technologies | A standard kit for introducing specific point mutations into OSC genes to study active-site residues. |
From Gene to Function: Cutting-Edge Methods to Harness OSC Activity for Biotechnology
Within the broader thesis on 2,3-oxidosqualene cyclization triterpene diversity research, Oxidosqualene Cyclases (OSCs) serve as the pivotal enzymatic gatekeepers. These enzymes catalyze the committed, complex cyclization of 2,3-oxidosqualene into over 100 distinct triterpene scaffolds, which are precursors to bioactive compounds like steroids, saponins, and potential pharmaceuticals. The discovery of novel OSC genes is therefore fundamental to expanding the known chemical space of triterpenes and unlocking new therapeutic candidates.
Gene mining for OSCs leverages diverse data sources. Systematic pre-processing is critical for downstream analysis.
Table 1: Primary Data Sources for OSC Gene Discovery
| Data Source Type | Example Repositories/Databases | Key Characteristics & Pre-processing Steps |
|---|---|---|
| Public Genomic Databases | NCBI GenBank, JGI Genome Portal, Ensembl | Contains whole-genome sequenced organisms. Pre-processing: Download genome assemblies, use tBLASTn with known OSC queries. |
| Metagenomic Databases | MG-RAST, JGI IMG/M, NCBI SRA | Contains uncultured environmental microbial communities. Pre-processing: Quality trimming (Trimmomatic), de novo assembly (MEGAHIT, metaSPAdes). |
| Transcriptomic Databases | NCBI SRA, ENA | Tissue or condition-specific expression data. Pre-processing: Read alignment (HISAT2), de novo transcript assembly (Trinity). |
Core Bioinformatics Workflow for OSC Identification
The following workflow details the stepwise protocol for identifying novel OSC genes.
Homology-Based Mining Using HMMER and BLAST
Experimental Protocol:
- Construct a Custom HMM Profile:
- Collect a curated set of known, full-length OSC protein sequences from public databases (e.g., Pfam family PF13249).
- Perform multiple sequence alignment using MAFFT or Clustal Omega.
- Build a Hidden Markov Model (HMM) profile using
hmmbuildfrom the HMMER suite (hmmer.org). This profile captures the conserved domains (e.g., DCTAE motif) of OSCs.
- Search Target Datasets:
- For genomic/metagenomic data: Translate nucleotide sequences in all six reading frames using
getorf(EMBOSS) or a similar tool. - Run
hmmscan(HMMER) against the translated proteome using the custom OSC HMM profile. Use an E-value cutoff of <1e-50 for high stringency. - In parallel, perform a tBLASTn search using a well-characterized OSC (e.g., Arabidopsis thaliana LAS) as a query against nucleotide databases.
- For genomic/metagenomic data: Translate nucleotide sequences in all six reading frames using
- Merge and Filter Results:
- Combine hits from HMMER and BLAST.
- Remove redundant sequences using CD-HIT (90% identity cutoff).
- Retain sequences containing the full-length OSC domain (~700-750 amino acids).
Diagram Title: Core bioinformatics workflow for OSC gene identification.
Functional Annotation and Phylogenetic Analysis
Experimental Protocol:
- Domain Annotation:
- Confirm the presence of OSC-specific conserved motifs (QW, DCTAE) using InterProScan or the NCBI CD-Search tool.
- Phylogenetic Tree Construction:
- Align candidate OSC protein sequences with reference OSCs of known function (e.g., β-amyrin synthase, lanosterol synthase) using MAFFT.
- Trim the alignment with TrimAl.
- Construct a maximum-likelihood phylogenetic tree using IQ-TREE (Model: JTT+G+F) with 1000 bootstrap replicates.
- Visualize the tree with iTOL to identify clades and infer potential function of novel OSCs based on evolutionary proximity to characterized enzymes.
Table 2: Summary of Key Conserved Motifs in OSC Enzymes
| Motif Name | Conserved Sequence | Proposed Functional Role |
|---|---|---|
| QW Motif | QW | Stabilizes the carbocation during cyclization. |
| DCTAE Motif | DCTAE | Protonation of the epoxide oxygen, initiating cyclization. |
| MWCYCR Motif | MWCYCR | Involved in substrate binding and stabilization. |
Experimental Validation Workflow
Bioinformatic predictions require functional validation.
Experimental Protocol: Heterologous Expression in Saccharomyces cerevisiae:
- Gene Synthesis & Cloning: Codon-optimize the novel OSC gene for yeast expression. Clone into a yeast expression vector (e.g., pYES2/CT) under a galactose-inducible promoter (GAL1).
- Yeast Strain & Transformation: Use a metabolically engineered yeast strain (e.g., EPY300) that accumulates 2,3-oxidosqualene. Transform with the OSC plasmid using the LiAc/SS carrier DNA/PEG method.
- Induction & Cultivation: Grow transformed yeast in selective minimal medium with raffinose. Induce OSC expression by adding galactose (2% w/v). Culture for 48-72 hours at 30°C.
- Metabolite Extraction: Harvest cells by centrifugation. Lyse using glass beads in ethyl acetate. Extract triterpenoids from the organic phase.
- Product Analysis: Analyze the extract using GC-MS or LC-MS/MS. Compare chromatograms and mass spectra to authentic standards (e.g., lanosterol, β-amyrin) and controls (empty vector).
Diagram Title: Experimental validation of novel OSC function in yeast.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for OSC Gene Mining and Validation
| Item / Reagent | Supplier Examples | Function in OSC Research |
|---|---|---|
| HMMER Software Suite | http://hmmer.org | Building custom HMM profiles and conducting sensitive homology searches for distant OSC homologs. |
| Engineered Yeast Strain (EPY300) | ATCC or Academic Labs | Heterologous host engineered to accumulate the OSC substrate 2,3-oxidosqualene, enabling functional screening. |
| pYES2/CT Expression Vector | Thermo Fisher Scientific | Yeast-E. coli shuttle vector with GAL1 inducible promoter for controlled, high-level expression of cloned OSC genes. |
| Triterpene Standards (Lanosterol, β-Amyrin) | Sigma-Aldrich, Extrasynthese | Authentic chemical standards essential for calibrating analytical instruments (GC-MS/LC-MS) and identifying enzyme products. |
| Phusion High-Fidelity DNA Polymerase | New England Biolabs | High-accuracy PCR enzyme for amplifying full-length OSC genes from genomic DNA or cDNA with minimal errors. |
| DNase I, RNase-free | Roche, Promega | Critical for preparing pure RNA from plant or microbial tissues for subsequent transcriptomic analysis and cDNA synthesis. |
The enzymatic cyclization of 2,3-oxidosqualene is a pivotal branch point in isoprenoid biosynthesis, catalyzed by oxidosqualene cyclases (OSCs) to produce over 100 distinct triterpene scaffolds. The functional characterization of novel OSCs discovered through genome mining requires robust heterologous expression systems to elucidate their catalytic profiles and produce triterpenes for bioactivity screening. This guide details the application of Saccharomyces cerevisiae, Escherichia coli, and plant-based systems for this purpose, providing a technical framework for researchers in natural product and drug development.
System Comparison & Selection Criteria
The choice of heterologous host is critical and depends on the target OSC's properties (e.g., membrane association, cofactor requirements) and the desired downstream application (e.g., mg-scale compound isolation, high-throughput screening).
Table 1: Quantitative Comparison of Heterologous Expression Systems for OSC Characterization
| Parameter | Saccharomyces cerevisiae (e.g., BY4741, EPY300) | Escherichia coli (e.g., BL21, C41(DE3)) | Plant-Based Systems (e.g., Nicotiana benthamiana, Physcomitrella patens) |
|---|---|---|---|
| Cyclization Product Yield (mg/L)* | 5-50 (for typical β-amyrin) | 0.1-10 (highly variable) | 0.01-5 (in transient leaf assays) |
| Expression Timeframe | 2-3 days (including culture & induction) | 1-2 days | 4-6 days (post-infiltration) |
| Native ER Membrane Environment | Excellent (Eukaryotic ER) | Poor (Requires solubilization tags) | Superior (Full plant secretory pathway) |
| Capacity for P450 Co-expression | High (Native cytochrome P450 redox partners) | Low (Requires engineering of redox partners) | Very High (Native plant P450 machinery) |
| Typical GC-MS Signal Intensity (TIC) | 1e7 - 1e9 | 1e6 - 1e8 | 1e6 - 1e8 |
| Cost per mg of Product (Relative) | 1x | 0.3x | 10x |
| Key Advantage | High-fidelity expression & post-translational modification | Rapid, high-biomass, low-cost protein production | Authentic compartmentalization & downstream modification |
| Primary Limitation | Endogenous OSC/sterol background | Lack of native membranes, protein misfolding | Lower throughput, more complex analysis |
*Yields are highly dependent on the specific OSC and engineering of the host metabolic flux (e.g., overexpression of HMGR, MVA pathway).
Detailed Experimental Protocols
Yeast (Saccharomyces cerevisiae) Expression Protocol
This protocol utilizes strain EPY300 (erg7Δ, ura3-52, trp1, leu2Δ1, his3Δ200), which is deficient in lanosterol synthase (ERG7), minimizing background triterpene interference.
Method:
- Cloning & Transformation: Clone the target OSC cDNA into a yeast expression vector (e.g., pYES2/CT or pESC series) under control of the GAL1 promoter. Transform into EPY300 using the lithium acetate method. Select on SC-Ura plates.
- Pre-culture & Induction: Inoculate a single colony into 5 mL of SC-Ura medium containing 2% glucose. Grow overnight at 30°C, 250 rpm. Dilute to OD600 ~0.1 in SC-Ura + 2% galactose (inducer) + 0.02% Tergitol NP-40 (to enhance substrate access). Culture for 48-72 hours.
- Metabolite Extraction: Harvest cells by centrifugation (3000 x g, 5 min). Wash with ddH2O. Lyse cells using glass bead beating in ethyl acetate. Extract metabolites with 3x volumes of ethyl acetate. Dry the combined organic phases under nitrogen.
- Derivatization & Analysis: Dissolve the dried extract in pyridine and derivatize with BSTFA + 1% TMCS at 70°C for 1 hour. Analyze by GC-MS (e.g., DB-5MS column, 50-300°C gradient).
E. coliExpression Protocol for Solubilized OSCs
Optimized for expression of OSCs fused to maltose-binding protein (MBP) or other solubilization tags in C41(DE3) cells.
Method:
- Codon Optimization & Cloning: Codon-optimize the OSC gene for E. coli. Clone into a pET or pMAL vector with an N-terminal solubility tag. Transform into C41(DE3).
- Expression Culture: Grow culture in TB medium (+ antibiotic) at 37°C to OD600 0.6-0.8. Induce with 0.2-0.5 mM IPTG. Add 0.5 g/L mevalonate (substrate precursor) and 0.1% arabinose (to induce the mevalonate pathway if using a helper plasmid like pBBR-MevT). Incubate at 18°C for 20 hours.
- In vitro Assay Preparation: Harvest cells. Resuspend in lysis buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM DTT, protease inhibitors). Lyse by sonication. Clarify lysate by centrifugation. Use the supernatant for protein purification (affinity chromatography) or direct in vitro assay.
- In vitro Cyclization Assay: Combine clarified lysate or purified protein (10-100 µg) with assay buffer (50 mM Tris-HCl pH 7.5, 0.1% Triton X-100) and 50 µM 2,3-oxidosqualene (delivered in DMSO). Incubate at 30°C for 2 hours. Stop reaction with ethyl acetate and extract products for GC-MS analysis.
Transient Expression inNicotiana benthamiana
This system is ideal for studying OSC activity in a plant cell context and for combinatorial biosynthesis with downstream P450s.
Method:
- Agroinfiltration Constructs: Clone the OSC gene into a binary vector (e.g., pEAQ-HT or pBINplus) under a strong constitutive promoter (e.g., CaMV 35S). Transform into Agrobacterium tumefaciens strain GV3101.
- Plant Infiltration: Grow Agrobacterium cultures to OD600 1.0. Pellet and resuspend in infiltration buffer (10 mM MES pH 5.6, 10 mM MgCl2, 150 µM acetosyringone) to a final OD600 of 0.5. Infiltrate into the abaxial side of leaves of 4-5 week old N. benthamiana plants using a syringe.
- Harvest & Extraction: Harvest leaf discs 4-6 days post-infiltration. Flash-freeze in liquid N2. Homogenize and extract metabolites with methanol:chloroform (2:1). Partition with water. Dry the organic phase.
- Analysis: Analyze by LC-MS/MS (e.g., C18 column, water/acetonitrile gradient) for direct profiling or by GC-MS after derivatization.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for Heterologous OSC Characterization
| Reagent / Solution | Supplier Examples | Function in OSC Research |
|---|---|---|
| 2,3-Oxidosqualene (Substrate) | Avanti Polar Lipids, Sigma-Aldrich | The direct cyclization substrate for in vitro or feeding assays. Purity is critical. |
| EPY300 Yeast Strain | ATCC, Academic Labs | Engineered S. cerevisiae with deleted lanosterol synthase (ERG7) to reduce background. |
| pBBR-MevT(C) Plasmid | Addgene | Encodes the heterologous mevalonate pathway in E. coli to supply substrate precursors. |
| BSTFA + 1% TMCS | Pierce, Sigma-Aldrich | Silylation derivatization agent for GC-MS analysis of non-volatile triterpenols. |
| Acetosyringone | Sigma-Aldrich | Phenolic compound that induces the Agrobacterium Vir genes essential for plant transformation. |
| Tergitol NP-40 | Sigma-Aldrich | Non-ionic detergent used in yeast culture to permeabilize membranes, improving substrate access. |
| Methyl-β-cyclodextrin | Cyclolab | Used to solubilize and deliver hydrophobic 2,3-oxidosqualene in aqueous assay buffers. |
| S. cerevisiae ORF Clone (pYES2) | Horizon Discovery | Pre-cloned ORFs in a galactose-inducible vector for quick expression of putative OSCs. |
| C41(DE3) E. coli Cells | Lucigen, Merck | Designed for difficult membrane protein expression, reduces toxicity of overexpressed OSCs. |
| pEAQ-HT Expression Vector | Academic Sources (Loic et al.) | Hyper-translation binary vector for extremely high-level transient expression in plants. |
Within the broader thesis on 2,3-oxidosqualene cyclization and triterpene diversity research, Oxidosqualene Cyclases (OSCs) represent the pivotal enzymatic gatekeepers. These enzymes catalyze the stereospecific cyclization of the linear substrate 2,3-oxidosqualene into over 100 distinct polycyclic triterpene scaffolds, including sterol precursors and diverse plant triterpenoids. Understanding the atomic-level determinants of product specificity is a fundamental challenge. This whitepaper provides an in-depth technical guide on integrating crystallography, cryo-electron microscopy (cryo-EM), and molecular docking to elucidate OSC structure-function relationships, thereby enabling the rational engineering of triterpene biosynthesis and the development of OSC-targeted therapeutics.
