Decoding Substrate Specificity: CCD1 vs. CCD4a Enzymes in Carotenoid Degradation Pathways
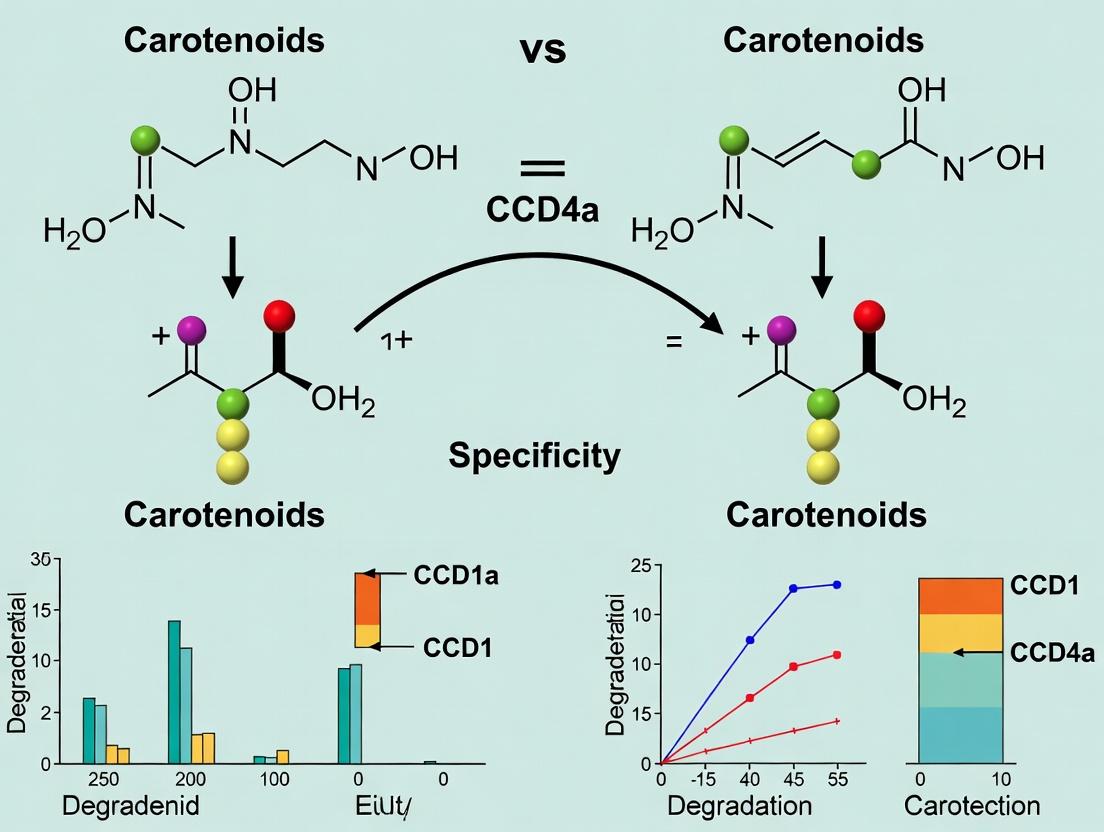
This article provides a comprehensive analysis of the structural and functional distinctions between carotenoid cleavage dioxygenases CCD1 and CCD4a, with a focus on their substrate specificity and degradation products.
Decoding Substrate Specificity: CCD1 vs. CCD4a Enzymes in Carotenoid Degradation Pathways
Abstract
This article provides a comprehensive analysis of the structural and functional distinctions between carotenoid cleavage dioxygenases CCD1 and CCD4a, with a focus on their substrate specificity and degradation products. We explore the foundational biology of these enzymes, detail current methodologies for studying their activity, address common experimental challenges, and present a comparative validation of their roles. Targeted at researchers and drug development professionals, this review synthesizes recent findings to highlight implications for biomedical research, including the development of therapies targeting retinoid-related pathways and nutritional interventions.
Unraveling the Core Biology: Structural and Functional Basics of CCD1 and CCD4a
Carotenoid Cleavage Dioxygenases (CCDs) are a family of non-heme iron enzymes that catalyze the oxidative cleavage of carotenoids, producing apocarotenoids with diverse biological functions. Research into their specificities, particularly comparative studies of CCD1 versus CCD4 subfamily members, is central to understanding apocarotenoid regulation in physiology and potential drug development. This guide compares key phylogenetic and functional characteristics of plant CCDs, with a focus on experimental data elucidating CCD1 and CCD4a substrate specificity.
Phylogenetic Classification & Nomenclature
CCDs are phylogenetically divided into two major clades: CCD1-like and CCD4/CCD7/CCD8-like enzymes, with CCD4 further subdivided (e.g., CCD4a, CCD4b). Nomenclature is primarily based on sequence homology and genetic analysis in model plants like Arabidopsis thaliana.
Table 1: Phylogenetic and Functional Comparison of Key Plant CCD Subfamilies
| Subfamily | Key Phylogenetic Marker | Primary Localization | Characterized Substrate Preference | Primary Apocarotenoid Products |
|---|---|---|---|---|
| CCD1 | Deep-branching clade; cytosolic | Cytoplasm, Peripheral to plastids | Symmetrical cleavage at 9,10(9',10') positions of multiple carotenoids | β-ionone, geranylacetone |
| CCD4 | Clade with CCD7/CCD8; plastid-targeting signal | Plastid (Chloroplast/Chromoplast) | Preferential cleavage at 9,10(9',10') of β-carotene, lutein; tissue-specific expression | β-ionone (in flowers), lutein-derived products |
| CCD7 (MAX3/HTD1) | Sister to CCD4 | Plastid | Asymmetric cleavage at 9,10 position of β-carotene | β-apo-10'-carotenal |
| CCD8 (MAX4/DAD1) | Sister to CCD4 | Plastid | Cleavage of β-apo-10'-carotenal (CCD7 product) | Carlactone (strigolactone precursor) |
Comparative Analysis: CCD1 vs. CCD4a Degradation Specificity
A core thesis in the field investigates the divergent substrate specificities of cytosolic CCD1 and plastidial CCD4a, despite catalyzing similar double-bond cleavages. Key experimental comparisons are summarized below.
Table 2: Experimental Comparison of CCD1 and CCD4a Substrate Specificity In Vitro
| Experimental Parameter | CCD1 (e.g., Arabidopsis AtCCD1) | CCD4a (e.g., Chrysanthemum CmCCD4a) | Experimental Implications |
|---|---|---|---|
| Recombinant Enzyme Source | E. coli expressing apoenzyme + Fe²⁺ reconstitution | E. coli expression with plastid-targeting peptide truncation | Both require in vitro iron incorporation for activity. |
| Substrate Range (Tested) | Broad: β-carotene, lycopene, zeaxanthin, lutein | Narrower: High activity on β-carotene, lower on lutein | CCD1 is a promiscuous cleaver; CCD4a shows higher selectivity. |
| Kinetic Data (β-carotene) | Km: ~10-20 µM; Vmax: ~50-100 pmol/min/µg | Km: ~2-5 µM; Vmax: ~150-200 pmol/min/µg | CCD4a may have higher affinity and turnover for β-carotene. |
| Primary In Vitro Product | β-ionone (from 9,10 cleavage) | β-ionone (from 9,10 cleavage) | Products identical; specificity differs in substrate access/selection. |
| Critical Inhibitor | 2,2'-Bipyridyl (iron chelator) - abolishes activity | 2,2'-Bipyridyl (iron chelator) - abolishes activity | Confirms both are iron-dependent dioxygenases. |
Experimental Protocols for Key Studies
Protocol 1: In Vitro Enzyme Assay for CCD Activity
- Enzyme Preparation: Heterologously express His-tagged CCD1 or truncated CCD4a in E. coli. Purify via Ni-NTA affinity chromatography.
- Apoenzyme Reconstitution: Incubate purified protein with 1-2 mM FeSO₄ and 2 mM ascorbate (reductant) for 1 hour at 4°C. Remove excess iron via desalting column.
- Substrate Delivery: Solubilize carotenoid substrate (e.g., β-carotene) in acetone. Deliver to assay buffer containing 0.1-0.5% (w/v) Tween 40. Final substrate concentration typically 5-50 µM.
- Reaction: Mix reconstituted enzyme with substrate emulsion. Incubate at 30°C with agitation for 30-120 minutes.
- Product Extraction & Analysis: Stop reaction with methanol. Extract apocarotenoids with hexane:ethyl acetate (9:1). Analyze via HPLC-PDA or GC-MS. Identify β-ionone by retention time and spectral match to authentic standard.
Protocol 2: In Planta Functional Validation via Overexpression
- Vector Construction: Clone full-length CCD1 or CCD4a cDNA under a constitutive promoter (e.g., CaMV 35S) in plant binary vector.
- Plant Transformation: Transform Arabidopsis (wild-type or carotenoid-accumulating mutant) or tobacco using Agrobacterium tumefaciens-mediated floral dip or leaf disc method.
- Phenotypic Screening: Select transgenic lines on antibiotic. Observe visible phenotypes (e.g., petal color whitening for CCD4a, altered volatile emission for CCD1).