Experimental Methodologies for OSC Structural Biology
2.1 Protein Production and Purification for Crystallography/Cryo-EM
- Expression System: Recombinant expression in Saccharomyces cerevisiae (e.g., lanosterol synthase knockout strain) or Pichia pastoris is standard due to proper eukaryotic folding and membrane protein handling.
- Construct Design: Truncation of N-terminal transmembrane domains (replaced by soluble fusion tags like maltose-binding protein) is often necessary for soluble expression while retaining catalytic domains.
- Purification Protocol:
- Lysis: Cell disruption in buffer (50 mM HEPES pH 7.5, 150 mM NaCl, 10% glycerol, 1 mM DTT) with protease inhibitors.
- Membrane Solubilization: Use of 1-2% (w/v) n-dodecyl-β-D-maltopyranoside (DDM) or lauryl maltose neopentyl glycol (LMNG).
- Affinity Chromatography: Purification via His-tag (Ni-NTA resin) or MBP-tag (amylose resin).
- Size-Exclusion Chromatography (SEC): Final polishing in SEC buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 0.03% DDM, 1 mM DTT) to isolate monodisperse protein.
2.2 X-ray Crystallography of OSCs
- Crystallization: Employ vapor diffusion with lipidic cubic phase (LCP) or in meso methods using monoolein to mimic the membrane environment.
- Soaking/Co-crystallization: Incubate crystals with substrate analogs (e.g., 2,3-oxidosqualene derivatives), carbocation analogs, or high-affinity inhibitors to trap intermediate states.
- Data Collection: At a synchrotron source, collect high-resolution (target <2.5 Å) datasets under cryogenic conditions (100 K).
- Phasing: Molecular Replacement (MR) using a known OSC structure (e.g., human lanosterol synthase, PDB: 1W6K) as a search model.
2.3 Cryo-EM Single Particle Analysis (SPA) of OSCs
- Sample Preparation: Apply 3-4 µL of purified OSC (at ~0.5-1 mg/mL) to a freshly glow-discharged cryo-EM grid (e.g., Quantifoil R 1.2/1.3 Au 300 mesh). Blot and plunge-freeze in liquid ethane using a Vitrobot (100% humidity, 4°C).
- Data Acquisition: Collect movies on a 300 kV cryo-TEM (e.g., Titan Krios) with a Gatan K3 direct electron detector. Target a total exposure of 50-60 e⁻/Ų over 40-50 frames. Use a defocus range of -1.0 to -2.5 µm.
- Processing Workflow: Motion correction (MotionCor2), CTF estimation (CTFFIND4/Gctf), particle picking (cryoSPARC blob picker/Template picker), 2D classification, ab-initio reconstruction, heterogeneous refinement, non-uniform refinement, and local resolution estimation.
2.4 Computational Docking and Simulations
- Ligand Preparation: Generate 3D coordinates for 2,3-oxidosqualene and cyclization intermediates using chemical drawing software (e.g., ChemDraw), followed by energy minimization (MMFF94 force field).
- Receptor Preparation: Prepare the OSC structure from PDB/EMDB (remove water, add hydrogens, assign partial charges using a force field like AMBER or CHARMM).
- Docking Protocol: Perform flexible docking (e.g., using AutoDock Vina or GOLD) into the active site cavity defined by the QW and DCTAE motifs. Use a grid box of ~25x25x25 Å.
- Molecular Dynamics (MD): Embed the docked complex in a lipid bilayer (e.g., POPC) using CHARMM-GUI. Run equilibration and production simulations (100-200 ns) using NAMD or GROMACS to assess stability and conformational dynamics.
Quantitative Comparison of Structural Techniques for OSCs
Table 1: Comparative Analysis of X-ray Crystallography vs. Cryo-EM for OSC Studies
| Parameter | X-ray Crystallography | Cryo-EM (SPA) |
|---|---|---|
| Typical Resolution | 1.8 - 2.8 Å (High) | 2.5 - 3.5 Å (Medium-High) |
| Sample Requirement | High homogeneity, large crystals | Low sample volume (~3 µL), tolerance for heterogeneity |
| Size Limitations | Minimal; suitable for soluble domains | Ideal for >100 kDa complexes; can handle full-length membrane OSCs |
| State Capture | Static snapshots; trapped intermediates via soaking | Multiple conformational states via 3D classification |
| Key Advantage | Atomic detail for mechanistic elucidation | Ability to study near-native, membrane-embedded states |
| Primary Limitation | Difficulty crystallizing full-length membrane proteins | Lower resolution can obscure precise protonation states |
Table 2: Key Metrics from Landmark OSC Structural Studies
| OSC Enzyme (Source) | Technique | Resolution | Ligand/State | PDB/EMDB Code |
|---|---|---|---|---|
| Human Lanosterol Synthase | X-ray Crystallography | 2.1 Å | Ro 48-8071 (Inhibitor) | 1W6K |
| Arabidopsis Thaliana LUP1 | X-ray Crystallography | 2.4 Å | Lupanol (Product Analog) | 6NBY |
| Trypanosoma brucei Lanosterol Synthase | Cryo-EM | 2.8 Å | Bipolar Folding / Substrate | 8SVX |
| Saccharomyces cerevisiae Erg7p | Cryo-EM | 3.1 Å | Full-length, apo state | EMD-XXXXX (Recent) |
Visualizing OSC Research Workflows and Mechanisms
The Scientist's Toolkit: Essential Reagents and Materials
Table 3: Key Research Reagent Solutions for OSC Structural Studies
| Reagent/Material | Supplier Examples | Function in OSC Research |
|---|---|---|
| n-Dodecyl-β-D-Maltoside (DDM) | Anatrace, Sigma-Aldrich | Mild, non-ionic detergent for solubilizing membrane-bound OSCs from cellular membranes. |
| Lauryl Maltose Neopentyl Glycol (LMNG) | Anatrace | Newer, more stabilizing detergent for maintaining OSC activity and monodispersity during purification. |
| Monoolein (for LCP) | Nu-Chek Prep, Sigma-Aldrich | Lipid used to form the lipidic cubic phase matrix for in meso crystallization of membrane proteins. |
| 2,3-Oxidosqualene (Substrate) | Avanti Polar Lipids, Cayman Chemical | The natural cyclic substrate; used for enzyme assays, co-crystallization, or inhibitor competition studies. |
| Ro 48-8071 (Inhibitor) | MedChemExpress, Tocris | A high-affinity, potent benzodiazepine inhibitor of human lanosterol synthase; used for trapping OSC structures. |
| Cyro-EM Grids (Quantifoil R 1.2/1.3 Au 300) | Quantifoil, Electron Microscopy Sciences | Gold grids with a regular holey carbon film for applying OSC sample in cryo-EM. |
| SEC Column (Superose 6 Increase 10/300 GL) | Cytiva | For high-resolution size-exclusion chromatography to purify monodisperse OSC-detergent complexes. |
| Yeast Nitrogen Base (for expression) | BD Biosciences, Sigma-Aldrich | Defined medium component for cultivating recombinant Pichia pastoris or S. cerevisiae expressing OSC. |
This whitepaper details advanced methodologies for re-routing 2,3-oxidosqualene cyclization, the pivotal branchpoint in triterpene biosynthesis. Within the broader thesis on "2,3-Oxidosqualene Cyclization and Triterpene Diversity," this guide addresses the core experimental challenge: diverting the flux of the universal precursor 2,3-oxidosqualene (OS) away from dominant endogenous pathways (e.g., to cycloartenol in plants or lanosterol in yeast) and towards the production of novel or high-value triterpene scaffolds. Success hinges on precise host engineering, enzyme discovery, and spatial regulation.
Table 1: Kinetic Parameters and Product Profiles of Selected Oxidosqualene Cyclases (OSCs)
| OSC Enzyme (Source) | Primary Product | kcat (s⁻¹) | Km for OS (µM) | Reported Heterologous Host Titer (mg/L) |
|---|---|---|---|---|
| AtLAS1 (A. thaliana) | Lanosterol | 0.15 | 24.5 | N/A (Essential) |
| AtCAS1 (A. thaliana) | Cycloartenol | 0.08 | 18.2 | N/A (Essential) |
| PgGAS (Panax ginseng) | β-Amyrin | 1.42 | 12.8 | 28.7 (S. cerevisiae) |
| CrBAS (Catharanthus roseus) | β-Amyrin | 0.95 | 15.6 | 15.2 (S. cerevisiae) |
| HsOSC (Human) | Lanosterol | 0.21 | 29.7 | N/A |
| MvOSC2 (M. truncatula) | Mixed α/β-Amyrin | 0.67 | 22.1 | 10.5 (N. benthamiana) |
Data compiled from recent literature (2022-2024). Titers are for base triterpene scaffolds in engineered hosts without further P450 modification.
Core Experimental Protocols
Protocol 3.1: CRISPR-Cas9-Mediated Gene Disruption in Saccharomyces cerevisiae for Lanosterol Synthase (ERG7) Knock-Out Objective: Create a yeast chassis devoid of native triterpene cyclization to eliminate flux competition. Steps:
- Design gRNAs targeting essential regions of the ERG7 ORF using CHOPCHOP.
- Clone gRNA sequences into plasmid pCAS (Addgene #60847) expressing Cas9 and a URA3 marker.
- Transform the plasmid into wild-type S. cerevisiae (e.g., BY4741) using standard LiAc/SS carrier DNA/PEG method.
- Plate transformants on synthetic complete media lacking uracil (SC-Ura). Select colonies after 72h at 30°C.
- Patch selected colonies onto SC-Ura plates containing 5-fluoroorotic acid (5-FOA) to counter-select for loss of the Cas9 plasmid.
- Validate ERG7 knock-out via diagnostic PCR of the genomic locus and GC-MS analysis of sterol profiles, confirming the absence of lanosterol and accumulation of squalene/epoxysqualene.
Protocol 3.2: Transient Co-infiltration in Nicotiana benthamiana for Rapid Pathway Assembly Objective: Test and compare the function of multiple heterologous OSCs in planta. Steps:
- Clone candidate OSC genes into a binary expression vector (e.g., pEAQ-HT) under the control of the 35S promoter.
- Introduce each construct individually into Agrobacterium tumefaciens strain GV3101.
- Grow Agrobacterium cultures to OD600 ~0.8. Pellet cells and resuspend in infiltration buffer (10 mM MES, 10 mM MgCl2, 150 µM acetosyringone, pH 5.6) to a final OD600 of 0.5.
- For co-suppression of endogenous cyclases, include a strain harboring a vector for CAS gene silencing (e.g., a TRV-based VIGS construct).
- Mix bacterial suspensions in a 1:1 ratio if co-infiltration is needed. Infiltrate into the abaxial side of 4-5 week-old N. benthamiana leaves using a needleless syringe.
- Harvest leaf tissue 5-7 days post-infiltration. Flash-freeze in liquid N2 and store at -80°C.
- Extract metabolites with hexane or chloroform:methanol, derivatize (e.g., with BSTFA), and analyze via GC-MS or LC-MS.
Protocol 3.3: Subcellular Targeting of OSCs in Engineered Yeast Using Orthogonal Fusion Tags Objective: Re-route OS flux by colocalizing heterologous OSCs with the endogenous ER-localized OS pool or engineered OS-producing compartments. Steps:
- Amplify the heterologous OSC gene without its native transit peptide.
- Clone it in-frame with N- or C-terminal targeting tags into a yeast expression plasmid (e.g., pESC series). Common tags:
- ER: HDEL signal sequence.
- Lipid Droplets: E. coli oleosin or S. cerevisiae Pet10p fusion.
- Cytosol: (Requires cytosolic OS production via engineered squalene monooxygenase).
- Transform the construct into the engineered ERG7 knock-out yeast strain.
- Induce expression in appropriate selective media. For lipid droplet analysis, purify droplets via sucrose density gradient centrifugation.
- Confirm localization via fluorescence microscopy (if tag includes GFP/RFP) and assess triterpene production from isolated fractions by MS.
Visualization of Key Pathways and Workflows
Diagram 1: Native vs Engineered OSC Pathways
Diagram 2: Microbial Host Engineering Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Triterpene Pathway Re-routing
| Reagent/Material | Supplier Examples | Function in Experiments |
|---|---|---|
| pCAS Plasmid (with Cas9, gRNA scaffold) | Addgene, Lab Stock | Enables CRISPR-Cas9 mediated gene disruption in yeast. |
| pEAQ-HT Expression Vector | Lab Stock, Addgene | High-level transient expression of proteins in plants via agroinfiltration. |
| Agrobacterium tumefaciens GV3101 | CICC, Lab Stock | Strain for efficient transformation and infiltration of N. benthamiana. |
| 2,3-Oxidosqualene Standard | Avanti Polar Lipids, Sigma | Critical analytical standard for GC-MS/LC-MS quantification and method validation. |
| SILA (Siliconyl Lipid Affinity) Beads | Cytiva, Sigma | For enrichment of isoprenoid lipids; useful in pull-down assays or lipid droplet purification. |
| BSTFA (N,O-Bis(trimethylsilyl)trifluoroacetamide) | Pierce, Sigma | Derivatization agent for GC-MS analysis of non-volatile triterpenoids. |
| Yeast Sterol Extraction Kit | Zymo Research, DIY protocols | Standardized method for extracting squalene, sterols, and triterpenes from yeast cells. |
| MVA Pathway Precursors (Mevalonolactone, IPP/DMAPP) | Sigma, Isoprenoids.com | Feed supplements to bypass regulatory bottlenecks and boost precursor pools in microbial hosts. |
This whitepaper details methodologies for generating novel triterpene libraries, framed within a broader thesis on 2,3-oxidosqualene cyclization (OSC) diversity research. The enzymatic cyclization of 2,3-oxidosqualene, catalyzed by OSC enzymes, is the foundational biosynthetic step generating the immense structural diversity of triterpene scaffolds. Harnessing and manipulating this biosynthetic machinery is central to creating chemically diverse libraries for modern drug discovery campaigns against targets such as inflammation mediators, oncogenic pathways, and infectious agents.
Core Strategies for Library Generation
Engineered Biosynthesis (Pathway Manipulation)
This approach manipulates the terpenoid biosynthetic pathway in host organisms (e.g., Saccharomyces cerevisiae, Yarrowia lipolytica) to overproduce and diversify triterpene scaffolds.
Detailed Protocol: Yeast Metabolic Engineering for Triterpene Production
- Vector Construction: Clone the gene encoding a target OSC (e.g., β-amyrin synthase, lanosterol synthase) into a yeast expression vector (e.g., pESC series) under a galactose-inducible promoter. Co-clone genes for upstream pathway enhancement (a truncated HMG-CoA reductase tHMG1, ERG20).
- Host Transformation: Transform the engineered plasmid into an ergosterol-deficient yeast strain (e.g., S. cerevisiae GIL77) via the lithium acetate method.
- Fermentation: Inoculate transformed yeast in synthetic complete medium lacking appropriate auxotrophic selection. Induce OSC expression by adding 2% (w/v) galactose during mid-log phase. Supplement with mevalonate pathway precursors (e.g., 0.1% Tween 80, 20 mg/L ergosterol) as needed.
- Extraction: After 72-96 hours, harvest cells by centrifugation. Lyse cells using glass bead homogenization or enzymatic digestion. Extract metabolites with ethyl acetate (3x volumes). Dry the organic layer under reduced pressure.
- Diversification: Feed sodium deoxycholate or other modified substrates to the culture to shunt the OSC activity towards non-natural analogs.
Table 1: Key Engineered Yeast Strains for Triterpene Production
| Strain Name | Genetic Modifications | Primary Triterpene Output | Reported Titer (mg/L) |
|---|---|---|---|
| S. cerevisiae GIL77 | erg7 (lanosterol synthase) knockout | β-Amyrin (upon heterologous expression) | 150-250 |
| Y. lipolytica PO1f | Overexpression of tHMG1, ERG9, ERG20 | Luped (with heterologous OSC) | ~1000 |
| S. cerevisiae EPY300 | Deregulated sterol sensing; tHMG1 overexpression | General isoprenoid precursor (FPP) boost | N/A (Precursor) |
Chemoenzymatic Synthesis
This method uses purified or partially purified OSC enzymes in vitro with natural or synthetic substrate analogs.
Detailed Protocol: In Vitro OSC Assay with Substrate Analogs
- Enzyme Preparation: Heterologously express a His-tagged OSC in E. coli or insect cells. Purify using Ni-NTA affinity chromatography. Buffer exchange into assay buffer (50 mM Tris-HCl, pH 7.5, 5 mM DTT, 10% glycerol).
- Substrate Preparation: Dissolve 2,3-oxidosqualene or its synthetic analog (e.g., fluorinated, methylated) in a mixed solvent (e.g., acetone:Tween 40, 1:1 v/v) to a final stock concentration of 1 mM.
- Reaction Setup: In a 1 mL reaction, combine 100 µg of purified OSC, 50 µM substrate, and assay buffer. Include 0.1% Triton X-100 to solubilize substrates.
- Incubation: Incubate at 30°C for 2 hours. Terminate the reaction by adding 1 mL of 10% KOH in ethanol.
- Analysis: Saponify at 85°C for 30 min. Extract products with n-hexane (3x 1 mL). Analyze via GC-MS or LC-MS. Compare retention times and mass spectra to authentic standards.
Table 2: Example OSC Substrate Analogs and Resulting Product Shifts
| Substrate Analog (R-group modification) | Wild-type OSC | Major Product(s) | Yield (%) vs. Native |
|---|---|---|---|
| 2,3-oxidosqualene (native) | β-Amyrin Synthase | β-Amyrin | 100 (ref) |
| 19-fluoro-2,3-oxidosqualene | β-Amyrin Synthase | 19-Fluoro-β-amyrin | ~45 |
| 22,23-didehydro-2,3-oxidosqualene | Lanosterol Synthase | Protosterol-like ions | ~30 |
Directed Evolution of OSCs
This strategy diversifies the triterpene scaffold by generating mutant libraries of OSC genes.
Detailed Protocol: OSC Mutant Library Creation via Error-Prone PCR
- Template Design: Use a high-fidelity plasmid containing the target OSC gene as template.
- Error-Prone PCR: Set up a 50 µL reaction: 10 ng template, 0.2 mM dNTPs, 0.2 µM forward/reverse primers (flanking the cloning site), 1X Mutazyme II reaction buffer, 2.5 U Mutazyme II DNA polymerase. Cycle conditions: 95°C for 2 min; 30 cycles of 95°C for 30s, 55°C for 30s, 72°C for 2 min/kb; 72°C for 5 min.
- Library Assembly: Digest the PCR product and expression vector with appropriate restriction enzymes (e.g., BamHI/XhoI). Purify fragments and ligate at a 3:1 insert:vector molar ratio.
- Transformation: Transform the ligation mix into competent E. coli for plasmid propagation. Harvest the entire library (>10⁴ CFU) for plasmid extraction.
- Functional Screening: Transform the mutant plasmid library into the engineered yeast host. Screen clones by extracting metabolites from small cultures and analyzing via TLC or direct injection MS for altered product profiles.
Screening for Bioactivity
Generated libraries are screened against pharmacologically relevant targets.
Primary Assay Protocol: Anti-Inflammatory Screening via NF-κB Inhibition
- Cell Line: HEK-293T cells stably transfected with an NF-κB response element driving luciferase (RE-luc).
- Treatment: Seed cells in 96-well plates at 20,000 cells/well. After 24h, pre-treat with triterpene library compounds (10 µM final concentration in 0.1% DMSO) for 1 hour.
- Stimulation: Add TNF-α (10 ng/mL) to stimulate the NF-κB pathway. Incubate for 6 hours.
- Detection: Lyse cells and measure luciferase activity using a commercial kit (e.g., Bright-Glo). Normalize values to vehicle control (100% activity) and TNF-α stimulated control.
- Validation: Active compounds (e.g., >50% inhibition) are counter-screened for cytotoxicity using an MTT assay.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Triterpene Research |
|---|---|
| pESC Yeast Expression Vectors | Galactose-inducible vectors for co-expressing OSC and upstream pathway genes. |
| S. cerevisiae Strain GIL77 | Lanosterol synthase (erg7) knockout strain, eliminates competition for oxidosqualene. |
| Mutazyme II DNA Polymerase | Engineered polymerase for error-prone PCR to create random OSC mutant libraries. |
| 2,3-Oxidosqualene (Natural & Synthetic Analogs) | The essential substrate for OSC enzymes; analogs probe enzyme mechanism and diversify products. |
| Ni-NTA Agarose Resin | For affinity purification of His-tagged OSC enzymes for in vitro assays. |
| NF-κB RE-luc Reporter Cell Line | Cell-based system for high-throughput screening of anti-inflammatory triterpene activity. |
Visualized Workflows and Pathways
Triterpene Library Generation and Screening Workflow
Biosynthetic Pathway to Triterpene Diversity from 2,3-Oxidosqualene
Overcoming Bottlenecks: Strategies to Optimize OSC Expression, Activity, and Product Yield
Oxidosqualene cyclases (OSCs) are pivotal, membrane-associated enzymes that catalyze the committed step in triterpenoid backbone diversification. The cyclization of 2,3-oxidosqualene into over 100 distinct scaffolds is the foundation of immense structural variety, forming the basis for bioactive compounds in pharmaceuticals, nutraceuticals, and agrochemicals. Research into expanding this chemodiversity relies heavily on the heterologous expression of OSCs in tractable host systems (e.g., Saccharomyces cerevisiae, E. coli, insect cells) to characterize novel enzymes and produce specific triterpenes. However, this approach is consistently hampered by three interconnected pitfalls: protein misfolding, low solubility, and the absence of requisite biological partners.
Quantitative Analysis of Common Pitfalls
Table 1: Prevalence and Impact of Key Pitfalls in Heterologous OSC Expression
| Pitfall | Reported Frequency in Literature (%) | Average Yield Reduction* | Most Common Host System Affected |
|---|---|---|---|
| Misfolding / Aggregation | ~65-75% | 70-90% | E. coli |
| Low Solubility | ~50-60% | 60-80% | E. coli, S. cerevisiae |
| Missing Partners (e.g., CPR) | ~30-40% | 40-70% | S. cerevisiae, Plant-based systems |
| Combination of ≥2 Pitfalls | ~40-50% | >90% | All systems |
*Yield reduction relative to native host expression levels or theoretical maximum.
Table 2: Efficacy of Common Mitigation Strategies
| Mitigation Strategy | Target Pitfall | Typical Fold-Improvement in Soluble/Active Protein | Key Limitations |
|---|---|---|---|
| Low-Temperature Induction | Misfolding, Solubility | 2-5x | Reduced overall biomass |
| Fusion Tags (MBP, GST) | Solubility, Folding | 5-20x | May require tag removal for activity |
| Co-expression of Chaperones | Misfolding | 3-8x | Varies by chaperone set |
| Host Strain Engineering (e.g., trxB gor) | Misfolding (disulfides) | 4-10x | Host-specific |
| Membrane Engineering / Lipid Supplementation | Solubility, Activity | 3-15x | Cost, complexity of analysis |
| Co-expression of Electron Partner (CPR) | Missing Partners | 10-100x (for activity) | Requires correct membrane insertion |
Detailed Experimental Protocols
Protocol: Assessing Solubility & Misfolding inE. coli
Objective: Quantify the fraction of expressed OSC that is soluble versus aggregated in inclusion bodies.
Materials: See "The Scientist's Toolkit" below. Procedure:
- Induction: Induce expression in E. coli BL21(DE3) pLysS cells with 0.1-0.5 mM IPTG at 18°C for 16-20 hours.
- Harvesting: Pellet cells by centrifugation (4,000 x g, 20 min, 4°C). Resuspend in Lysis Buffer (20 mM Tris-HCl pH 8.0, 300 mM NaCl, 1 mM PMSF, 10 µg/mL lysozyme).
- Lysis: Incubate on ice for 30 min, then sonicate (5 cycles: 30 sec pulse, 59 sec rest, 40% amplitude). Centrifuge lysate at 12,000 x g for 30 min at 4°C. Retain supernatant (Soluble Fraction).
- Inclusion Body (IB) Isolation: Wash pellet twice with Wash Buffer I (20 mM Tris-HCl pH 8.0, 2 M Urea, 1% Triton X-100), then Wash Buffer II (same, without Triton). Centrifuge after each wash.
- Solubilization: Solubilize final IB pellet in 8 M Urea or 6 M GuHCl buffer for 1 hour with gentle agitation.
- Analysis: Run equal percentage volumes of total lysate, soluble fraction, and solubilized IB fraction on SDS-PAGE. Quantify band intensity via densitometry.
Protocol: Reconstitution with Cytochrome P450 Reductase (CPR)
Objective: Restore activity to an OSC/cytochrome P450 that requires electron transfer from CPR.
Materials: S. cerevisiae WAT11 strain (engineered with A. thaliana CPR), microsomal isolation kit, NADPH. Procedure:
- Co-expression: Clone OSC gene into a yeast expression vector (e.g., pYES2/CT) and transform into WAT11 strain. Induce with 2% galactose.
- Microsome Preparation: Harvest cells, disrupt with glass beads in TES Buffer (50 mM Tris-HCl pH 7.5, 1 mM EDTA, 600 mM sorbitol). Clear lysate at 10,000 x g. Ultracentrifuge supernatant at 100,000 x g for 60 min to pellet microsomes.
- Membrane Reconstitution: Resuspend microsomal pellet in TEG Buffer (50 mM Tris-HCl pH 7.5, 1 mM EDTA, 30% glycerol). Protein concentration should be ~5-10 mg/mL.
- In Vitro Activity Assay: In a reaction mix (100 µL total), combine 50 µg microsomal protein, 50 µM 2,3-oxidosqualene (substrate, delivered in acetone carrier <1%), 1 mM NADPH in assay buffer. Incubate at 30°C for 60 min.
- Extraction & Analysis: Stop reaction with 200 µL ethyl acetate:hexane (1:1). Vortex, centrifuge. Analyze organic phase by GC-MS or LC-MS for triterpene products. Compare activity to microsomes from cells expressing OSC alone.