- Metabolite Profiling: Extract carotenoids and apocarotenoids from transgenic tissues. Quantify via LC-MS/MS to confirm substrate depletion and product formation.
Visualizations
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CCD Functional Analysis
| Reagent/Material | Function in Research | Example Application |
|---|---|---|
| Heterologous Expression System | High-yield production of recombinant CCD protein. | pET vectors in E. coli BL21(DE3) for enzyme purification. |
| FeSO₄ & Ascorbic Acid | Cofactor supply for reconstituting active iron center in apo-CCD enzymes. | In vitro enzyme reconstitution post-purification. |
| Carotenoid Substrates | Native enzyme substrates for activity assays. | β-carotene, lutein, lycopene for in vitro cleavage assays. |
| Tween 40/Emulsifiers | Facilitate delivery of hydrophobic carotenoids to enzyme in aqueous assay buffer. | Creating substrate emulsions for in vitro reactions. |
| β-ionone Standard | Authentic chemical standard for product identification and quantification. | HPLC/GC-MS calibration to confirm enzymatic product. |
| 2,2'-Bipyridyl | Specific iron chelator; negative control to confirm dioxygenase mechanism. | Inhibition of CCD activity in control experiments. |
| Plant Binary Vectors | For stable or transient expression of CCD genes in plant models. | pBI121 or pEAQ vectors for Arabidopsis/tobacco transformation. |
| LC-MS/MS System | Sensitive identification and quantification of carotenoids/apocarotenoids. | Profiling metabolites in transgenic plants or enzyme assays. |
This comparison guide examines the gene structure and protein localization mechanisms of carotenoid cleavage dioxygenases (CCDs), with a specific focus on CCD1 and CCD4a. Understanding the chromosomal context and targeting signals of these enzymes is fundamental to elucidating their distinct substrate specificities and physiological roles in carotenoid degradation, a key area in plant metabolic engineering and nutraceutical development.
Comparative Analysis: Chromosomal Context and Gene Structure
The genomic organization of CCD1 and CCD4a genes influences their regulation and evolutionary divergence.
Table 1: Comparison of Gene Structure and Genomic Context
| Feature | CCD1 (e.g., Arabidopsis thaliana AtCCD1) | CCD4a (e.g., Arabidopsis thaliana AtCCD4) | Functional Implication |
|---|---|---|---|
| Chromosomal Location | Chromosome 4 (AT4G19170) | Chromosome 4 (AT4G19150) | Physical clustering suggests gene duplication events. |
| Exon/Intron Structure | 6 exons, 5 introns | 7 exons, 6 introns | Alternative splicing potential differs; CCD4a structure is more complex. |
| Upstream Regulatory Elements | Contains light-responsive (G-box), stress-responsive elements | Enriched in ethylene-responsive (ERE) and circadian-related elements | Differential transcriptional regulation: CCD1 linked to general stress/development; CCD4a tied to senescence & specific tissue responses. |
| Paralogous Gene Family | Member of a small subclade with CCD7, CCD8 | Member of a distinct subclade with CCD4b, CCD4c | Subfunctionalization after duplication led to divergent substrate preferences. |
Comparative Analysis: Subcellular Targeting Signals
Subcellular localization dictates access to carotenoid substrates, critically defining CCD1 and CCD4a function.
Table 2: Comparison of Targeting Signals and Localization
| Feature | CCD1 | CCD4a | Supporting Experimental Evidence |
|---|---|---|---|
| Predicted Signal | N-terminal lacking canonical organelle transit peptides. | N-terminal chloroplast transit peptide (cTP). | In silico prediction tools (TargetP, ChloroP). |
| Experimentally Confirmed Localization | Cytoplasm (soluble). | Chloroplast stroma (plasticidic). | Transient expression of GFP-fusions in Nicotiana benthamiana leaves. |
| Key Targeting Sequence | None identifiable. | First ~50-70 amino acids are necessary and sufficient. | Truncation/mutation analysis via GFP fusions shows loss of plastid import. |
| Impact on Substrate Access | Accesses cytosolic/apoplastic carotenoids or derived products (e.g., xanthophylls). | Accesses chloroplast-localized carotenoid pools (e.g., β-carotene, lutein). | In vitro assays with isolated chloroplasts show CCD4a, but not CCD1, degrades plastid pigments. |
Detailed Experimental Protocol: Subcellular Localization via Transient GFP Expression
Aim: To experimentally determine the localization of CCD1 and CCD4a proteins. Materials: cDNA clones, GFP vector (e.g., pSAT6-GFP), Agrobacterium tumefaciens strain GV3101, Nicotiana benthamiana plants, confocal laser scanning microscope. Method:
- Fusion Construct Creation: Amplify full-length and truncated coding sequences of CCD1 and CCD4a without stop codons. Clone in-frame into the GFP vector.
- Agrobacterium Transformation: Introduce constructs into Agrobacterium.
- Plant Infiltration: Grow N. benthamiana plants for 4-5 weeks. Infiltrate leaves with Agrobacterium cultures (OD600 ~0.5) using a needleless syringe.
- Incubation: Maintain plants for 48-72 hours post-infiltration under light.
- Microscopy: Visualize GFP fluorescence (excitation 488 nm) alongside chloroplast autofluorescence (excitation 633 nm) using a confocal microscope. Key Control: Co-infiltrate with a known plastid marker (RFP-peroxisome marker) for co-localization analysis.
Visualization of Localization Pathways
Title: CCD1 and CCD4a Protein Synthesis and Localization Pathways
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CCD Localization and Function Studies
| Reagent/Material | Function/Benefit | Example/Supplier |
|---|---|---|
| Gateway-Compatible GFP Vectors | Enables rapid, in-frame cloning of CCD genes for transient expression. Reduces cloning artifacts. | pSAT6-GFP, pEarleyGate series (Addgene). |
| Agrobacterium tumefaciens GV3101 | Standard strain for high-efficiency transient transformation of Nicotiana benthamiana leaves. | Common lab strain, available from culture collections. |
| Confocal Microscope with Spectral Detection | Allows precise co-localization by separating GFP signal from chloroplast autofluorescence. | Zeiss LSM 980, Leica SP8. |
| Chloroplast Isolation Kit | Provides purified, intact chloroplasts for in organello enzyme activity assays. | Merck Chloroplast Isolation Kit, Abcam kits. |
| Carotenoid Standards | Essential for calibrating HPLC/MS systems to identify and quantify cleavage products. | β-carotene, lutein, β-ionone (CaroteNature, Sigma-Aldrich). |
| Protease Inhibitor Cocktails | Preserves protein integrity during chloroplast import assays and protein extraction. | cOmplete, EDTA-free (Roche). |
| cTP Prediction Software | In silico identification of targeting signals to guide experimental design. | TargetP-2.0, ChloroP, Predotar. |
This comparison guide, situated within a broader thesis on CCD1 vs. CCD4a carotenoid degradation specificity, objectively analyzes the structural determinants of substrate selectivity in these carotenoid cleavage dioxygenases (CCDs). The focus is on active site architecture, supported by experimental structural biology data.
Comparative Analysis of Active Site Structural Parameters
The defining differences in substrate specificity between cytosolic CCD1 and plastidial CCD4a are rooted in their distinct active site geometries, as revealed by X-ray crystallography and mutagenesis studies. CCD1 favors symmetric cleavage of linear carotenes like lycopene, while CCD4a preferentially catalyzes asymmetric cleavage of bicyclic substrates like β-carotene and specific apocarotenoids.
Table 1: Key Structural Features of CCD1 vs. CCD4a Active Sites
| Feature | CCD1 | CCD4a | Functional Implication |
|---|---|---|---|
| Overall Cavity Shape | Long, narrow, and linear. | Wider, more enclosed, and bent. | Accommodates linear vs. bicyclic substrates. |
| Volume (ų) | ~700-800 | ~500-600 | Limits size and orientation of substrate. |
| Entry Portal | Relatively open. | Partially obstructed by flexible loops. | Influences substrate access and regiospecificity. |
| Key Gating Residues | Smaller residues (e.g., Leu, Val). | Bulky aromatic residues (e.g., Phe, Trp). | Creates steric hindrance, defining cleavage position. |
| Substrate-Binding Flexibility | High; allows sliding. | Restricted; locks substrate in specific pose. | Determines symmetric vs. asymmetric cleavage outcome. |
| Catalytic Fe²⁺ Coordination | Conserved 4-His motif. | Conserved 4-His motif. | Essential for dioxygen activation; conserved mechanism. |
Experimental Protocols for Key Cited Studies
Protocol 1: X-ray Crystallography for Active Site Determination
- Protein Expression & Purification: Recombinant CCD1 (e.g., from A. thaliana) and CCD4a (e.g., from C. morifolium) are expressed in E. coli with a cleavable His-tag. Proteins are purified via immobilized metal affinity chromatography (IMAC) followed by size-exclusion chromatography (SEC).
- Crystallization: Purified proteins are concentrated to 10-20 mg/mL. Crystals are obtained via vapor diffusion using conditions containing PEGs as precipitants. For substrate-complex structures, crystals are soaked with 1-5 mM substrate (e.g., β-apo-8'-carotenal) or an unreactive analog.