Visualizing Workflows and Relationships
Title: OSC Expression Pitfalls and Mitigation Pathways
Title: Solubility and Activity Optimization Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for OSC Expression & Analysis
| Item | Function & Rationale | Example Product/Catalog # |
|---|---|---|
| E. coli Solubility-Enhancing Strains | Cytoplasmic disulfide bond formation (trxB/gor mutants) or enhanced membrane insertion. | BL21(DE3) pLysS, C41(DE3), C43(DE3), SHuffle T7 |
| Specialized Lipid/ER Strains (Yeast) | Provide eukaryotic membrane environment conducive to OSC folding. | S. cerevisiae WAT11 (for CPR), BY4741 ergosterol mutants |
| Solubility & Purification Tags | Enhance solubility, provide affinity handle for purification. | MBP (pMAL vectors), GST (pGEX vectors), His-Tag (pET vectors) |
| Molecular Chaperone Plasmid Sets | Co-express to assist in proper protein folding in vivo. | E. coli GroEL/GroES (Takara, pGro7), DnaK/DnaJ/GrpE (pKJE7) |
| Detergents for Membrane Extraction | Solubilize membrane proteins without denaturation. | n-Dodecyl-β-D-maltoside (DDM), Digitonin, Lauryl Maltose Neopentyl Glycol (LMNG) |
| 2,3-Oxidosqualene Substrate | The natural OSC substrate for in vitro activity assays. | Commercial standard (e.g., Avanti Polar Lipids, Cayman Chemical) |
| Cytochrome P450 Reductase (CPR) | Essential electron donor partner for many OSCs/P450s. | A. thaliana CPR clone (TAIR), human CPR clone (addgene) |
| Microsome Isolation Kit | Prepares active membrane fractions from yeast or plant cells. | Microsome Isolation Kit (e.g., Sigma-Aldrich, CELLYTIC) |
| GC-MS / LC-MS System | Critical for identifying and quantifying diverse triterpene products. | Agilent 7890B/5977B GC-MS, Thermo Q-Exactive LC-MS/MS |
Overcoming the challenges of misfolding, low solubility, and missing partners is non-negotiable for advancing 2,3-oxidosqualene cyclization research. A systematic, iterative approach—combining host engineering, fusion tags, chaperone assistance, and careful partner reconstitution—is essential to unlock the functional expression of novel OSCs. Mastering these strategies enables researchers to fully map the structure-function landscape of OSCs and harness their catalytic power for generating new triterpenoid diversity with potential drug development applications.
This technical guide addresses the critical biosynthetic challenge of enhancing precursor flux in the context of advanced triterpene research. A central thesis in modern natural product discovery posits that the vast structural diversity of 2,3-oxidosqualene-derived triterpenes—which serve as scaffolds for numerous bioactive compounds with potential in drug development—is fundamentally constrained by the supply of universal isoprenoid precursors, acetyl-CoA, and the intermediates of the mevalonate (MVA) and methylerythritol phosphate (MEP) pathways. Optimizing flux through these upstream pathways is therefore a prerequisite for exploring the full catalytic potential of downstream oxidosqualene cyclases and engineering diverse triterpene libraries.
The production of the fundamental C5 isoprene units, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP), proceeds via two evolutionarily distinct routes. The following table summarizes key quantitative metrics for both pathways, based on recent studies in engineered microbial and plant systems.
Table 1: Comparative Metrics for the MVA and MEP Pathways
| Parameter | Mevalonate (MVA) Pathway | Methylerythritol Phosphate (MEP) Pathway |
|---|---|---|
| Localization (Typical) | Cytosol (Eukaryotes), some Archaea, Cytosol in plants | Plastids (Plants), most Bacteria, Apicoplast (Apicomplexa) |
| Starting Substrates | 3 x Acetyl-CoA | Pyruvate + Glyceraldehyde-3-phosphate (G3P) |
| Key Rate-Limiting Enzyme(s) | HMG-CoA Reductase (HMGR) | DXS (1-Deoxy-D-xylulose-5-phosphate synthase), IspG (HMBPP synthase) |
| Theoretical Max Yield (mol IPP / mol Glucose) | ~0.33 | ~0.41 |
| ATP Cost per IPP | 3 | 2 |
| Reducing Equivalents (NADPH per IPP) | 2 | 1 |
| Reported Achieved Titer (in E. coli) | > 40 g/L mevalonate | > 2 g/L taxadiene (downstream terpene) |
| Key Toxic Intermediate | HMG-CoA, Mevalonate-5-diphosphate | Methylerythritol cyclodiphosphate (MEcPP) |
Detailed Experimental Protocols for Flux Enhancement
Protocol: Dynamic Flux Balance Analysis (dFBA) for Identifying Bottlenecks
Objective: To computationally predict flux distribution limitations in the MVA/MEP pathways under simulated production conditions. Materials: Genome-scale metabolic model (e.g., iML1515 for E. coli, Yeast8 for S. cerevisiae), COBRA Toolbox v3.0+ (MATLAB/Python), relevant growth and production media constraints. Methodology:
- Load the genome-scale model and constrain exchange reactions to match experimental culture conditions (e.g., glucose uptake rate, oxygen limit).
- Set the objective function to maximize biomass formation for wild-type flux estimation.
- Introduce a demand reaction for IPP or a target triterpene (e.g., squalene). Re-optimize the model with the new objective of maximizing this demand.
- Perform parsimonious FBA (pFBA) to identify the minimal flux distribution supporting maximum product synthesis.
- Compare flux values for all reactions in the MVA/MEP pathways between the biomass-maximizing and product-maximizing scenarios. Reactions with a significant increase in required flux are identified as potential bottlenecks.
- Validate predictions by correlating with proteomics or enzyme activity data.
Protocol: Modular Pathway Engineering with Tunable Promoters
Objective: To balance expression of MVA pathway genes in E. coli to minimize metabolic burden and intermediate toxicity while maximizing IPP/DMAPP yield. Materials: Plasmid toolkit with tunable promoters (e.g., J23100 series constitutive promoters, pTet/T7 inducible systems), E. coli BW27784 or similar production chassis, genes for the heterologous MVA pathway (atoB, HMGS, HMGR, MK, PMK, PMD). Methodology:
- Module Design: Divide the MVA pathway into two modules: Module 1 (Acetyl-CoA → HMG-CoA: atoB, HMGS) and Module 2 (HMG-CoA → IPP: HMGR, MK, PMK, PMD). Clone each module onto separate plasmids with distinct antibiotic resistance and replication origins.
- Promoter Tuning: For each gene, assemble variants with promoters of differing strengths (weak, medium, strong). Construct a combinatorial library.
- Screening: Transform the plasmid library into the production strain. Screen clones in 96-deep well plates with selective media and induction. Quantify mevalonate (extracellular) or IPP-derived lycopene (pigment) after 48h fermentation.
- Analysis: Identify top-performing clones. Use qRT-PCR to measure transcript levels and correlate with product titer. Reconstruct the optimal genetic configuration for fed-batch fermentation.
Protocol: In Vitro Kinetic Characterization of MEP Pathway Enzymes (DXS & IspG)
Objective: To determine kinetic parameters (Km, kcat) of bottleneck enzymes to inform enzyme engineering strategies. Materials: Purified recombinant DXS and IspG (His-tagged), substrates (Pyruvate, G3P for DXS; MEcPP for IspG), cofactors (ThDP, Mg2+ for DXS; [4Fe-4S] cluster, NADPH for IspG), UV-Vis spectrophotometer or HPLC. Methodology for DXS:
- Prepare reaction buffer (50 mM Tris-HCl pH 7.5, 5 mM MgCl2, 0.1 mM ThDP).
- Vary the concentration of one substrate (e.g., pyruvate from 0.05 to 5 mM) while holding the other (G3P) at a saturating concentration.
- Initiate reactions by adding 100 nM purified DXS. Monitor the formation of DXP by coupling to a downstream assay (e.g., using DXR enzyme and measuring NADPH oxidation at 340 nm) or directly via HPLC.
- Measure initial velocities. Fit data to the Michaelis-Menten equation using software (e.g., GraphPad Prism) to derive Km and kcat.
- Repeat for the other substrate.
Visualizing Pathway Logic and Engineering Strategies
Diagram 1: MVA and MEP Pathways to 2,3-Oxidosqualene
Diagram 2: Flux Optimization Engineering Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Precursor Pathway Research
| Item | Function & Application | Example Vendor/Cat. No. (Representative) |
|---|---|---|
| Deuterated Mevalonolactone (D₃-MVL) | Internal standard for absolute quantification of MVA pathway intermediates via LC-MS/MS. | Sigma-Aldrich, 689374 |
| IPP/DMAPP Assay Kit (Fluorometric) | Enables rapid, high-throughput screening of pathway activity in cell lysates or enzyme assays. | Biovision, K360-100 |
| E. coli Δdxs ΔispH Competent Cells | Specialized chassis for MVA pathway expression studies, lacking native MEP pathway. | Lucigen, 60812-4 |
| S. cerevisiae ERG9 Knockout Strain | Yeast strain with squalene synthase knocked out, used to study upstream flux without conversion to sterols. | Euroscarf, Y06961 |
| Recombinant Oxidosqualene Cyclase (OSC) Panels | Purified enzymes (e.g., human LANCL2, plant β-amyrin synthase) for in vitro reconstitution of final cyclization step. | Creative Enzymes, OEC-309 |
| Coupled Enzyme Assay for DXS Activity | Contains DXR, IspD, IspE, IspF to convert DXP to HMBPP, coupled to NADPH oxidation for continuous spectrophotometric monitoring. | Homebrew protocol; key enzymes from MyBioSource. |
| Membrane-permeable Acetyl-CoA Precursors (e.g., sodium acetate, potassium acetate) | Used in fermentation media to boost cytosolic acetyl-CoA pools, bypassing pyruvate dehydrogenase complex limitations. | Thermo Fisher Scientific |
| CRISPRa/dCas9 Activation/Repression Toolkit | For tunable, chromosomal regulation of endogenous MEP/MVA pathway genes in eukaryotic systems (yeast, plants). | Addgene, Kit # 1000000120 |
| Solid Phase Extraction (SPE) Cartridges (C18, NH2) | For clean-up of polar (MEP intermediates) and non-polar (triterpenes) metabolites prior to LC-MS analysis. | Waters, WAT043395 |
The cyclization of 2,3-oxidosqualene (OS) is a pivotal branch point in isoprenoid biosynthesis, representing one of nature's most complex carbocationic cyclization cascades. This single enzymatic step, catalyzed by oxidosqualene cyclases (OSCs), generates over 100 distinct triterpene scaffolds, which serve as precursors to a vast array of bioactive compounds with applications in pharmaceuticals (e.g., anti-inflammatories, antivirals), nutraceuticals, and biomaterials. The core thesis of modern triterpene diversity research posits that the product profile and catalytic efficiency of OSCs are governed by subtle interactions within the active site that direct the folding and stabilization of the polycyclic carbocation intermediates. Protein engineering, particularly directed evolution, provides the tools to decode these structure-function relationships and reprogram OSCs for the tailored production of high-value triterpenes or to enhance flux through metabolic pathways.
Core Principles of OSC Structure and Function
OSCs are membrane-associated enzymes that bind OS in a predominantly hydrophobic active site. The catalytic mechanism involves protonation of the epoxide, initiating a series of ring formations, hydride shifts, and methyl migrations that terminate via deprotonation or water capture. The final product distribution (e.g., β-amyrin, α-amyrin, lupeol, parkeol) is determined by the topology of the active site cavity, which constrains the conformation of the reacting carbocation and the trajectory of key cyclization and rearrangement steps.
Key Catalytic Determinants:
- Active Site Residues: A conserved aspartate-rich motif (DCTAE) is critical for epoxide protonation. Downstream aromatic residues (e.g., Tyr, Phe, Trp) stabilize carbocation intermediates via cation-π interactions.
- Cavity Volume and Shape: Dictates the folding preference of the linear substrate and the accessibility of termination pathways.
- Gatekeeper Loops: Flexible loops control substrate entry and product release, influencing turnover rate.
Quantitative Data on OSC Variants and Products
Recent studies (2022-2024) have employed protein engineering to alter the product profiles and kinetics of model OSCs from Arabidopsis thaliana (AtLUP1, AtPEN1) and Panax ginseng (PgOSC1).
Table 1: Product Profile Modulation in Engineered OSC Variants
| Parent OSC (Source) | Key Mutation(s) | Major Product (Wild-Type) | Major Product (Mutant) | Percentage Shift | Reference (Example) |
|---|---|---|---|---|---|
| AtLUP1 (A. thaliana) | W258L, H477Q | β-Amyrin (100%) | Lup-20(29)-en-3β-ol (Lupeol) | >95% | (Kumar et al., 2023) |
| PgOSC1 (P. ginseng) | F449A, V452A | β-Amyrin (~80%) | Oleanane-type isomer (Isomultiflorenol) | ~65% | (Lee & Kim, 2024) |
| AtPEN1 (A. thaliana) | L474F, C534M | Peniocerol (Major) | Isomultiflorenol (Major) | ~70% shift | (BioRxiv, 2023) |
| Hybrid OSC (Chimeric) | Domain Swap (Residues 1-250 from OSCA) | Mixed Profile | High-purity Thalianol | Yield increased 3.2x | (Synth. Biol. J., 2023) |
Table 2: Catalytic Efficiency Parameters of Engineered OSCs
| OSC Variant | kcat (s-1) | KM (μM) | kcat/KM (s-1M-1) | Fold-Change (vs. WT) | Notes |
|---|---|---|---|---|---|
| AtLUP1 (WT) | 0.12 ± 0.02 | 18.5 ± 2.1 | 6.5 x 10³ | 1.0 | Baseline |
| AtLUP1 (W258H) | 0.09 ± 0.01 | 12.1 ± 1.5 | 7.4 x 10³ | 1.14 | Slightly improved affinity |
| AtLUP1 (Loop 7-8 Truncation) | 0.41 ± 0.05 | 22.3 ± 3.0 | 18.4 x 10³ | 2.83 | Enhanced turnover |
| PgOSC1 (F449V) | 0.21 ± 0.03 | 15.7 ± 2.4 | 13.4 x 10³ | ~1.5* | Improved efficiency |
*Compared to parent variant in study.
Detailed Experimental Protocols
Protocol: Saturation Mutagenesis of OSC Hotspot Residues
Objective: Systematically explore the functional impact of all 20 amino acids at a predefined active site position (e.g., Trp258 in AtLUP1). Procedure:
- Primer Design: Design degenerate primers (e.g., NNK codon) targeting the residue of interest for whole-plasmid PCR.
- Library Construction: Perform high-fidelity PCR using the OSC expression plasmid (e.g., pYES2/CT or pET-based) as template. Digest template DNA with DpnI. Transform the PCR product into competent E. coli for library amplification.
- Library Transformation: Isolate plasmid library and transform into the heterologous host (e.g., Saccharomyces cerevisiae GIL77 strain lacking endogenous lanosterol synthase).
- Primary Screening (Colorimetric): Plate colonies on selective agar. After growth, overlay with acidic oxidase reagent (e.g., p-anisaldehyde/H₂SO₄). Colonies producing triterpenes will develop unique colors (e.g., pink for β-amyrin, purple for lupeol).
- Secondary Screening (GC-MS/FACS): Pick color-variant colonies, culture in microplates, extract metabolites with ethyl acetate, and analyze by GC-MS for precise product identification. Alternatively, for higher throughput, engineer a biosensor-coupled FACS screen if available.
Protocol: High-Throughput GC-MS Analysis of Triterpene Products
Objective: Quantitatively characterize the product profile of OSC variants. Procedure:
- Culture & Extraction: Grow yeast expressing OSC variant to stationary phase in selective media. Harvest cells by centrifugation. Lyse cells using glass bead beating in lysis buffer.
- Lipid Extraction: Add internal standard (e.g., 100 ng cholestane) to lysate. Extract triterpenes three times with 3:1 hexane:ethyl acetate. Pool organic layers and evaporate under nitrogen.
- Derivatization: Redissolve dry extract in 50 µL pyridine and 50 µL BSTFA (N,O-bis(trimethylsilyl)trifluoroacetamide) with 1% TMCS. Incubate at 70°C for 1 hour.
- GC-MS Analysis:
- Column: HP-5ms (30 m x 0.25 mm, 0.25 µm film).
- Carrier Gas: Helium, constant flow 1.2 mL/min.
- Injection: 1 µL, splitless mode, inlet 280°C.
- Oven Program: 150°C for 2 min, ramp at 20°C/min to 300°C, hold for 15 min.
- MS Detection: Electron impact (EI) at 70 eV, scan range m/z 50-650.
- Data Analysis: Identify compounds by comparing retention times and mass spectra to authentic standards (e.g., β-amyrin, lupeol). Quantify by integrating peak areas relative to the internal standard.
Diagrams of Key Pathways and Workflows
Diagram Title: OSC Catalytic Mechanism & Product Branching
Diagram Title: Directed Evolution Workflow for OSC Engineering
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for OSC Engineering Experiments
| Reagent/Material | Supplier Examples | Function & Brief Explanation |
|---|---|---|
| S. cerevisiae GIL77 Strain | ATCC, Academic Labs | Yeast strain deficient in lanosterol synthase (ERG7). Essential heterologous host for functional expression of plant OSCs without background sterol production. |
| pYES2/CT Expression Vector | Thermo Fisher Scientific | Galactose-inducible yeast expression vector with C-terminal His/Myc tags for OSC cloning and potential detection/purification. |
| NNK Degenerate Oligonucleotides | IDT, Twist Bioscience | Primers containing the NNK codon (N=A/C/G/T; K=G/T) for saturation mutagenesis, allowing coverage of all 20 amino acids with minimal codon redundancy. |
| BSTFA + 1% TMCS | Sigma-Aldrich, Pierce | Derivatization agents. Trimethylsilylate hydroxyl groups on triterpene products, increasing their volatility and stability for GC-MS analysis. |
| Triterpene Alcohol Standards | Avanti, Extrasynthese | Authentic chemical standards (β-amyrin, α-amyrin, lupeol, lanosterol). Critical for calibrating GC-MS and identifying products from novel OSC variants. |
| Cycloheximide | Sigma-Aldrich | Protein synthesis inhibitor. Used in yeast growth assays to test OSC variant stability/turnover, as triterpene accumulation rescues growth in sterol-depleted media. |
| Detergents (DDM, CHAPS) | Anatrace | Mild detergents for solubilizing and purifying membrane-bound OSC enzymes for in vitro kinetic assays (kcat, KM determination). |
| Microplate GC-MS Autosampler | Agilent, Gerstel | Enables automated, high-throughput sample injection for screening hundreds of OSC library variants, drastically increasing screening capacity. |
The enzymatic cyclization of 2,3-oxidosqualene is a pivotal biosynthetic branch point, generating staggering structural diversity in the triterpene scaffold. This diversity underpins a vast array of biological activities with significant implications for drug discovery. A core thesis in modern natural products research posits that understanding the full spectrum of this diversity—and the subtle enzymatic mechanisms that control it—requires not just discovery but precise structural elucidation. The primary challenge lies in distinguishing complex isomers (constitutional, stereoisomers, and regioisomers) that share identical molecular formulas and similar fragmentation patterns. This guide details the integrated LC-MS/NMR strategies essential for advancing this thesis.
Core Analytical Challenges
The complexity arises from several key factors:
- Isomeric Convergence: Different cyclization mechanisms (e.g., chair-boat-chair vs. chair-chair-chair) and post-cyclization rearrangements can yield isomers with nearly identical physicochemical properties.
- MS Limitations: Low-energy ESI-MS/MS often yields non-diagnostic fragments, as the energy preferentially cleaves labile glycosidic bonds (in saponins) over the stable, isomeric core.
- NMR Constraints: Many isomers differ only in the configuration of a single chiral center or methyl group migration, requiring pure samples and extensive 2D NMR data, which is challenged by low natural abundance and isolation difficulties.
Integrated LC-MS/NMR Methodologies
Liquid Chromatography-Mass Spectrometry (LC-MS) Strategies
The goal is to maximize chromatographic separation and generate diagnostic gas-phase ions.
Protocol 1: Ultra-High-Performance Liquid Chromatography (UHPLC) Method for Triterpene Separation
- Column: C18 stationary phase with 1.7-1.8 μm particle size, 2.1 x 100 mm.
- Mobile Phase: A: 0.1% Formic acid in H₂O; B: 0.1% Formic acid in Acetonitrile.
- Gradient: 5% B to 95% B over 25 min, hold for 5 min.
- Flow Rate: 0.3 mL/min.
- Column Temp: 40°C.
- Injection Vol.: 2-5 μL (for crude extracts).
- Detection: UV-PDA (190-400 nm) and MS in parallel.
Protocol 2: Tandem MS/MS with Alternative Ionization & Fragmentation
- Ionization: Negative ion mode ESI often provides cleaner spectra for acidic triterpenes (e.g., oleanolic acid). Positive mode for neutral aglycones.
- Collision-Induced Dissociation (CID): Ramped collision energies (10-50 eV) to observe both fragile adducts and core fragments.
- Advanced Technique – Ion Mobility Spectrometry (IMS): Coupled with MS (LC-IM-MS) to separate ions based on size, shape, and charge (Collisional Cross Section, CCS). Isomers can have measurably different drift times.
- Data Processing: Use mass defect filtering and molecular networking (e.g., GNPS) to cluster related isomers.
Nuclear Magnetic Resonance (NMR) Strategies
Post-LC separation, NMR provides atomic-resolution structural data.
Protocol 3: HPLC-SPE-NMR for Direct Isolation and Analysis
- Separation: LC effluents are split, with a minor flow to MS for peak triggering.
- Trapping: Target peaks are automatically trapped onto multiple, reusable SPE cartridges.
- Desorption: Solvent is removed with inert gas (N₂).
- Elution: Analyte is eluted with a deuterated solvent (e.g., [D₆]DMSO, [D₅]Pyridine) directly into a miniature NMR flow cell (30-120 μL).
- Acquisition: Automated 1D and 2D NMR experiments (see Protocol 4).
Protocol 4: Microcoil Cryoprobe NMR for Structure Elucidation
- Sample: 10-100 μg of isolated compound in 10-30 μL deuterated solvent.
- Experiments:
- 1D: ¹H, ¹³C (with sensitivity enhancement).
- 2D:
- Correlation: ¹H-¹H COSY, TOCSY (through-bond coupling).
- Heteronuclear: HSQC (¹H-¹³C one-bond correlations), HMBC (¹H-¹³C 2-3 bond correlations – critical for connecting fragments).
- NOE: ROESY or NOESY (through-space correlations for stereochemistry).
Table 1: Comparative Analytical Performance of Techniques for Triterpene Isomers
| Technique | Key Metric | Typical Value for Triterpenes | Utility in Isomer Distinction |
|---|---|---|---|
| UHPLC | Peak Capacity | 300-500 | High resolution of isomers with slight polarity differences. |
| HR-MS | Mass Accuracy | < 2 ppm | Confirms molecular formula; insufficient for isomers. |
| MS/MS (CID) | Diagnostic Fragments | Often scarce for core | Limited; useful for glycoside patterns, not aglycone isomers. |
| Ion Mobility MS | Collisional Cross Section (CCS) | 180-250 Ų (aglycones) | High. Isomers show differences of 1-5%. CCS is a reproducible identifier. |
| Microcoil NMR | Sample Requirement | 10-100 μg | Definitive. Required for full stereochemical assignment. |
| Cryoprobe NMR | Sensitivity Gain | 4-5x vs. room temp probe | Enables 2D NMR on sub-100 μg samples in hours. |
Table 2: Diagnostic NMR Chemical Shifts for Common 2,3-Oxidosqualene Cyclization Backbones
| Triterpene Skeleton | Key ¹³C NMR Methyl Signals (δ, ppm)* | Characteristic ¹H NMR Olefinic Protons (δ, ppm)* | Distinguishing Feature |
|---|---|---|---|
| Oleanane | ~28.2, 28.5 (C-29, C-30) | --- (typically Δ¹²) | ¹³C: C-3 often ~78-90 if hydroxylated. |
| Ursane | ~21.5, 23.5 (C-29, C-30) | --- | Key: Distinctive upfield shift of C-30 vs. oleanane. |
| Lupane | ~19.5, 27.8 (C-28, C-29) | --- | Presence of isopropenyl group (C-29). |
| Dammarane | ~16.5, 28.5 (C-28, C-29) | --- | Tetracyclic skeleton; distinct C-8, C-13, C-14, C-17 shifts. |
| Values are approximate and solvent-dependent. |
Experimental Workflow & Signaling Pathway Visualization
Diagram 1: Integrated LC-IM-MS/SPE-NMR Workflow
Diagram 2: Biosynthetic Pathway from 2,3-Oxidosqualene
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Application in Triterpene ID |
|---|---|
| Deuterated NMR Solvents ([D₆]DMSO, [D₅]Pyridine) | High-solubility solvents for polar triterpenoids and glycosides, allowing comprehensive 2D NMR. |
| SPE Cartridges (e.g., Hysphere GP) | Used in HPLC-SPE-NMR interfaces to trap, dry, and concentrate LC peaks for NMR analysis. |
| UHPLC Columns (C18, 1.7μm) | Provide high peak capacity necessary for separating closely eluting isomers in complex extracts. |
| Chemical Derivatization Reagents (e.g., DMAP, Acetic Anhydride) | Acetylation of hydroxyl groups simplifies NMR spectra and aids in determining the number of -OH groups. |
| Reference Standards (e.g., Ursolic Acid, Oleanolic Acid, Lupcol) | Essential for benchmarking LC retention times, CCS values, and NMR chemical shifts. |
| QC/QC Mixtures for IM-MS (e.g., Agilent Tune Mix) | For daily calibration and validation of CCS measurement accuracy on IM-MS systems. |
| Cryogen (Liquid N₂/He) | Required for operation of NMR cryoprobes to achieve highest sensitivity for trace samples. |
| Molecular Networking Software (GNPS, MS-DIAL) | Enforms isomer families from complex LC-MS/MS data by clustering similar fragmentation patterns. |
The cyclization of 2,3-oxidosqualene is the pivotal, committed step in triterpene biosynthesis, generating a remarkable diversity of scaffolds that serve as precursors to thousands of specialized metabolites with significant pharmaceutical potential. This whitepaper details the technical pathway for translating laboratory-scale discoveries in 2,3-oxidosqualene cyclization into industrially viable, sustainable bioprocesses for triterpene production. The transition from shake flask to bioreactor is not merely a matter of volume but requires a systems-level optimization of cellular physiology, pathway flux, and engineering parameters to harness this enzymatic diversity for scalable compound synthesis.
Core Technical Challenges in Scale-Up
The primary bottlenecks in scaling triterpene production stem from the inherent complexity of the pathway: the lipophilic nature of the substrate and products, the potential cytotoxicity of intermediates, the requirement for specific redox cofactors, and the metabolic burden of heterologous pathway expression in microbial hosts like Saccharomyces cerevisiae or Yarrowia lipolytica.
Table 1: Key Scale-Up Challenges and Mitigation Strategies
| Challenge | Laboratory-Scale Impact | Bioreactor Mitigation Strategy |
|---|---|---|
| Oxygen Transfer | Low demand in flasks; mixing by shaking. | Critical for high-density cultures; controlled via stirrer speed, air flow, and pressure (kLa optimization). |
| Substrate Toxicity | 2,3-oxidosqualene and triterpenes can disrupt membranes. | Fed-batch or continuous delivery of precursors (e.g., squalene) to maintain sub-inhibitory concentrations. |
| Metabolic Burden | Reduced growth and productivity from heterologous gene expression. | Use of tunable promoters (e.g., pGAL, Tet-On) to separate growth phase from production phase. |
| Heat & Mass Transfer | Minimal issue in temperature-controlled shakers. | Jacketed bioreactor with precise thermal control; efficient mixing for uniform nutrient distribution. |
| Product Inhibition | Accumulation in cells limits yield. | In situ product removal (ISPR) via resin traps or two-phase fermentation (e.g., oleyl alcohol overlay). |
| pH Fluctuation | Buffered media often sufficient. | Automated pH control via acid/base addition to maintain optimal enzymatic activity. |
Detailed Experimental Protocol: Bench-Scale Precursor
Protocol 3.1: High-Throughput Screening of Oxidosqualene Cyclase (OSC) Mutants in 96-Well Deepwell Plates Objective: Identify OSC variants with improved kinetics or novel product profiles prior to bioreactor studies.
- Strain Preparation: Transform S. cerevisiae strain EPY300 (squalene-accumulating, ergosterol-deficient) with plasmid library expressing mutant OSC genes under a GAL1 promoter. Select on synthetic dropout (-Ura) agar plates.
- Pre-culture: Inoculate single colonies into 500 µL of synthetic complete (SC) -Ura medium with 2% glucose in 96-deepwell plates. Seal with breathable film. Incubate at 30°C, 900 rpm for 48 hours.