- Data Collection & Structure Solution: Diffraction data are collected at a synchrotron source (e.g., 100 K). Structures are solved by molecular replacement using a known CCD (e.g., AtCCD1) as a search model. Iterative rounds of refinement and model building are performed.
- Analysis: Active site cavities are defined using software like CASTp or CAVER. Residue interactions and distances are measured in PyMOL or Chimera.
Protocol 2: Site-Directed Mutagenesis Coupled with Activity Assays
- Mutagenesis: Target residues in the CCD4a active site (e.g., Phe, Trp) are mutated to alanine or to their CCD1 counterparts using PCR-based mutagenesis kits.
- Protein Production: Wild-type and mutant proteins are expressed and purified as in Protocol 1.
- In vitro Enzymatic Assay: Reactions contain 5 µM enzyme, 20 µM substrate (e.g., β-carotene solubilized in 0.1% Tween-40), 50 mM Tris-HCl (pH 7.5), and 150 mM NaCl. Reactions proceed at 30°C for 1 hour.
- Product Analysis: Products are extracted with ethyl acetate and analyzed by reverse-phase HPLC with diode-array detection, comparing retention times and spectra to authentic standards. Kinetic parameters (Km, kcat) are determined.
Visualization of Structural Determinants of Specificity
Diagram 1: Substrate Binding and Cleavage Outcomes
Diagram 2: CCD4a Active Site Architecture
Research Reagent Solutions Toolkit
Table 2: Essential Materials for CCD Structural & Functional Analysis
| Item | Function in Research | Example/Catalog Consideration |
|---|---|---|
| Recombinant Expression Vector | High-yield protein production in E. coli. | pET-28a(+) with TEV protease site for His-tag removal. |
| Affinity Chromatography Resin | One-step purification of His-tagged proteins. | Ni-NTA or Co²⁺-based TALON resin. |
| Size-Exclusion Column | Final polishing step for monodisperse, pure protein. | Superdex 200 Increase for SEC. |
| Crystallization Screen Kits | Initial search for protein crystallization conditions. | JCSG+, Morpheus, or PEG/Ion screens. |
| Carotenoid/Apocarotenoid Standards | HPLC reference for product identification. | β-Carotene, β-ionone, crocetin dialdehyde. |
| Site-Directed Mutagenesis Kit | Engineering point mutations in active site residues. | Q5 Hot Start or KAPA HiFi kits for high fidelity. |
| Iron Chelator/Assay | Verifying Fe²⁺ incorporation in active site. | Bathophenanthroline disulfonate (BPDS) assay. |
| HPLC System with DAD | Separation and spectral analysis of hydrophobic products. | C30 reverse-phase column for carotenoid separation. |
Within the expanding thesis on carotenoid cleavage dioxygenase (CCD) specificity—particularly comparing the divergent substrate preferences of CCD1 and CCD4a enzymes—understanding their in vivo roles is critical. This guide compares the endogenous functions of key carotenoid-cleaving enzymes across model organisms, focusing on the physiological consequences of their substrate specificity.
Comparison of Endogenous Substrates and Physiological Outputs
Table 1: In Vivo Substrate Specificity and Functional Roles of Selected CCDs
| Enzyme (Organism) | Primary Endogenous Substrate(s) | Biological Function & Phenotype of Loss-of-Function Mutant | Key Experimental Evidence |
|---|---|---|---|
| CCD1 (Arabidopsis thaliana) | β-carotene, lutein (in vitro); likely volatile apocarotenoid precursors in planta. | Minimal impact on plant pigmentation. Major role in producing volatile apocarotenoid signals (e.g., β-ionone) for pollinator/defense communication. | Mutant ccd1 plants show >80% reduction in β-ionone emissions (measured by GC-MS). No change in leaf or petal carotenoid levels. |
| CCD4a (Arabidopsis thaliana) | Specific carotenoids in chloroplasts: Lutein, β-carotene. | Regulates photosynthetic apparatus and photoprotection. Mutants exhibit photo-oxidative stress under high light and altered non-photochemical quenching (NPQ). | HPLC analysis shows 30-40% higher lutein in ccd4a mutant chloroplasts. Increased sensitivity to photooxidative bleaching. |
| CCD4 (Chrysanthemum morifolium) | β-carotene in chromoplasts. | Direct determinant of petal color. Cleaves β-carotene, preventing its accumulation. Mutants have white → yellow petals due to carotenoid accumulation. | Genetic mapping; white-flowered cultivars contain a non-functional CCD4 allele. In vitro assay confirms β-carotene cleavage. |
| NCED3 (Arabidopsis thaliana) | 9-cis-violaxanthin & 9'-cis-neoxanthin (xanthophylls). | Central regulator of abscisic acid (ABA) biosynthesis for drought stress response. Mutants are wilty and ABA-deficient. | LC-MS shows >90% reduction in ABA in water-stressed nced3. Precursor (cis-xanthophylls) accumulation confirmed. |
| CCD7/CCD8 (Arabidopsis; Strigolactone Biosynthesis) | Unknown carotenoid precursor(s). | Produces strigolactone hormones, inhibiting shoot branching. Mutants exhibit highly branched phenotype. | Grafting experiments; rootstock of ccd7/ccd8 mutants fails to rescue wild-type branching in scion. |
Detailed Experimental Protocols
1. Protocol: Gas Chromatography-Mass Spectrometry (GC-MS) for Apocarotenoid Volatiles (CCD1 Function)
- Objective: Quantify volatile apocarotenoids (e.g., β-ionone) emitted from plant tissues.
- Materials: Headspace sampling chamber, Tenax TA adsorption tubes, GC-MS system.
- Steps:
- Enclose intact flowers or leaves of wild-type and
ccd1mutant in a sealed glass vessel. - Purge volatile organic compounds onto a Tenax TA adsorbent trap for 60 minutes.
- Desorb trapped compounds thermally into the GC inlet.
- Separate compounds on a non-polar capillary column (e.g., DB-5) with a temperature gradient (40°C to 250°C).
- Detect and identify compounds via electron impact mass spectrometry. Quantify β-ionone by comparing peak area to an authentic standard curve.
- Enclose intact flowers or leaves of wild-type and
2. Protocol: Chloroplast Isolation and Carotenoid Profiling via HPLC (CCD4a Function)
- Objective: Analyze carotenoid composition within isolated chloroplasts.
- Materials: Sucrose density gradient media, Potter-Elvehjem homogenizer, refrigerated centrifuge, C30 reverse-phase HPLC column.
- Steps:
- Homogenize fresh leaf tissue in ice-cold grinding buffer.
- Filter homogenate and centrifuge at low speed to remove debris.
- Pellet chloroplasts via centrifugation at 2,500 x g for 10 min.
- Purify chloroplasts on a discontinuous sucrose gradient.
- Extract pigments from the chloroplast pellet with acetone.
- Separate carotenoids by HPLC using a C30 column with a methanol/MTBE/water gradient.
- Identify and quantify peaks using diode-array detection, comparing retention times and spectra to known standards.
Visualization of Pathways and Workflows
Diagram Title: Physiological Outputs from CCD Substrate Cleavage
Diagram Title: Workflow for Analyzing CCD Substrates and Products
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents and Kits for CCD Functional Analysis
| Item | Function in Research | Example Application in Protocols Above |
|---|---|---|
| C30 Reverse-Phase HPLC Columns | Superior separation of geometric and structural carotenoid isomers. | Quantifying lutein vs. β-carotene in chloroplast extracts. |
| Deuterated Internal Standards (e.g., d5-ABA, d3-β-ionone) | Allows precise, matrix-effect-corrected quantification via mass spectrometry. | Accurate measurement of ABA or apocarotenoid volatiles in complex samples. |
| Chloroplast Isolation Kits | Provide optimized buffers and density media for intact organelle preparation. | Isolating pure chloroplasts for CCD4a substrate analysis. |
| Carotenoid & Apocarotenoid Reference Standards | Essential for identifying HPLC/GC peaks and constructing calibration curves. | Identifying lutein, β-carotene, β-apo-8'-carotenal, β-ionone. |
| Tenax TA Adsorbent Tubes/Traps | Reliable capture of volatile organic compounds for thermal desorption. | Headspace sampling of floral volatiles in CCD1 studies. |
| LC-MS Grade Solvents (Acetone, Methanol, MTBE) | Minimize background noise and ion suppression in sensitive MS detection. | Extracting and separating carotenoids for NCED3/CCD product profiling. |
Evolutionary Conservation and Divergence of CCD1 and CCD4a Across Species
Within the broader thesis investigating the carotenoid degradation specificity of CCD1 versus CCD4a, this guide provides a comparative analysis of their evolutionary conservation and divergence. This comparison is critical for understanding functional specialization, informing protein engineering, and identifying species-specific metabolic pathways relevant to nutraceutical and pharmaceutical development.
Structural and Functional Comparison
The core enzymatic activity of CCD1 and CCD4a involves the oxidative cleavage of carotenoids at specific double bonds, but their substrate preferences and biological roles have diverged significantly.