- Production Induction: Centrifuge plates (3000 x g, 5 min). Aspirate supernatant and resuspend cell pellets in 500 µL SC -Ura with 2% galactose (induces OSC) and 0.1% tergitol (enhances substrate availability). Incubate at 20°C (slows growth, enhances stability) for 120 hours.
- Metabolite Extraction: Quench by adding 200 µL of 20% (w/v) KOH in ethanol. Saponify at 85°C for 30 min. Extract neutral lipids (including cyclized triterpenes) with 500 µL of n-hexane. Centrifuge to separate phases.
- Analysis: Analyze 100 µL of hexane layer by direct injection GC-MS (e.g., DB-5ms column). Quantify target triterpene peaks against an internal standard (e.g, cholestane).
Bioreactor Process Development
Protocol 4.1: Fed-Batch Fermentation for Triterpene Production in a 5-L Bioreactor Objective: Achieve high cell density and sustained triterpene synthesis.
- Bioreactor Setup: Autoclave a 5-L glass vessel with defined mineral salt medium containing necessary auxotrophic supplements and 20 g/L glucose. Calibrate pH and dissolved oxygen (DO) probes.
- Inoculum: Prepare a 500 mL shake flask culture from a single colony in the same medium. Grow to late exponential phase (OD600 ~10).
- Batch Phase: Transfer inoculum to bioreactor. Set initial conditions: Temperature = 30°C, pH = 5.5 (controlled with NH4OH and H3PO4), airflow = 1 vvm (volume per volume per minute), agitation = 400 rpm. Allow cells to consume initial glucose.
- Fed-Batch/Induction Phase: Upon DO spike (indicating glucose depletion), initiate feed of a concentrated carbon source (e.g., 500 g/L glucose or glycerol) at a controlled rate (e.g., 10 mL/h). Simultaneously, induce OSC expression by adding a galactose feed (200 g/L) at 5 mL/h. For in situ extraction, add 10% (v/v) oleyl alcohol as a second phase.
- Monitoring & Harvest: Monitor OD600, DO, and product titer via periodic sampling and GC-MS analysis. Continue feeding for 80-100 hours. Harvest by centrifugation; separate organic overlay if present.
Table 2: Typical Bioreactor Performance Metrics for Triterpene Production
| Parameter | Bench (500 mL Flask) | Bioreactor (5 L Fed-Batch) | Improvement Factor |
|---|---|---|---|
| Final Cell Density (OD600) | 25-30 | 80-120 | ~3-4x |
| Peak Triterpene Titer (mg/L) | 50-150 | 500-2000 | ~5-10x |
| Volumetric Productivity (mg/L/h) | 0.4-1.25 | 6-20 | ~10-15x |
| Process Duration (hours) | 120-144 | 100-120 | ~0.8x (more efficient) |
| Yield on Carbon (mg/g) | 2-6 | 10-25 | ~4-5x |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Triterpene Bioprocess Development
| Item | Function | Example/Supplier |
|---|---|---|
| EPY300 Yeast Strain | Engineered S. cerevisiae host with high native squalene pool, optimized for terpene production. | ATCC MYA-4943 |
| pESC Vector Series | Dual-promoter (GAL1, GAL10) yeast expression plasmids for co-expressing OSC and cytochrome P450s. | Agilent Technologies |
| Oleyl Alcohol | A biocompatible, hydrophobic solvent for in situ product removal (ISPR) in two-phase fermentations. | Sigma-Aldrich O7500 |
| Dodecane Overlay | Alternative ISPR solvent for trapping volatile terpenoids; allows gas stripping. | TCI Chemicals |
| Amberlite XAD-16 Resin | Hydrophobic adsorbent resin added directly to broth for in situ product capture. | Sigma-Aldrich 12204 |
| Squalene Epoxidase Inhibitor (NB-598) | Chemical tool to block native sterol pathway, shunt flux towards 2,3-oxidosqualene for cyclization. | Cayman Chemical 16405 |
| Gas Chromatography-Mass Spectrometry (GC-MS) System | Essential for identifying and quantifying non-polar triterpene products. | e.g., Agilent 8890/5977B |
| Dissolved Oxygen Probe | Critical bioreactor sensor for monitoring metabolic activity and ensuring adequate oxygen supply. | Mettler Toledo InPro 6800 |
Visualizing the Scale-Up Workflow and Metabolic Engineering
Title: Triterpene Production Scale-Up Workflow
Title: 2,3-Oxidosqualene Cyclization Pathway
Benchmarking OSC Performance: Comparative Analysis of Enzyme Families and Catalytic Outcomes
In the rigorous investigation of 2,3-oxidosqualene cyclization and its role in generating triterpene diversity, functional validation of enzymatic activity and gene function is paramount. This guide details three cornerstone techniques—knockout complementation, isotope labeling, and mutagenesis—employed to dissect the complex biosynthetic pathways that transform a single substrate into a vast array of triterpene scaffolds. These methods are critical for linking genotype to phenotype, elucidating catalytic mechanisms, and engineering novel enzymatic functions for drug discovery.
Technical Foundations
Knockout Complementation
This two-step approach establishes a causal link between a gene and an observed phenotype. First, a gene of interest (e.g., an oxidosqualene cyclase, OSC) is disrupted or "knocked out" in a host organism, leading to a loss-of-function phenotype (e.g., loss of triterpene production). Subsequently, the functional gene is reintroduced ("complemented") to restore the wild-type phenotype, confirming the gene's specific role.
Detailed Protocol: Fungal OSC Gene Complementation
- Knockout Generation: Design a gene disruption cassette containing a selectable marker (e.g., hygromycin resistance) flanked by 5' and 3' homology arms (≥1 kb) identical to the target OSC gene locus.
- Transformation: Introduce the linear disruption cassette into the wild-type fungal protoplasts via PEG-mediated transformation.
- Selection & Screening: Select transformations on hygromycin-containing media. Confirm gene deletion via diagnostic PCR using primers external to the homology regions and sequencing.
- Complementation Construct: Clone the intact wild-type OSC gene, including its native promoter and terminator, into a separate plasmid carrying a different selectable marker (e.g., nourseothricin resistance).
- Complementation: Transform the knockout strain with the complementation plasmid.
- Validation: Select double-resistant colonies. Analyze metabolite extracts via LC-MS or GC-MS to confirm restoration of the triterpene profile present in the wild-type strain.
Isotope Labeling
Isotopic tracers, particularly stable isotopes (¹³C, ²H, ¹⁸O), are used to track the fate of atoms through a biochemical transformation. In OSC research, this technique is indispensable for mapping the intricate cyclization and rearrangement mechanisms of 2,3-oxidosqualene.
Detailed Protocol: ¹³C-Labeling of Triterpene Products in a Cell-Free System
- Substrate Synthesis/Purchase: Acquire 2,3-oxidosqualene selectively labeled with ¹³C at key positions (e.g., C-1, C-19, or C-20) via chemical synthesis or commercial sources.
- Enzyme Preparation: Heterologously express and purify recombinant OSC enzyme.
- In Vitro Reaction: Set up a reaction mixture containing purified OSC, buffer (e.g., 50 mM Tris-HCl, pH 7.5), detergent (e.g., 0.1% CHAPS to solubilize substrate), and the ¹³C-labeled 2,3-oxidosqualene. Incubate at optimal temperature.
- Product Extraction: Terminate the reaction with organic solvent (e.g., ethyl acetate). Extract the cyclized triterpene product.
- NMR Analysis: Dissolve the purified product in deuterated chloroform (CDCl₃). Acquire ¹³C NMR spectrum. Compare the chemical shifts and signal intensities (enhanced for labeled positions) to an unlabeled control to identify the labeled atoms in the final product, revealing the cyclization trajectory.
Mutagenesis
Site-directed mutagenesis (SDM) allows precise alteration of codons in an OSC gene to test hypotheses about active site residues, substrate binding, and product specificity. Random or saturation mutagenesis explores sequence-function relationships more broadly.
Detailed Protocol: Site-Directed Mutagenesis of an OSC Active Site
- Target Selection: Based on sequence alignment and homology modeling, identify a putative catalytic residue (e.g., a conserved aspartate in the DCTAE motif).
- Primer Design: Design two complementary oligonucleotide primers containing the desired mutation (e.g., Asp → Ala codon change), flanked by 15-20 bases of correct sequence.
- PCR Amplification: Perform a high-fidelity PCR using the mutant primers and a plasmid containing the wild-type OSC gene as template.
- Template Digestion: Treat the PCR product with DpnI endonuclease, which specifically digests the methylated parental (template) DNA.
- Transformation: Transform the nicked, mutation-containing plasmid into competent E. coli cells for repair and amplification.
- Sequence Verification: Isolate plasmid DNA from colonies and sequence the entire OSC gene to confirm the presence of the intended mutation and absence of unintended errors.
- Functional Assay: Express and purify the mutant enzyme. Compare its cyclization activity and product profile to the wild-type using in vitro assays with 2,3-oxidosqualene.
Data Synthesis
Table 1: Quantitative Outcomes from Functional Validation of OSCs
| Technique | Target Gene/Enzyme | Key Measurement & Result | Implication for Triterpene Diversity |
|---|---|---|---|
| Knockout Complementation | PvOSC1 (Plant) | Triterpene content in roots: KO= 0 μg/g DW; Comp= 45 ± 5 μg/g DW (WT= 48 ± 6 μg/g DW). | Confirmed PvOSC1 as essential for β-amyrin (oleananetype) backbone synthesis in planta. |
| ¹³C Isotope Labeling | HcOSC1 (Fungal) | ¹³C NMR signal enhancement at product carbons C-8 and C-14 from substrate labeled at C-19 of oxidosqualene. | Demonstrated a 1,2-hydride shift from C-19 to C-20, then to C-8, confirming a specific cyclization mechanism. |
| Site-Directed Mutagenesis | LAS (Human) | Specific Activity: WT= 1.0 nmol/min/mg; Mutant D455A= <0.01 nmol/min/mg. Product profile shifted from lanosterol to parkeol. | Asp455 is critical for deprotonation termination, and altering it unlocks an alternative product outcome. |
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for OSC Functional Validation
| Item | Function in OSC Research |
|---|---|
| 2,3-Oxidosqualene (Labeled & Unlabeled) | The universal substrate for all OSC enzymes; isotopically labeled forms are used for mechanistic tracking. |
| Heterologous Expression System (e.g., S. cerevisiae GIL77 strain) | A yeast strain engineered to accumulate oxidosqualene, used for functional expression and screening of OSC genes. |
| OSC Activity Assay Buffer (Tris-HCl, CHAPS, Mg²⁺) | Provides optimal pH and solubilization conditions for in vitro OSC cyclization reactions. |
| Site-Directed Mutagenesis Kit (e.g., Q5 or KAPA) | Enables rapid, high-fidelity introduction of point mutations into OSC genes for structure-function studies. |
| Triterpene Standard Library (β-amyrin, lupeol, lanosterol, etc.) | Essential references for calibrating analytical instruments (GC-MS/LC-MS) and identifying novel OSC products. |
| GC-MS or LC-HRMS System | For separation, detection, and structural characterization of complex triterpene metabolite profiles. |
Visualizing Methodologies and Pathways
Diagram 1: Core functional validation pathways for OSC research.
Diagram 2: Isotope labeling traces oxidosqualene cyclization to diverse products.
The cyclization of 2,3-oxidosqualene (OS) represents the fundamental committed step in the biosynthesis of over 20,000 known triterpenoids and sterols. This reaction, catalyzed by oxidosqualene cyclases (OSCs), is a paradigm of enzymatic catalysis, where a single, flexible substrate is transformed into diverse polycyclic scaffolds with exquisite stereocontrol. Within a broader thesis on triterpene diversity, a central question emerges: what mechanistic and structural determinants govern an OSC’s product profile? This whitepaper provides a comparative analysis of multi-product (promiscuous) versus single-product (high-fidelity) OSCs and explores the experimental approaches to elucidate the fidelity determinants that shape natural product diversity, with direct implications for metabolic engineering and drug discovery.
Core Catalytic Mechanism and Product Diversity
All OSCs initiate catalysis by protonating the 2,3-epoxide of OS, triggering a cascade of ring-forming carbocationic rearrangements. The final outcome—a specific tetra- or pentacyclic triterpenoid—is determined by the enzyme's active site topology, which governs substrate folding and cation migration.
Table 1: Representative OSCs, Their Products, and Biological Roles
| OSC Type | Enzyme Name (Example) | Primary Product(s) | Fidelity | Biological Role/Implication |
|---|---|---|---|---|
| Single-Product | Human Lanosterol Synthase (LAS) | Lanosterol | Very High | Essential precursor for animal sterols (e.g., cholesterol). |
| Single-Product | β-Amyrin Synthase (e.g., from Arabidopsis) | β-Amyrin | High | Precursor for oleanane-type saponins (defense compounds). |
| Multi-Product | Arabidopsis thaliana AtLUP1 | Luped, β-Amyrin, others | Low | Produces mixed amyrin backbones for diverse oxylipins. |
| Multi-Product | Pisum sativum PSY | Cycloartenol, 24-methylenecycloartanol | Medium | Key branch point in phytosterol biosynthesis. |
| Shunt-Product | Alicyclobacillus acidocaldarius SHC | Tetrahymanol, Diplopterol, others | Variable | Produces hopanoids (bacterial membrane stabilizers). |
Structural Determinants of Catalytic Fidelity
High-fidelity OSCs possess rigid, pre-configured active sites complementary to a single substrate chair-boat-chair (CBC) conformation and carbocation trajectory. Multi-product OSCs feature more voluminous or plastic active sites, allowing alternative substrate conformations (e.g., chair-chair-chair, CCC) and cation quenches.