Table 1: Key Functional Characteristics of CCD1 and CCD4a
| Feature | CCD1 | CCD4a |
|---|---|---|
| Primary Cleavage Site | 9,10 (9',10') | 9,10 (9',10') |
| Typical Substrates | Lycopene, β-carotene, Zeaxanthin | β-carotene, Lutein, Neoxanthin |
| Main Product(s) | C13 apo-carotenoids (e.g., pseudoionone) | C10 apo-carotenoids (e.g., β-ionone) & C27 |
| Subcellular Localization | Cytosol | Plastid |
| Primary Physiological Role | Volatile scent/flavor production; general cleavage | Color pigmentation (e.g., saffron crocin, white flowers) |
| Phylogenetic Distribution | Widely conserved in plants; found in some bacteria and fungi | Primarily in plants; CCD4 clade expanded in specific lineages (e.g., Rosaceae) |
Table 2: Quantitative Kinetic Parameters from Recombinant Enzymes (Representative Data)
| Enzyme (Species) | Substrate | Km (µM) | kcat (s⁻¹) | kcat/Km (M⁻¹s⁻¹) | Reference Context |
|---|---|---|---|---|---|
| CCD1 (A. thaliana) | Lycopene | 8.2 | 0.15 | 1.83 x 10⁴ | In vitro assay with E. coli membranes |
| CCD1 (C. roseus) | β-carotene | 5.5 | 0.08 | 1.45 x 10⁴ | Recombinant protein in yeast |
| CCD4a (C. sativus) | β-carotene | 2.1 | 0.32 | 1.52 x 10⁵ | Affinity-purified enzyme |
| CCD4a (M. domestica) | Zeaxanthin | 12.7 | 0.04 | 3.15 x 10³ | Transient expression in N. benthamiana |
Evolutionary Analysis
Phylogenetic reconstruction reveals that CCD1 and CCD4a belong to distinct subfamilies within the larger CCD enzyme family. CCD1 is more ancient, with homologs in cyanobacteria, suggesting an origin in primary carotenoid metabolism. CCD4a evolved later in plants, likely from a gene duplication event, and has undergone more recent lineage-specific expansions, correlating with novel pigmentation traits.
Table 3: Conservation of Key Motifs and Residues
| Motif/Residue | CCD1 Conservation | CCD4a Conservation | Proposed Function |
|---|---|---|---|
| Fe²⁺-Binding His Residues | 100% (HxxxH) | 100% (HxxxH) | Dioxygenase active site coordination |
| Plastid Transit Peptide | Absent | Highly conserved N-terminal signal | Subcellular targeting |
| Substrate Channel Residues | Variable | More conserved, hydrophobic | Substrate specificity determination |
Experimental Protocols for Key Studies
Protocol 1: Heterologous Expression and In Vitro Enzyme Assay
- Objective: Determine kinetic parameters (Km, kcat) for CCD1/CCD4a.
- Methodology:
- Clone full-length cDNA (without transit peptide for CCD4a) into a prokaryotic (e.g., pET) or yeast expression vector.
- Express protein in E. coli BL21(DE3) or Saccharomyces cerevisiae. Induce with IPTG or galactose.
- Isolate membranes/protein via centrifugation and purification (Ni-NTA if His-tagged).
- Prepare substrate by embedding carotenoid (e.g., β-carotene) in membrane vesicles (from E. coli or liposomes).
- Incubate purified enzyme with substrate in reaction buffer (Tris-HCl, pH 7.5, with DTT, FeSO₄).
- Extract products with hexane/ethyl acetate and analyze via HPLC-PDA or GC-MS.
- Calculate kinetic parameters by varying substrate concentration.
Protocol 2: Subcellular Localization Visualization
- Objective: Confirm cytosol vs. plastid localization.
- Methodology:
- Fuse full-length CCD4a (with transit peptide) or CCD1 to GFP at the N- or C-terminus.
- Construct transiently expressed in Arabidopsis mesophyll protoplasts or Nicotiana benthamiana leaves via PEG-mediated transformation or agroinfiltration.
- After 24-48 hours, visualize GFP fluorescence using confocal microscopy alongside chloroplast autofluorescence (chlorophyll channel).
- Co-localization analysis confirms plastid targeting for CCD4a, diffuse cytosolic signal for CCD1.
Protocol 3: Phylogenetic Tree Construction
- Objective: Analyze evolutionary relationships.
- Methodology:
- Retrieve CCD protein sequences from public databases (NCBI, Phytozome) using BLAST.
- Perform multiple sequence alignment with Clustal Omega or MUSCLE.
- Trim alignment to conserved regions using Gblocks.
- Construct phylogenetic tree using Maximum Likelihood method (e.g., RAxML) or Bayesian inference (MrBayes).
- Visualize tree with FigTree or iTOL, annotating CCD1 and CCD4/CCD4a clades.
Visualizations
Diagram 1: Proposed evolutionary pathway of CCD1 and CCD4.
Diagram 2: Experimental workflow for subcellular localization.
The Scientist's Toolkit
Table 4: Essential Research Reagent Solutions
| Item | Function in CCD1/CCD4a Research | Example/Note |
|---|---|---|
| Carotenoid Substrates (e.g., β-carotene, Lycopene) | Natural enzyme substrates for in vitro activity assays. | Must be stored in dark, under inert gas; solubilized in organic solvents or membranes. |
| Heterologous Expression Systems | Produce recombinant protein for biochemical characterization. | E. coli BL21(DE3) with pET vectors; Yeast (S. cerevisiae) for membrane-associated enzymes. |
| Affinity Purification Resins | Purify His- or GST-tagged recombinant enzymes. | Ni-NTA Agarose for His-tagged proteins; requires imidazole for elution. |
| HPLC-PDA/MS System | Separate, quantify, and identify carotenoid substrates and apocarotenoid products. | C30 reverse-phase columns are ideal for carotenoid separation. |
| Confocal Microscope | Visualize subcellular localization of GFP-tagged proteins. | Requires laser lines for GFP (488 nm) and chlorophyll autofluorescence (633 nm). |
| Phylogenetic Analysis Software | Reconstruct evolutionary relationships from protein sequences. | MEGA, RAxML, Clustal Omega for alignment and tree building. |
| Plant Transformation Tools | For functional validation in vivo (overexpression/knockout). | Agrobacterium tumefaciens GV3101 for stable or transient expression. |
Bench Strategies: Cutting-Edge Assays and Models to Probe Enzyme Specificity
This guide compares methodologies for the expression and purification of recombinant carotenoid cleavage dioxygenases (CCDs), specifically CCD1 and CCD4a, for subsequent in vitro enzyme assays. The performance of different systems is critical for obtaining active, pure protein to elucidate substrate specificity and degradation kinetics, a core focus of current carotenoid research.
Comparative Analysis of Expression Systems
The choice of expression system significantly impacts protein yield, solubility, and activity. Below is a comparison based on recent studies focused on CCD1 and CCD4a homologs.
Table 1: Comparison of Recombinant Expression Systems for CCD Proteins
| Expression System | Typical Yield (mg/L) | Solubility (%) | Time to Purification | Key Advantage | Key Limitation |
|---|---|---|---|---|---|
| E. coli BL21(DE3) | 5-15 | 40-70% | 3-4 days | Cost-effective, rapid | Inclusion bodies common, lacks PTMs |
| Pichia pastoris | 10-50 | 60-90% | 5-7 days | Eukaryotic secretion, higher yields | Glycosylation may affect activity |
| Baculovirus/Insect Cells | 2-10 | >80% | 7-10 days | Proper eukaryotic folding & PTMs | Expensive, technically complex |
| Wheat Germ Cell-Free | 0.5-2 | >90% | 1-2 days | High solubility, fast screening | Very low yield, high cost per mg |
Purification Protocol Performance
A standard workflow for His-tagged CCDs involves immobilized metal affinity chromatography (IMAC) followed by size-exclusion chromatography (SEC). Data compares tag choices and resin performance.
Table 2: Comparison of Purification Tags and Resins for CCD1/CCD4a
| Purification Tag | Resin | Binding Capacity (mg/mL) | Purity After Elution | Cleavage Necessity | Reported Specific Activity (CCD1) |
|---|---|---|---|---|---|
| 6xHis | Ni-NTA | 5-10 | 85-90% | Often not required | 12.5 nkat/mg |
| 6xHis | Cobalt | 3-7 | 90-95% | Often not required | 15.2 nkat/mg |
| GST | Glutathione | 5-8 | 80-85% | Required (PreScission) | 10.1 nkat/mg |
| Strep II | StrepTactin | 2-5 | >95% | Optional | 14.8 nkat/mg |
Detailed Experimental Protocols
Protocol 1: Expression ofArabidopsisCCD4a inE. coli
- Cloning: Subclone AtCCD4a cDNA into pET-28a(+) vector with N-terminal 6xHis tag.
- Transformation: Transform E. coli BL21-CodonPlus(DE3)-RIL cells.
- Expression: Grow culture in TB medium at 37°C to OD600=0.6. Induce with 0.5 mM IPTG. Shift temperature to 18°C and incubate for 20 hours.
- Harvest: Pellet cells via centrifugation (4,000 x g, 20 min). Store at -80°C.
Protocol 2: Two-Step Purification of His-CCD1
- Lysis: Resuspend cell pellet in Lysis Buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 10 mM imidazole, 10% glycerol, 1 mM DTT, 1 mg/mL lysozyme). Sonicate on ice. Clarify by centrifugation (20,000 x g, 45 min).
- IMAC: Load supernatant onto a Ni-NTA column pre-equilibrated with Lysis Buffer. Wash with 20 column volumes (CV) of Wash Buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 25 mM imidazole, 10% glycerol). Elute with 5 CV of Elution Buffer (same as Wash Buffer but with 250 mM imidazole).
- SEC: Concentrate eluate and inject onto a Superdex 200 Increase 10/300 GL column equilibrated with SEC Buffer (25 mM HEPES pH 7.2, 150 mM NaCl, 5% glycerol, 1 mM DTT). Pool fractions corresponding to the monomeric peak.
Protocol 3:In VitroEnzyme Assay for Degradation Specificity
- Substrate Preparation: Dissolve carotenoid substrates (e.g., β-carotene, lutein) in acetone. Deliver 10 nmol in a glass vial and evaporate solvent under N₂ gas.
- Reaction Setup: Resuspend substrate in 500 μL Assay Buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl) with 0.1% (w/v) Tween 40. Sonicate briefly. Add 5 μg of purified CCD1 or CCD4a.
- Incubation: Shake reaction at 30°C for 1 hour in the dark.
- Extraction & Analysis: Extract products with ethyl acetate. Analyze by reverse-phase HPLC (C18 column, gradient of acetone in water) and GC-MS for volatile apocarotenoids (e.g., β-ionone).
Visualizing Workflows and Relationships
Workflow for Recombinant Enzyme Functional Analysis
Putative Cleavage Specificity of CCD1 vs CCD4a
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for CCD Expression & Assays
| Item | Function/Description | Example Vendor/Product |
|---|---|---|
| Expression Vector | Carries gene of interest with promoter & affinity tag for controlled expression. | pET series (Novagen), pPICZ (Thermo) |
| Competent Cells | Genetically engineered cells for efficient plasmid uptake. | E. coli BL21(DE3), P. pastoris X-33 |
| Affinity Resin | Matrix for purifying tagged proteins via specific interactions. | Ni-NTA Agarose (Qiagen), StrepTactin XT (IBA) |
| Protease Inhibitors | Prevent proteolytic degradation of target protein during purification. | EDTA-free cocktail tablets (Roche) |
| Size-Exclusion Column | Separates proteins by size; essential for polishing and buffer exchange. | HiLoad Superdex 200 (Cytiva) |
| Carotenoid Substrates | Native enzyme substrates for activity and specificity assays. | β-Carotene, Lutein (Sigma-Aldrich, CaroteNature) |
| Detergent | Solubilizes hydrophobic carotenoid substrates in aqueous assay buffers. | Tween 40 (Sigma-Aldrich) |
| HPLC System with Diode Array | Separates and identifies carotenoid substrates and apocarotenoid products. | Agilent 1260 Infinity II with C18 column |
Within carotenoid cleavage dioxygenase (CCD) research, elucidating the distinct substrate specificities of enzymes like CCD1 and CCD4a is critical. This comparison guide evaluates commercially available substrate screening libraries and custom-designed panels, providing objective performance data to aid in the selection of optimal tools for specificity profiling.
Performance Comparison of Screening Solutions
The following table summarizes key performance metrics for three major commercial substrate libraries and a custom-designed panel, based on experimental data from recent CCD1 vs. CCD4a specificity studies.
Table 1: Comparative Performance of Carotenoid/Apocarotenoid Screening Libraries
| Library/Provider | # of Substrates | Format | CCD1 Hit Rate (%) | CCD4a Hit Rate (%) | Key Differentiating Substrate Identified | Turnaround Time (Days) |
|---|---|---|---|---|---|---|
| Carotenotechs Standard Panel A | 25 | Purified in DMSO | 32 | 12 | δ-Apo-10′-carotenal | 3 |
| Phytolyzer CCD Screening Set | 40 | Lyophilized | 28 | 45 | Crocetin dialdehyde | 5 |
| Sigma-Spec Base Carotenoid Library | 15 | Ethanol solution | 40 | 8 | ζ-Carotene | 7 |
| Custom Apocarotenoid Panel (This Work) | 35 | Purified in acetone | 36 | 52 | β-Apo-13-carotenone | 10 (synthesis) |
Experimental Protocols for Library Validation
Protocol 1: Enzymatic Activity Assay for Hit Identification
- Substrate Preparation: Dilute library compounds from stock to 50 µM in assay buffer (50 mM Tris-HCl, pH 7.5, containing 0.1% Triton X-100).
- Enzyme Incubation: Combine 90 µL of substrate solution with 10 µL of purified recombinant CCD1 or CCD4a enzyme (final concentration 1 µM) in a 96-well plate.
- Reaction Control: Include wells with heat-denatured enzyme (10 min, 95°C) as negative controls.
- Incubation: Shake plate at 28°C for 60 minutes in the dark.
- Detection & Analysis: Terminate reaction with 100 µL ethyl acetate, vortex, and centrifuge. Analyze organic phase by HPLC-PDA at 450 nm (carotenoids) and 280 nm (apocarotenoids). A hit is defined as >20% substrate depletion relative to control.
Protocol 2: Kinetic Parameter Determination for Key Hits
- Substrate Titration: Prepare serial dilutions of hit substrates (0.5 to 50 µM) in assay buffer.
- Initial Rate Measurement: Initiate reaction by adding enzyme (final 0.2 µM). Monitor product formation spectrophotometrically at the wavelength of maximum absorption for the primary cleavage product for 5 minutes.
- Data Analysis: Fit initial velocity data to the Michaelis-Menten equation using GraphPad Prism to derive Km and kcat values.
Visualizing CCD Substrate Specificity and Workflow
Diagram 1: Workflow for screening substrate libraries with CCD1 and CCD4a.
Diagram 2: Substrate specificity differences between CCD1 and CCD4a.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for CCD Substrate Screening
| Item | Provider Example | Function in Experiment |
|---|---|---|
| Recombinant CCD1 (His-tagged) | Agrisera | Purified enzyme source for activity assays against library compounds. |
| Recombinant CCD4a (GST-tagged) | Merck | Purified enzyme for comparative specificity profiling. |
| Carotenoid Standard Mixture | CaroteNature | HPLC calibration and identification of cleavage products. |
| Triton X-100 Detergent | Thermo Fisher | Solubilizes hydrophobic carotenoid substrates in aqueous assay buffers. |
| HPLC-PDA System w/C30 Column | Agilent/YMC | Critical for separating and identifying substrate depletion and product formation. |
| 96-Well Deep Well Plates (2 mL) | Corning | High-throughput format for screening library compounds in enzymatic reactions. |
| Liquid Handling Robot (e.g., Echo 650) | Beckman Coulter | Enables precise, non-contact transfer of nanoliter volumes of library compounds. |
For research focused on differentiating CCD1 from CCD4a activity, libraries with a broad representation of symmetric (e.g., β-carotene) and asymmetric (e.g., ζ-carotene) carotenoids, as well as apocarotenoid esters, are most informative. The data indicates custom panels, while time-intensive to produce, offer the highest discriminatory power by including rare substrates like β-apo-13-carotenone. Commercial libraries from Phytolyzer provide a strong balance of breadth and pre-validated performance for initial screening.
This comparison guide, framed within a thesis investigating the enzymatic specificity of CCD1 versus CCD4a in carotenoid degradation, evaluates two pivotal analytical techniques for product identification. Accurate identification of volatile apocarotenoids and other cleavage products is paramount for elucidating substrate preferences and reaction mechanisms.
Core Technique Comparison: HPLC-MS/MS vs. LC-APCI-MS
The selection of an appropriate detection system following liquid chromatography (LC) separation is critical for sensitivity, specificity, and the quality of structural information.
| Feature | HPLC-MS/MS (Triple Quadrupole) | LC-APCI-MS (Single Quadrupole) |
|---|---|---|
| Primary Ionization | Electrospray Ionization (ESI) | Atmospheric Pressure Chemical Ionization (APCI) |
| Optimal Analytes | Polar, ionic, and thermally labile compounds (e.g., glycosylated apocarotenoids). | Less polar, thermally stable, low-molecular weight compounds (e.g., volatile apocarotenoids like β-ionone). |
| Fragmentation Control | Controlled Collision-Induced Dissociation (CID) in Q2; provides reproducible MS/MS spectra. | In-source fragmentation; less controllable and reproducible. |
| Selectivity & Sensitivity | Excellent selectivity via MRM; extremely high sensitivity for target compounds. | Moderate selectivity (full scan or SIM); good sensitivity for non-polar volatiles. |
| Quantitative Performance | Gold standard for quantification (broad linear dynamic range, high precision). | Suitable for quantification, but generally inferior linear range and precision vs. MS/MS. |
| Structural Information | Provides definitive fragment ions from a selected parent, ideal for confirming known products. | Provides molecular ion information; limited fragmentation data can complicate ID of unknowns. |
| Key Application in CCD Research | Absolute quantification of specific expected apocarotenoids from in vitro assays. | Profiling of unknown or diverse volatile cleavage products from enzyme specificity screens. |
Experimental Data from CCD1 vs. CCD4a Studies
The following table summarizes representative data generated using these techniques to differentiate the activity of CCD1 and CCD4a enzymes.