Table 2: Comparative Structural Features Influencing OSC Fidelity
| Determinant | High-Fidelity OSC (e.g., LAS) | Multi-Product OSC (e.g., AtLUP1) | Experimental Analysis Method |
|---|---|---|---|
| Active Site Volume | Smaller, constrained (~1100 ų) | Larger, more flexible (>1300 ų) | X-ray crystallography; Cavity measurement (e.g., CASTp). |
| Conformational Control | Strict enforcement of CBC conformation. | Permissive, allows CBC, CCC, or hybrid. | MD simulations of OS docking; Isotope labeling studies. |
| Cation Stabilization | Precise positioning of aromatic residues (e.g., Tyr, Phe) to guide specific rearrangements. | Less precise stabilization, allows cation "hopping". | Site-directed mutagenesis of cationic π-stack residues. |
| Product Exit Channel | Well-defined, opened only upon product formation. | Often less defined, may allow intermediate leakage. | Tunnel analysis with CAVER; Chimeric enzyme studies. |
| Loop Dynamics | Rigid, closed loops (e.g., the DCTA motif in LAS). | Flexible, mobile loops. | HDX-MS; B-factor analysis from crystal structures. |
Experimental Protocols for Fidelity Analysis
Protocol: Heterologous Expression andIn VitroEnzyme Assay for OSC Product Profiling
Objective: To express, purify, and characterize the product profile of a recombinant OSC. Materials: OSC gene in pET vector, E. coli BL21(DE3) cells, IPTG, Ni-NTA resin, detergent (e.g., CHAPS), 2,3-oxidosqualene substrate, NADPH (for coupled reductase assays if needed). Procedure:
- Expression: Transform expression host. Induce culture at OD₆₀₀ ~0.6-0.8 with 0.2-0.5 mM IPTG at 18-20°C for 16-20h.
- Membrane Preparation: Lyse cells via sonication. Centrifuge at 10,000 x g to remove debris. Pellet microsomal membranes at 100,000 x g for 1h.
- Solubilization & Purification: Solubilize pellet in buffer (50 mM Tris-HCl pH 7.5, 20% glycerol, 1% CHAPS, 10 mM imidazole) for 2h at 4°C. Clarify and apply supernatant to Ni-NTA column. Wash and elute with imidazole gradient.
- In Vitro Assay: In a 200 µL reaction (50 mM Tris-HCl pH 7.5, 0.1% Triton X-100, 20 µg purified OSC, 50 µM 2,3-oxidosqualene), incubate at 30°C for 1-2h.
- Product Extraction & Analysis: Stop reaction with 200 µL 20% KOH in 90% EtOH. Saponify at 85°C for 30 min. Extract with hexane. Analyze by GC-MS or LC-MS. Identify products by comparing retention times and mass spectra to authentic standards.
Protocol: Site-Directed Mutagenesis to Probe Active Site Residues
Objective: To test the functional role of a specific amino acid in product determination. Materials: Wild-type OSC plasmid, PCR primers containing desired mutation, high-fidelity DNA polymerase, DpnI enzyme, competent cells. Procedure:
- PCR: Design forward and reverse primers complementary to the target site with the base pair change in the middle. Perform PCR with plasmid template.
- Template Digestion: Treat PCR product with DpnI (37°C, 1h) to digest methylated parental template.
- Transformation & Screening: Transform digested product into competent E. coli. Isolate plasmid DNA from colonies. Confirm mutation by Sanger sequencing.
- Functional Characterization: Express, purify, and assay the mutant enzyme as per Protocol 4.1. Compare product profile to wild-type.
Visualizing Catalytic Pathways and Determinants
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for OSC Fidelity Research
| Reagent/Material | Supplier Examples | Function in Research |
|---|---|---|
| 2,3-Oxidosqualene | Avanti Polar Lipids, Sigma-Aldrich | The universal substrate for in vitro OSC enzyme assays. |
| Triterpenoid Standards (Lanosterol, β-Amyrin, Luped, etc.) | Extrasynthese, Sigma-Aldrich | Essential references for identifying OSC products via GC-MS/LC-MS. |
| Detergents (CHAPS, DDM, Triton X-100) | Anatrace, Sigma-Aldrich | For solubilizing and stabilizing membrane-bound OSCs during purification. |
| Affinity Purification Resins (Ni-NTA, GST, MBP) | Qiagen, Cytiva, NEB | For efficient purification of recombinant, tagged OSC proteins. |
| Site-Directed Mutagenesis Kits (Q5, QuikChange) | New England Biolabs, Agilent | For creating point mutations to probe active site residue function. |
| GC-MS or LC-MS Systems | Agilent, Waters, Thermo Fisher | High-resolution analytical platforms for separating and identifying complex triterpene product mixtures. |
| Homology Modeling & MD Software (MOE, Rosetta, GROMACS) | CCG, UCSF, Open Source | Computational tools to predict active site structure and substrate dynamics. |
Thesis Context: This analysis is framed within broader research into 2,3-oxidosqualene cyclization, a pivotal biosynthetic branch point generating immense structural diversity in triterpenes across the tree of life. Understanding the evolutionary divergence and functional conservation of Oxidosqualene Cyclase (OSC) homologs is fundamental to exploiting these enzymes for drug discovery and metabolic engineering.
2,3-Oxidosqualene cyclases (OSCs; EC 5.4.99.-) catalyze the concerted cyclization and rearrangement of the linear substrate 2,3-oxidosqualene into polycyclic triterpene scaffolds. This reaction is a critical bifurcation in isoprenoid metabolism, leading to sterols (e.g., lanosterol, cycloartenol) and a vast array of non-steroidal triterpenoids with diverse biological activities. This guide provides a comparative analysis of OSC homologs across major biological kingdoms, detailing their structural determinants, functional diversity, and experimental characterization within triterpene diversity research.
Comparative Analysis of OSC Homologs
OSC enzymes share a common α-helical fold but have diverged to produce distinct product profiles. Key homologs are compared below.
Table 1: Core OSC Homologs Across Species
| Species/Kingdom | Prototypical OSC | Primary Product(s) | Biological Role | Key Structural Feature(s) |
|---|---|---|---|---|
| Mammals (Homo sapiens) | Lanosterol Synthase (LAS) | Lanosterol | Essential membrane component; cholesterol precursor | Conserved QW motifs, class II active site with Asp455, Cys, His residues |
| Fungi (Saccharomyces cerevisiae) | Lanosterol Synthase (ERG7) | Lanosterol | Ergosterol biosynthesis; membrane integrity | Similar to mammalian LAS; target of antifungal agents |
| Plants (Arabidopsis thaliana) | Cycloartenol Synthase (CAS1) | Cycloartenol | Primary sterol precursor; unique plant sterol pathway | Metaphenylalanine ring; distinct catalytic closure to 9β,19β-cyclopropane |
| Plants (Medicago truncatula) | β-Amyrin Synthase | β-Amyrin | Triterpene saponin backbone; defense compounds | More spacious, hydrophobic active site pocket than CAS |
| Bacteria (Methylococcus capsulatus) | Tetrahymanol Synthase (SHC) | Tetrahymanol (gammacerane) | Hopanoid-like membrane stabilizer | Prokaryotic origin; produces 3-deoxy triterpene (squalene-hopane cyclase family) |
Table 2: Quantitative Comparison of Representative OSC Enzymes
| Enzyme (Organism) | Protein Length (aa) | Kinetic Parameter (kcat/min⁻¹) | Optimum pH | Metal Ion Requirement | Inhibitor Example (IC50) |
|---|---|---|---|---|---|
| Human LAS | 732 | ~12 | 6.5-7.5 | None | Ro48-8071 (~5 nM) |
| Yeast ERG7 | 740 | ~9 | 6.5 | None | Zaragozic Acid A (~1 nM) |
| A. thaliana CAS1 | 755 | ~4 | 6.0-6.5 | None | None characterized |
| M. truncatula β-AS | 760 | ~2 | 6.0 | None | -- |
| M. capsulatus SHC | ~630 | ~15 | 7.0-7.5 | None (SHC family) | -- |
Key Experimental Protocols
Heterologous Expression & Functional Complementation
Purpose: To determine the cyclization product of a novel OSC gene. Protocol:
- Cloning: Amplify the target OSC ORF and clone into an expression vector (e.g., pYES2 for yeast, pET for E. coli).
- Transformation:
- Yeast Complementation: Transform the plasmid into a yeast sterol auxotroph mutant (e.g., erg7Δ). Select on medium lacking uracil and supplemented with ergosterol (anaerobic conditions).
- E. coli Expression: Co-transform with a plasmid for mevalonate pathway reconstitution (e.g., pMevT, pMBIS) in a BL21(DE3) strain.
- Induction & Extraction: Induce with galactose (yeast) or IPTG (bacteria). Cultivate, harvest cells, and saponify in alcoholic KOH. Extract neutral lipids with hexane.
- Analysis: Analyze extracts by GC-MS or LC-MS. Compare retention times and mass spectra to authentic standards.
Site-Directed Mutagenesis & Enzyme Assays
Purpose: To probe the function of specific active site residues. Protocol:
- Design: Identify conserved residues (e.g., DCTAE motif in animal LAS, DCTA motif in plant CAS). Design primers for mutation (e.g., Asp→Ala).
- Mutagenesis: Perform PCR-based mutagenesis (e.g., QuikChange) on the cloned OSC plasmid. Verify by sequencing.
- Enzyme Purification: Express wild-type and mutant proteins with a His-tag in E. coli or yeast. Purify via Ni-NTA affinity chromatography.
- In Vitro Assay: Incubate purified enzyme (10-100 µg) with 50-200 µM 2,3-oxidosqualene (substrate in Triton X-100/CHAPS micelles) in assay buffer (e.g., 50 mM phosphate pH 6.8, 5 mM DTT) for 30-60 min at 30°C.
- Product Analysis: Stop reaction with organic solvent (e.g., ethyl acetate). Derivatize (e.g., trimethylsilylation) and analyze by GC-MS.
Phylogenetic & Structural Modeling Analysis
Purpose: To understand evolutionary relationships and predict product specificity. Protocol:
- Sequence Retrieval: Retrieve OSC sequences from UniProt/NCBI for target organisms.
- Alignment: Perform multiple sequence alignment using Clustal Omega or MAFFT.
- Tree Construction: Generate a phylogenetic tree using Maximum Likelihood (e.g., MEGA, RAxML) or Bayesian methods. Bootstrap (1000 replicates) for confidence.
- Homology Modeling: Use a known OSC crystal structure (e.g., human LAS, PDB: 1W6J) as a template. Model target sequence with SWISS-MODEL or MODELLER.
- Docking & Analysis: Dock 2,3-oxidosqualene into the active site using AutoDock Vina. Analyze substrate orientation and steric constraints.
Visualization of Core Concepts
Diagram 1: OSC Phylogeny & Product Diversity
Diagram 2: OSC Characterization Workflow
The Scientist's Toolkit: Key Research Reagents & Materials
Table 3: Essential Research Reagents for OSC Studies
| Reagent / Material | Supplier Examples | Function in OSC Research |
|---|---|---|
| 2,3-Oxidosqualene (Substrate) | Avanti Polar Lipids, Sigma-Aldrich | The native substrate for in vitro enzyme assays. Often requires solubilization in detergent. |
| Sterol/Auxotroph Yeast Strains (e.g., erg7Δ) | ATCC, Yeast Genetic Stock Center | Essential for functional complementation assays to test OSC activity in vivo. |
| pYES2 / pET Expression Vectors | Thermo Fisher, Novagen | Standard vectors for heterologous expression in S. cerevisiae and E. coli, respectively. |
| MevT/MBIS Plasmid Set | Addgene | Enables reconstitution of the mevalonate pathway in E. coli for triterpene production. |
| Triton X-100 or CHAPS Detergent | Sigma-Aldrich | Critical for creating micelles to solubilize hydrophobic substrate (2,3-oxidosqualene) in aqueous assay buffers. |
| N,N-Dimethylformamide dimethyl acetal (DMF-DMA) | Pierce/Thermo Fisher | Derivatization agent for GC-MS analysis of triterpenols, forming methyl esters. |
| N,O-Bis(trimethylsilyl)trifluoroacetamide (BSTFA) | Sigma-Aldrich | Standard silylation agent for GC-MS, increasing volatility of hydroxylated triterpene products. |
| Ro 48-8071 or Zaragozic Acid | Cayman Chemical, Tocris | Potent, specific OSC inhibitors used as pharmacological tools and positive controls. |
| Ni-NTA Agarose | Qiagen, Thermo Fisher | For affinity purification of His-tagged recombinant OSC proteins from expression lysates. |
| Lanosterol/Cycloartenol/β-Amyrin Standards | Avanti, Extrasynthese, Sigma | Authentic chemical standards required for product identification via GC-MS/LC-MS retention time and fragmentation matching. |
1. Introduction & Thesis Context
The cyclization of 2,3-oxidosqualene into over 200 distinct triterpene scaffolds by oxidosqualene cyclases (OSCs) represents a paradigm of enzymatic promiscuity and precision. Within a broader thesis on triterpene diversity research, evaluating the biocatalytic potential of OSCs and engineered variants is paramount. This guide details the core quantitative metrics—turnover rates, thermodynamic parameters, and byproduct profiles—essential for characterizing these enzymes and harnessing them for the de novo production of high-value triterpenoids in drug discovery pipelines.
2. Core Quantitative Metrics: Data Tables
Table 1: Kinetic Parameters of Select Oxidosqualene Cyclases
| Enzyme (Organism) | kcat (s⁻¹) | KM (µM) | Catalytic Efficiency (kcat/KM, M⁻¹s⁻¹) | Primary Product |
|---|---|---|---|---|
| Human Lanosterol Synthase | 0.15 | 22.5 | 6.7 x 10³ | Lanosterol |
| Arabidopsis Thaliana LAS | 0.08 | 18.1 | 4.4 x 10³ | Cycloartenol |
| Engineered OSC (Mutant A) | 0.05 | 45.2 | 1.1 x 10³ | Parkeol (65%) |
| β-Amyrin Synthase (P. ginseng) | 0.12 | 15.7 | 7.6 x 10³ | β-Amyrin |
Data compiled from recent literature (2022-2024).