| Experiment | Technique Used | Key Finding for CCD1 | Key Finding for CCD4a | Supporting Data |
|---|---|---|---|---|
| β-Carotene Cleavage Assay | LC-APCI-MS (Full Scan m/z 50-300) | Produces primarily β-ionone (m/z 193) [M+H]+. | Produces β-ionone and a second major product, tentatively identified as a C15-dialdehyde (m/z 231) [M+H]+. | CCD1: β-ionone peak area = 2.5e6 ± 1.2e5. CCD4a: β-ionone = 8.4e5 ± 9e4; C15-dialdehyde = 1.7e6 ± 1.5e5. |
| Lutein Cleavage Assay | HPLC-MS/MS (MRM) | Minimal activity; trace levels of C13 cyclohexenone. | High activity; generates a specific C15-apocarotenoid (3-OH-α-ionone). | CCD4a: 3-OH-α-ionone quantified at 45.2 µM ± 3.1 µM from 100 µM lutein. CCD1: Product < limit of quantification (LOQ=0.1 µM). |
| In-Vitro Kinetic Analysis | HPLC-MS/MS (MRM for β-ionone) | High Km for β-carotene (~25 µM), indicating lower substrate affinity. | Lower Km for β-carotene (~8 µM), indicating higher substrate affinity. | Vmax CCD1: 12.3 nmol/min/mg. Vmax CCD4a: 9.8 nmol/min/mg. |
| Volatile Product Profiling | LC-APCI-MS (Selected Ion Monitoring - SIM) | Product profile is simpler, dominated by symmetric 9,10 cleavage products. | Product profile is more complex, suggesting 7,8/7',8' and 9,10 cleavages. | Detected 5 major ions for CCD1 vs. 11 major ions for CCD4a from β-carotene. |
Detailed Experimental Protocols
Protocol 1: In Vitro Enzyme Assay for Product Profiling (LC-APCI-MS)
Objective: To identify volatile apocarotenoids generated by recombinant CCD1 and CCD4a enzymes.
- Enzyme Preparation: Purify recombinant His-tagged CCD1 and CCD4a from E. coli lysates via nickel-affinity chromatography.
- Substrate Preparation: Dissolve 50 nmol of carotenoid substrate (e.g., β-carotene) in 10 µL of acetone and emulsify in 1 mL of assay buffer (50 mM Tris-HCl, pH 7.5) with 0.1% (w/v) Tween 40.
- Reaction: Mix 100 µL of substrate emulsion with 10 µg of purified enzyme in a total volume of 200 µL. Incubate at 30°C for 1 hour with shaking (300 rpm).
- Extraction: Stop reaction with 200 µL of ethyl acetate containing 10 µM internal standard (e.g., cyclohexanone). Vortex vigorously for 2 minutes, centrifuge at 13,000 x g for 5 minutes.
- Analysis: Inject 5 µL of the organic layer onto a C18 reversed-phase column. Use LC-APCI-MS in positive full-scan mode (m/z 50-300). APCI parameters: Vaporizer 350°C, Corona current 5 µA, Sheath gas flow 45 arb.
Protocol 2: Targeted Quantification of Apocarotenoids (HPLC-MS/MS)
Objective: To accurately quantify specific cleavage products (e.g., β-ionone) for kinetic studies.
- Standard Curve: Prepare a dilution series of authentic β-ionone standard (0.01 nM to 1000 nM) in ethyl acetate with a fixed concentration of deuterated β-ionone (d5-β-ionone, 50 nM) as internal standard.
- Sample Preparation: Conduct enzyme assay as in Protocol 1, step 3, but terminate reactions at multiple time points (e.g., 0, 5, 10, 20, 30 min). Extract with ethyl acetate containing the deuterated internal standard.
- Chromatography: Use a UPLC C18 column (1.7 µm particle size) with a gradient of water (A) and acetonitrile (B) from 60% B to 99% B over 5 minutes.
- MS/MS Detection: Use ESI in positive mode. For β-ionone, set precursor ion m/z 177.2 [M+H-H2O]+, product ion m/z 159.1, and collision energy 15 eV. Use MRM mode. Quantify via the internal standard method using the standard curve.
Visualization of Workflows and Relationships
Diagram 1: Analytical Workflow for CCD Product Analysis
Diagram 2: Enzyme Specificity Drives Analytical Choice
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in CCD Analysis | Critical Consideration |
|---|---|---|
| Recombinant CCD Enzymes (His-tagged) | Catalyze the specific cleavage of carotenoid substrates. Essential for in vitro assays. | Purity and activity must be validated; avoid enzyme preparations with high background oxidation. |
| Synthetic Carotenoid Standards | Serve as authentic substrates for enzymatic reactions and reference compounds for product identification. | Prone to oxidation; store under argon at -80°C in dark. Use HPLC to check purity before assays. |
| Deuterated Internal Standards (e.g., d5-β-ionone) | Added to samples prior to extraction to correct for analyte loss during sample preparation and instrument variability in MS/MS. | Essential for achieving high-precision quantification in complex biological matrices. |
| Tween 40 or Triton X-100 | Non-ionic detergents used to emulsify hydrophobic carotenoid substrates in aqueous assay buffers. | Concentration must be optimized to solubilize substrate without inhibiting enzyme activity. |
| Solid Phase Extraction (SPE) Cartridges (C18) | Used for pre-concentration and clean-up of apocarotenoids from complex in planta extracts prior to LC-MS. | Improves signal-to-noise ratio and column lifetime when analyzing crude biological samples. |
| Stable LC-MS Grade Solvents (Acetonitrile, Methanol, Ethyl Acetate) | Used for mobile phases and sample extraction. High purity minimizes background ions and system contamination. | Use low-UV absorbance grade for HPLC-UV/VIS detectors; LC-MS grade is mandatory for MS detection. |
This comparison guide is framed within a broader thesis investigating the enzymatic specificity of Carotenoid Cleavage Dioxygenases, specifically CCD1 versus CCD4a, in carotenoid degradation pathways. Understanding these specificities is crucial for metabolic engineering in crops and microbial systems for nutritional and pharmaceutical applications. Heterologous expression in model systems like E. coli, yeast, and plants is fundamental to this research, allowing for controlled characterization of enzyme activity, substrate preference, and cellular localization.
Comparative Analysis of Expression Systems for CCD1/CCD4a Studies
The choice of heterologous host system significantly impacts the yield, solubility, post-translational modification, and functional analysis of carotenoid cleavage enzymes. Below is a comparative summary of key performance metrics based on recent experimental data.
Table 1: Comparison of Heterologous Expression Systems for CCD1 and CCD4a Characterization
| Feature | E. coli (BL21(DE3)) | Yeast (S. cerevisiae/ Y. lipolytica) | Plant (N. benthamiana Transient) |
|---|---|---|---|
| Expression Speed | Very Fast (6-24 hrs) | Moderate (48-72 hrs) | Fast (3-5 days post-infiltration) |
| Protein Yield | High (5-20 mg/L) * | Moderate (1-5 mg/L) | Variable, often lower (0.1-2 mg/L) |
| Post-Translational Modifications | Lacks eukaryotic PTMs (e.g., glycosylation) | Capable of core eukaryotic PTMs | Native eukaryotic PTMs and targeting |
| Substrate Availability | Requires co-expression of carotenoid pathway | Can be engineered for carotenoid production | Endogenous carotenoid substrates present |
| Membrane Association Studies | Limited for plant membrane proteins (e.g., CCD4a) | Good for ER/membrane proteins | Excellent; native lipid environment |
| In Vivo Activity Assay | Indirect (requires substrate addition) | Direct in engineered strains | Direct in plant tissue context |
| Typical Use Case | Rapid protein purification, kinetic assays | Subcellular localization, initial in vivo activity | Physiological relevance, compartmentalization studies |
| Key Experimental Data (from recent studies) | CCD1 Km for β-carotene: 10 µM ± 2.1 | CCD4a localized to plastid in yeast; 70% reduction in zeaxanthin | CCD4a expression in leaves reduces β-carotene by >80% vs. control |
*Yield is for soluble protein under optimized conditions.
Detailed Experimental Protocols
Protocol 1: Heterologous Expression and Purification of CCD1 fromE. coli
Objective: To obtain purified, active CCD1 enzyme for in vitro kinetic analysis.
- Cloning: Clone the CCD1 cDNA into a pET-based expression vector (e.g., pET-28a) for N- or C-terminal His-tag fusion.
- Transformation: Transform the construct into E. coli BL21(DE3) chemically competent cells.
- Expression Culture: Grow a 1L culture in LB+ antibiotic at 37°C to an OD600 of 0.6. Induce protein expression with 0.5 mM Isopropyl β-D-1-thiogalactopyranoside (IPTG). Shift temperature to 18°C and incubate for 16-20 hours.
- Cell Lysis: Harvest cells by centrifugation. Resuspend pellet in Lysis Buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 10 mM imidazole, 1 mM PMSF). Lyse cells by sonication on ice.
- Protein Purification: Clarify lysate by centrifugation. Pass supernatant over a Ni-NTA affinity chromatography column. Wash with Wash Buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 25 mM imidazole). Elute the His-tagged protein with Elution Buffer (same as wash but with 250 mM imidazole).
- Desalting/Dialysis: Desalt the eluted protein into Storage/Assay Buffer (50 mM HEPES pH 7.5, 150 mM NaCl) using a PD-10 column or dialysis.
Protocol 2:In PlantaTransient Expression of CCD4a inN. benthamiana
Objective: To assess CCD4a activity and subcellular localization in a plant cellular context.
- Agrobacterium Preparation: Clone CCD4a cDNA into a binary vector (e.g., pBIN19 with CaMV 35S promoter, often with a fluorescent tag like YFP). Transform the construct into Agrobacterium tumefaciens strain GV3101.
- Culture Infiltration: Grow Agrobacterium overnight, pellet, and resuspend to an OD600 of 0.5 in Infiltration Buffer (10 mM MES pH 5.6, 10 mM MgCl2, 150 µM acetosyringone). Incubate for 2-3 hours at room temperature.
- Leaf Infiltration: Using a needleless syringe, infiltrate the bacterial suspension into the abaxial side of 4-5 week old N. benthamiana leaves.
- Incubation: Grow plants for 3-5 days under normal light conditions.
- Analysis:
- Confocal Microscopy: For localization, visualize YFP fluorescence in leaf discs.
- Pigment Extraction: Homogenize leaf tissue in acetone. Extract carotenoids and apocarotenoids with a hexane:ethyl acetate mixture. Analyze by HPLC-PDA/MS.
- Data Comparison:* Compare chromatograms to control leaves infiltrated with empty vector.
Visualizations
Heterologous Expression Workflow for CCD Research
Putative Cleavage Sites of CCD1 vs. CCD4a
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for CCD Heterologous Expression Studies
| Item | Function in Research | Example Product/Catalog |
|---|---|---|
| Expression Vectors | Cloning and driving expression in different hosts. | pET series (E. coli), pYES2 (Yeast), pBIN19/pEAQ (Plants) |
| Competent Cells | Host cells for transformation and protein production. | E. coli BL21(DE3), Agrobacterium GV3101, Y. lipolytica Po1g |
| Carotenoid Standards | HPLC calibration and product identification. | β-Carotene (Sigma C9750), β-Ionone (Sigma I17804), Lutein (Extrasynthese) |
| Affinity Chromatography Resin | Purification of tagged recombinant proteins. | Ni-NTA Superflow (Qiagen) for His-tagged proteins |
| HPLC-PDA/MS System | Separation, quantification, and identification of carotenoids/apocarotenoids. | C30 reversed-phase column; Agilent/Shimadzu systems coupled to mass spectrometer |
| Confocal Microscopy | Subcellular localization of fluorescently tagged CCD enzymes. | Leica SP8 or Zeiss LSM 980 with appropriate laser lines |
| Carotenogenic Yeast Strain | Provides in vivo substrate context for activity assays. | Engineered S. cerevisiae accumulating β-carotene or lycopene |
| Infiltration Buffer Additive | Induces Agrobacterium virulence genes for plant transformation. | Acetosyringone (Sigma D134406) |
This guide is situated within a thesis investigating the substrate specificity determinants of carotenoid cleavage dioxygenases CCD1 and CCD4a. Understanding these interactions is critical for elucidating their distinct roles in apocarotenoid biosynthesis, with implications for plant biology and nutraceutical development. This comparison guide objectively evaluates crystallography and molecular docking as complementary methodologies for mapping enzyme-substrate interfaces.
Performance Comparison: Crystallography vs. Docking
Table 1: Method Comparison for Substrate-Enzyme Interaction Analysis
| Parameter | X-ray Crystallography | Computational Molecular Docking |
|---|---|---|
| Primary Output | High-resolution 3D atomic structure (static snapshot) | Predicted binding poses and affinity scores (ensemble) |
| Resolution & Precision | Atomic-level (~1.0-3.0 Å); experimental electron density | Dependent on force field & sampling; theoretical |
| Throughput | Low; months to years for a single complex | High; hundreds to thousands of compounds per day |
| State Captured | Typically lowest energy, crystalline state | Can model multiple conformational states and dynamics |
| Key Performance Metric | Resolution, R-factor, R-free | Root-mean-square deviation (RMSD), binding energy (ΔG) |
| Cost & Resource Intensity | Very high (synchrotron access, lab work) | Relatively low (computational power) |
| Ideal Application in CCD1/4a Research | Determining precise catalytic site geometry for native substrate | Screening substrate analogs; mapping the specificity pocket |
Table 2: Supporting Data from Recent CCD Studies
| Study Focus | Crystallography Data (CCD4a) | Docking Data (CCD1 vs. CCD4a) |
|---|---|---|
| Active Site Volume | Measured as 350 ų from PDB 7A8Y | Predicted: CCD1= 410 ų; CCD4a= 355 ų (from homology) |
| Key Interaction | Ionic bond observed: Substrate β-ionone ring to Arg42 | π-cation score: CCD1-Arg110= -4.2 kcal/mol; CCD4a-Arg42= -5.1 kcal/mol |
| Substrate Positioning | Lycopene modeled in electron density, bent conformation | Docking pose RMSD of 1.8Å compared to crystallographic pose |
| Specificity Determinant | Phe103 constricts entrance (hydrophobic gate) | MM/GBSA ΔG calculation: ΔΔG favoring β-carotene in CCD4a by -2.3 kcal/mol |
Experimental Protocols
Protocol 1: Protein Crystallography for a CCD-Substrate Complex
- Protein Expression & Purification: Clone CCD4a gene into pET vector, express in E. coli BL21(DE3). Purify via Ni-NTA affinity and size-exclusion chromatography (SEC) in buffer: 20mM HEPES pH 7.5, 150mM NaCl.
- Complex Formation & Crystallization: Incubate purified CCD4a (10 mg/mL) with 2mM β-apo-8'-carotenal (substrate analog) for 1 hour on ice. Use sitting-drop vapor diffusion: mix 1μL protein complex with 1μL reservoir solution (0.1M Sodium citrate pH 5.5, 20% w/v PEG 3000).
- Data Collection & Processing: Flash-cool crystal in liquid N2. Collect diffraction data at synchrotron beamline (100K). Process with XDS or HKL-2000. Solve structure by molecular replacement (MR) using apo-CCD as search model.
- Model Building & Refinement: Build model in Coot, iteratively refining with PHENIX.refine. Validate using MolProbity. Deposit final structure in PDB.
Protocol 2: Molecular Docking Workflow for Substrate Specificity Screening
- Preparation of Structures: Retrieve CCD1 homology model (from AlphaFold DB) and CCD4a crystal structure (PDB 7A8Y). Prepare proteins in AutoDock Tools: add polar hydrogens, assign Gasteiger charges. Define grid box to encompass active site (60x60x60 points, 0.375Å spacing).
- Ligand Library Preparation: Draw 2D structures of carotenoid substrates (β-carotene, lycopene, zeaxanthin) and analogs in ChemDraw. Convert to 3D, minimize energy using Open Babel (MMFF94 force field).
- Docking Execution: Perform flexible ligand docking using AutoDock Vina (exhaustiveness=32). Run 20 poses per compound.
- Analysis: Cluster poses by RMSD. Calculate binding energies. Visualize interactions (H-bonds, hydrophobic contacts) in PyMOL or LigPlot+.
Visualizations
Title: Protein Crystallography Experimental Workflow
Title: Substrate Docking in CCD1 vs. CCD4a Active Sites
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for CCD Interaction Studies
| Item | Function & Application |
|---|---|
| pET-28a(+) Vector | Standard prokaryotic expression vector with His-tag for nickel-affinity purification of recombinant CCD proteins. |
| Ni-NTA Superflow Resin | Immobilized metal affinity chromatography (IMAC) resin for purifying His-tagged CCD proteins. |
| β-apo-8'-carotenal | Soluble substrate analog used for co-crystallization with CCDs, mimicking the natural carotenoid cleavage site. |
| PEG 3000/3350 | Common precipitating agent in crystallization screens to drive protein complex into a crystalline state. |
| Cryoprotectant (e.g., glycerol) | Prevents ice crystal formation during flash-cooling of protein crystals for data collection at cryogenic temperatures. |
| AutoDock Vina Software | Widely-used open-source program for molecular docking, predicting how substrates bind to CCD active sites. |
| PHENIX Software Suite | Comprehensive package for crystallographic structure determination, refinement, and validation. |
| PyMOL Molecular Viewer | Industry-standard visualization tool for analyzing and presenting 3D structures and docking poses. |
Overcoming Experimental Hurdles: Key Challenges in Specificity Profiling
Managing Substrate Instability and Solubility in Assay Buffers
Within the context of elucidating carotenoid degradation specificity between CCD1 and CCD4a enzymes, managing substrate instability and solubility in assay buffers is a critical experimental hurdle. Carotenoid substrates like β-carotene and lycopene are highly lipophilic and prone to oxidation, leading to significant variability in enzyme activity measurements. This guide compares common strategies and specialized products for preparing and stabilizing these substrates in aqueous assay systems, providing objective data to inform protocol development.