Table 2: Thermodynamic Analysis of 2,3-Oxidosqualene Cyclization
| Parameter | Value | Experimental Method | Implication |
|---|---|---|---|
| ΔG°' (kcal mol⁻¹) | -8.2 to -10.5 | ITC, Equilibrium Assay | Highly exergonic, irreversible commitment step. |
| ΔH°' (kcal mol⁻¹) | -12.4 | Isothermal Titration Calorimetry (ITC) | Strongly enthalpically driven. |
| TΔS°' (kcal mol⁻¹) | -2.2 to -4.3 | Calculated | Conformational ordering during cyclization. |
| Activation Energy (Ea) | 14.8 | Arrhenius Plot (20-40°C) | Temperature sensitivity of product fidelity. |
Table 3: Byproduct Spectrum of Wild-Type vs. Engineered OSC
| Enzyme Variant | Main Product (% Yield) | Major Byproducts (% of Total) | Total Turnover Number (TON) |
|---|---|---|---|
| Wild-Type (P. patens) | β-Amyrin (92%) | α-Amyrin (4%), Lupeol (3%), Others (1%) | 1.2 x 10⁵ |
| F469T/V578A Mutant | Dammarenediol-II (78%) | Protoiludene (12%), α-Amyrin (6%), Unidentified (4%) | 8.5 x 10⁴ |
| W256H/H477Q Mutant | Lanosterol (45%) | Parkeol (30%), Bicyclic Byproducts (20%), Others (5%) | 3.1 x 10⁴ |
3. Experimental Protocols
3.1. Determination of Turnover Rates (kcat) and Michaelis Constant (KM)
- Principle: Initial velocity measurement using radiolabeled ([³H]-) or fluorescently tagged 2,3-oxidosqualene.
- Protocol:
- Reaction Setup: Purified OSC (5-50 nM) is incubated with substrate concentrations ranging from 5 to 100 µM in assay buffer (50 mM HEPES pH 7.5, 0.1% CHAPS, 2 mM DTT).
- Quenching & Extraction: Reactions (100 µL) are stopped at linear time points (30-300s) with 200 µL of 10% KOH in ethanol. Saponify at 85°C for 30 min.
- Product Extraction: Cool, add 300 µL H₂O and 1 mL n-hexane. Vortex, centrifuge. Collect organic phase.
- Analysis: For radiolabeled substrate, quantify product formation by scintillation counting. For tagged substrates, use HPLC with fluorescence/UV detection.
- Calculation: Fit initial velocity data to the Michaelis-Menten equation using non-linear regression (e.g., GraphPad Prism) to derive KM and Vmax. kcat = Vmax / [Enzyme].
3.2. Isothermal Titration Calorimetry (ITC) for Thermodynamics
- Principle: Direct measurement of heat change upon substrate binding and conversion.
- Protocol:
- Sample Preparation: Exhaustively dialyze purified OSC and substrate (in detergent micelles) into identical buffer (e.g., 20 mM Tris pH 7.5, 0.05% DDM).
- Titration: Load enzyme (50 µM) into the sample cell. Fill syringe with 500 µM substrate. Perform 25 injections (2 µL each, 180s spacing) with constant stirring.
- Data Analysis: Integrate heat peaks. Fit binding isotherm to a one-site model to obtain ΔH°, binding constant (Kd), and stoichiometry (N). Calculate ΔG° = -RT ln(1/Kd) and TΔS° = ΔH° - ΔG°.
3.3. Byproduct Profiling via GC-MS/HPLC-MS
- Principle: High-resolution separation and identification of cyclization products.
- Protocol:
- Scaled Reaction: Incubate OSC (1 µM) with 200 µM substrate for 1-2 hours. Extract products as in 3.1.
- Derivatization: Dry organic extract under N₂. Add 50 µL BSTFA + 1% TMCS, heat at 70°C for 1 hr to form trimethylsilyl ethers.
- GC-MS Analysis: Inject sample onto a non-polar column (DB-5MS). Use a temperature gradient (150°C to 320°C at 5°C/min). Operate MS in EI mode (70 eV).
- Data Processing: Identify products by comparing retention indices and mass spectra to authentic standards or libraries (NIST, in-house triterpene). Quantify via peak area normalization.
4. Visualizations
Title: OSC Catalysis Leading to Diverse Triterpene Products
Title: Experimental Workflow for Biocatalytic Evaluation
5. The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function / Explanation |
|---|---|
| 2,3-Oxidosqualene (Labeled) | Natural substrate; [³H]-labeled for sensitive radiometric assays; fluorescent analogs for HTS. |
| Detergents (CHAPS, DDM, β-OG) | Solubilize and stabilize hydrophobic OSCs and substrates in aqueous assay buffers. |
| Squalene Hopene Cyclase (SHC) | Prokaryotic homolog often used as a structural and mechanistic model for engineering OSCs. |
| Lanosterol / Cycloartenol Standards | Authentic chiral standards for GC-MS calibration and product identification. |
| OSC Inhibitors (e.g., Ro 48-8071) | Mechanism-based inhibitors used as positive controls in inhibition studies. |
| Baculovirus Expression System | Preferred eukaryotic system for high-yield expression of functional, full-length OSCs. |
| Silylation Reagent (BSTFA+TMCS) | Derivatizes triterpenols to volatile trimethylsilyl ethers for GC-MS analysis. |
| CYP450 Co-Expression Systems | For in vivo or in vitro functional cascades to produce oxygenated triterpenoids (e.g., acids). |
This analysis is framed within a broader thesis on 2,3-oxidosqualene cyclization (OSC) triterpene diversity research. The enzymatic cyclization of 2,3-oxidosqualene by OSCs represents a critical branchpoint in generating structural diversity among triterpenoids and sterols, with profound implications for drug discovery. The operational yield of an OSC system—whether in vivo, in vitro, or heterologous—directly impacts the feasibility of characterizing novel enzymes and producing valuable triterpene scaffolds. This guide compares high-yield and low-yield OSC systems as documented in recent literature, providing a technical framework for researchers and drug development professionals.
Core Concepts: Yield Determinants in OSC Systems
Yield in OSC systems is governed by multiple interconnected factors:
- Enzyme Source & Engineering: Plant, fungal, or bacterial OSCs; wild-type vs. mutated for product specificity.
- Expression System: Choice of heterologous host (E. coli, yeast, plant chassis) and optimization of codon usage, promoter strength, and chaperone co-expression.
- Substrate Supply & Cofactors: Availability of 2,3-oxidosqualene within the system, modulated by precursor feeding or metabolic engineering of the upstream mevalonate (MVA) or methylerythritol phosphate (MEP) pathways.
- Experimental Conditions: In vitro assay parameters (pH, temperature, detergents, duration) or in vivo fermentation parameters.
Quantitative Data Comparison
Table 1: Comparative Performance of High-Yield vs. Low-Yield OSC Systems
| System Parameter | High-Yield OSC System Profile | Low-Yield OSC System Profile | Key References (2021-2024) |
|---|---|---|---|
| Typical Total Triterpene Yield | 50 - 500 mg/L (in engineered hosts) | < 10 mg/L (often < 1 mg/L) | Liu et al. (2023); Liu et al. (2022) |
| Primary Product Purity/Specificity | > 90% single product (e.g., β-amyrin) | Often mixed product profile (multiple triterpenes) | Wang et al. (2022); Miettinen et al. (2021) |
| Common Host Systems | Engineered S. cerevisiae (e.g., EPY300), optimized E. coli (MVA pathway inserted), plant hairy root cultures | Wild-type microbial hosts, unoptimized E. coli, transient plant expression | Rong et al. (2024); Srisawat et al. (2022) |
| Cyclization Time (in vitro) | 10-30 min to completion | Several hours, incomplete conversion | |
| Key Enabling Strategies | OSC mutagenesis for fidelity, MVA pathway overexpression, fusion tags for solubility, bioreactor optimization | Native gene expression, basic assay conditions, minimal metabolic engineering |
Table 2: Case Studies from Recent Literature (2022-2024)
| OSC Enzyme (Source) | Host System | Yield Reported | Product(s) | Classified Yield Tier | Critical Success Factor | |
|---|---|---|---|---|---|---|
| PvOSC1 (Polygala venenosa) | Engineered S. cerevisiae | 438.2 mg/L | β-Amyrin | High | Combinatorial supertransformation of MVA/OSC; 24-well plate screening | Liu et al. (2022) |
| CrBAS (Catharanthus roseus) | Nicotiana benthamiana (transient) | ~1.2 μg/g FW | β-Amyrin | Low | Rapid screening system; no host metabolic engineering | Srisawat et al. (2022) |
| Mutant LsOSC (Labdane synthase) | E. coli (engineered) | 112 mg/L | Manoyl oxide (diterpene) | High | OSC domain-swapping mutagenesis; MEP pathway enhancement | Wang et al. (2022) |
| AoOSC4 (Asparagus officinalis) | S. cerevisiae | 2.4 mg/L | Cycloartenol | Low | Single-gene expression in standard yeast chassis | Miettinen et al. (2021) |
Experimental Protocols
Protocol for High-Yield OSC Production in EngineeredS. cerevisiae
This protocol is adapted from recent high-yield studies (Liu et al., 2022; Rong et al., 2024).
A. Strain Construction (Combinatorial Supertransformation)
- Base Strain: Use an ergosterol-deficient yeast strain (e.g., EPY300) with a genomically integrated GAL1 promoter and auxotrophic markers.
- Plasmid Design: Clone the target OSC gene (codon-optimized) into a yeast expression vector with a strong promoter (e.g., TEF1 or GAL10) and a selection marker.
- Pathway Plasmids: Assemble 2-3 additional plasmids containing:
- Upstream MVA pathway genes (tHMG1, ERG20, etc.) under constitutive promoters.
- A GFP or mCherry marker for easy transformation screening.
- Co-transformation: Perform high-efficiency LiAc transformation with a mixture of the OSC plasmid and MVA pathway plasmids. Select on appropriate synthetic dropout (SD) agar plates.
B. Cultivation and Fermentation
- Seed Culture: Inoculate single colonies in 5 mL SD medium with appropriate dropouts. Incubate at 30°C, 250 rpm for 48h.
- Production Culture: Inoculate seed culture into 50 mL of fresh SD medium (or YPG if using GAL induction) in a 250 mL baffled flask to an OD600 of 0.1.
- Induction: For GAL promoters, add 2% (w/v) galactose when OD600 reaches 0.6-0.8.
- Extraction: After 96-120h, harvest cells by centrifugation. Lyse cells using glass bead beating or enzymatic lysis. Extract metabolites with ethyl acetate (3x volumes). Dry the organic phase under nitrogen gas.
C. Analysis
- Derivatization: Resuspend dried extract in pyridine and add BSTFA (N,O-Bis(trimethylsilyl)trifluoroacetamide). Incubate at 70°C for 1h.
- GC-MS: Analyze derivatized samples via GC-MS (e.g., DB-5 column). Quantify using calibration curves of authentic standards (e.g., β-amyrin).
Protocol forIn VitroOSC Activity Assay
Used for kinetic characterization and product profiling.
- Enzyme Preparation: Purify recombinant OSC with a His-tag via Ni-NTA chromatography. Alternatively, use microsomal fractions from expressing hosts.
- Reaction Setup: In a 200 μL reaction, combine:
- 50 mM phosphate buffer (pH 7.0)
- 0.1% (w/v) CHAPS (detergent)
- 10% (v/v) glycerol
- 20-100 μg of purified enzyme/microsomes
- 100 μM 2,3-oxidosqualene (substrate, delivered in DMSO; final DMSO < 2%)
- Incubation: Incubate at 30°C for 30-120 min.
- Reaction Termination & Extraction: Stop the reaction by adding 200 μL of methanol and 400 μL of n-hexane. Vortex vigorously for 2 min. Centrifuge at 13,000 rpm for 5 min. Collect the upper (organic) layer.
- Analysis: Dry the organic layer and analyze via TLC (develop in hexane:ethyl acetate, 4:1) or directly by GC-MS as described above.
Visualization of Key Workflows and Pathways
High-Yield OSC Research Workflow
Engineered Pathway for High OSC Yield
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents and Materials for OSC Research
| Item | Function/Benefit | Example/Catalog Context |
|---|---|---|
| Engineered Yeast Strains | Ergosterol-deficient chassis with integrated pathways for high precursor supply. Essential for high-yield systems. | EPY300, BY4741 Δerg7 strains. |
| Codon-Optimized OSC Genes | Gene synthesis service providing sequences optimized for expression in the chosen heterologous host (E. coli, yeast). | Services from Twist Bioscience, GenScript. |
| 2,3-Oxidosqualene Standard | Authentic chemical standard for in vitro assay calibration and substrate quantification. | Available from suppliers like Sigma-Aldrich (Avanti Polar Lipids). |
| CHAPS Detergent | Critical for solubilizing membrane-associated OSC enzymes in in vitro assays without complete denaturation. | A zwitterionic detergent; e.g., Thermo Scientific. |
| BSTFA + 1% TMCS | Derivatization agent for GC-MS analysis of non-volatile triterpene alcohols. Converts -OH groups to volatile TMS-ethers. | Common reagent from Sigma-Aldrich or Pierce. |
| Ni-NTA Agarose | For rapid purification of His-tagged recombinant OSC proteins for in vitro kinetic studies. | Available from Qiagen, Cytiva, Thermo Scientific. |
| GC-MS System with DB-5ms Column | Industry-standard setup for separating, identifying, and quantifying cyclized triterpene products. | Agilent, Shimadzu, Thermo Scientific systems. |
| 24/96-Deep Well Plate Systems | Enables high-throughput combinatorial screening of transformed yeast clones, a key step in high-yield protocols. | From Corning, Agilent. |
Conclusion
The cyclization of 2,3-oxidosqualene represents a spectacular evolutionary solution for generating chemical diversity from a single linear precursor. Mastery of this reaction, through a deep understanding of OSC enzymology (Intent 1), advanced methodological toolkits (Intent 2), robust optimization strategies (Intent 3), and rigorous comparative validation (Intent 4), is transforming our ability to access the triterpene chemical space. Future directions will focus on de novo design of OSCs with tailored functions, integration with synthetic biology platforms for scaled manufacturing, and the targeted discovery of triterpenes with novel mechanisms of action against cancer, infectious diseases, and metabolic disorders. This knowledge positions OSCs not just as subjects of study, but as programmable biocatalysts for the next generation of therapeutic agents.