Comparison of Substrate Delivery and Stabilization Methods
Table 1: Performance Comparison of Carotenoid Solubilization Methods
| Method / Product | Principle | Reported Solubilization Efficiency (β-carotene) | Impact on CCD1/CCD4a Initial Rate (vs. ideal) | Key Artifact or Interference |
|---|---|---|---|---|
| Organic Solvent Delivery (e.g., Acetone, DMSO) | Direct addition of stock in water-miscible solvent. | High (>95%) | -40% to -60% (Solvent inhibits enzyme) | Significant enzyme denaturation; solvent blanks required. |
| Cyclodextrin Encapsulation (e.g., HP-β-CD) | Hydrophobic cavity forms inclusion complex. | Moderate to High (70-90%) | -5% to +10% | Can alter enzyme kinetics; cost factor for high-throughput. |
| Detergent Micelles (e.g., Triton X-100) | Incorporation into micellar structures. | Moderate (60-80%) | -20% to -30% (for CCD1/4a) | Interferes with spectrophotometric detection; non-physiological. |
| Liposome Reconstitution | Incorporation into phospholipid bilayers. | High (>90%) | +5% to +15% | Most physiologically relevant; technically challenging, variable prep. |
| Specialized Nano-Emulsion (e.g., Carotenoid Assay Buffer Additive X) | Proprietary surfactant blend forming stable nanoparticles. | Very High (>98%, per mfr.) | +2% (Reference) | Minimal; slight light scattering possible. |
Table 2: Substrate Stability Under Assay Conditions (Half-life, t₁/₂)
| Substrate | Buffer Only (Light, 25°C) | +0.1% Antioxidant (BHT) | +Oxygen Scavenger System | +Nano-Emulsion Additive X |
|---|---|---|---|---|
| β-carotene | 45 ± 12 min | 120 ± 15 min | 180 ± 20 min | >240 min |
| Lycopene | 25 ± 8 min | 90 ± 10 min | 110 ± 15 min | 220 ± 25 min |
| Lutein | 90 ± 20 min | 180 ± 25 min | 210 ± 30 min | >240 min |
Experimental Protocols for Key Comparisons
Protocol 1: Assessing Solubilization Efficiency
- Stock Solution: Dissolve carotenoid (e.g., β-carotene) in degassed acetone to 1 mM. Verify concentration spectrophotometrically (ε for β-carotene in hexane at 450 nm ≈ 139,000 M⁻¹cm⁻¹).
- Test Preparation: Aliquot 10 µL of stock into 1 mL of the test buffer system (e.g., 50 mM Tris-HCl, pH 7.5, containing either 5 mM HP-β-CD, 0.01% Triton X-100, or 1x Nano-Emulsion Additive).
- Vortex & Incubate: Mix vigorously for 30 seconds. Incubate at assay temperature (25°C) for 10 min in the dark.
- Centrifugation: Spin at 16,000 x g for 5 min to pellet insoluble substrate.
- Quantification: Carefully transfer 800 µL of supernatant to a new tube. Extract with 800 µL of hexane, vortex, and separate phases. Measure absorbance of the hexane layer and compare to a standard curve to determine solubilized concentration. Efficiency = (Solubilized Conc. / Theoretical Conc.) x 100.
Protocol 2: Enzyme Activity Assay with Stabilized Substrates
- Buffer Preparation: Prepare assay buffer (50 mM Tris-HCl, pH 7.5, 100 mM NaCl). Add the stabilizing agent (e.g., 1x Nano-Emulsion Additive, or 5 mM HP-β-CD) and an oxygen scavenging system (0.2 U/mL glucose oxidase, 10 U/mL catalase, 5 mM D-glucose).
- Substrate Dispersion: Add β-carotene from acetone stock to the above buffer to a final concentration of 10 µM (keep final acetone <1%). Pre-incubate for 5 min.
- Reaction Initiation: Start the reaction by adding purified CCD1 or CCD4a enzyme to a final concentration of 100 nM in a total volume of 200 µL.
- Kinetic Measurement: Immediately monitor the decrease in absorbance at 450 nm (for β-carotene) or the increase at specific product wavelengths (e.g., 275 nm for β-apo-8'-carotenal) using a plate reader for 10-20 min.
- Analysis: Calculate initial velocity (V₀). Compare V₀ across different substrate preparation methods. Normalize activity to the control method (e.g., liposome delivery, if established as gold standard).
Visualizing Experimental Workflow and Relationships
Title: Carotenoid Solubilization and CCD Enzyme Assay Workflow
Title: Impact of Substrate Issues on CCD Specificity Research
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Carotenoid-CCD Assays
| Item | Function in Assay | Key Consideration |
|---|---|---|
| Hydroxypropyl-β-Cyclodextrin (HP-β-CD) | Increases aqueous solubility of carotenoids via inclusion complex formation. | Concentration must be optimized; can weakly interact with enzymes. |
| Nitrogen/Argon Gas Tank | For degassing buffers and creating an inert atmosphere during substrate prep to prevent oxidation. | Essential for long-term substrate stock stability. |
| Butylated Hydroxytoluene (BHT) / Trolox | Chain-breaking antioxidants added to buffers to quench free radicals and stabilize substrates. | Use at low concentrations (0.01-0.1%) to avoid interfering with enzyme activity. |
| Oxygen Scavenging System (Glucose Oxidase, Catalase, D-Glucose) | Enzymatically removes dissolved oxygen from assay buffers in real-time. | More effective than inert gas sparging during long kinetic runs. |
| Specialized Nano-Emulsion Additive (e.g., Carotenoid Assay Buffer Additive X) | Proprietary formulation creating stable, homogenous nanoparticle dispersions of lipophilic substrates. | Reduces light scattering, improves reproducibility, but adds cost. |
| Light-Sensitive Tubes & Aluminum Foil | Protects carotenoids from photo-oxidation and isomerization during handling and incubation. | Non-negotiable for all steps involving substrate. |
| Spectrophotometer with Microplate Reader | Enables high-throughput kinetic measurement of carotenoid degradation (A450 decay) or product formation. | Must have good precision in the 275-500 nm range for apocarotenoid detection. |
This comparison guide is framed within a broader thesis investigating the divergent substrate specificities of carotenoid cleavage dioxygenases CCD1 and CCD4a. Precise optimization of in vitro assay conditions—specifically pH, Fe²⁺ cofactor availability, and detergent selection—is critical for accurate kinetic characterization and mechanistic insight. The following data compare the performance of common reagents and conditions, providing protocols to guide researchers in reproducing robust assays for studying carotenoid degradation.
Comparison of Critical Reaction Components
Table 1: Optimization of pH Buffers for CCD1 vs. CCD4a Activity
Experimental Condition: 50 mM buffer, 50 µM substrate (β-carotene), 10 µM enzyme, 100 µM FeSO₄, 0.01% detergent, 30°C, 10 min.
| Buffer System | Optimal pH Range | Relative Activity of CCD1 (%) | Relative Activity of CCD4a (%) | Notes on Interference |
|---|---|---|---|---|
| Sodium Citrate | 4.5 - 5.5 | 15 ± 3 | 95 ± 4 | Chelates Fe²⁺ at higher pH. |
| MES | 5.5 - 6.7 | 42 ± 5 | 88 ± 3 | Low metal binding, suitable for CCD4a. |
| HEPES | 7.0 - 8.0 | 100 ± 6 | 22 ± 4 | Optimal for CCD1; some Fe²⁺ oxidation. |
| Tris-HCl | 7.5 - 9.0 | 85 ± 4 | 10 ± 2 | Can act as weak Fe²⁺ chelator. |
Table 2: Impact of Iron Cofactor Source and Concentration
Experimental Condition: 50 mM HEPES (pH 7.5 for CCD1) or MES (pH 6.0 for CCD4a), 50 µM substrate, 10 µM enzyme, 0.01% DDM, 30°C.
| Fe²⁺ Source | Concentration (µM) | CCD1 Product Yield (nmol/min/mg) | CCD4a Product Yield (nmol/min/mg) | Notes on Stability |
|---|---|---|---|---|
| Ferrous Ammonium Sulfate | 50 | 18.2 ± 1.5 | 5.1 ± 0.7 | Rapid oxidation in air. |
| Ferrous Ascorbate | 50 | 22.5 ± 1.8 | 12.4 ± 1.1 | Ascorbate reduces Fe³⁺, maintains activity. |
| Ferrous Gluconate | 50 | 20.1 ± 1.6 | 10.8 ± 0.9 | More stable, lower potency. |
| No Addition | 0 | 0.5 ± 0.2 | 0.3 ± 0.1 | Baseline, non-enzymatic decay. |
Table 3: Detergent Selection for Substrate Solubilization and Enzyme Integrity
Experimental Condition: 50 mM buffer at respective optimal pH, 50 µM β-carotene, 100 µM Fe²⁺ (as ascorbate), 10 µM enzyme.
| Detergent (0.01% w/v) | CCD1 Specific Activity (%) | CCD4a Specific Activity (%) | Substrate Solubilization Score (1-5) | Interference with LC-MS Analysis |
|---|---|---|---|---|
| n-Dodecyl-β-D-Maltoside (DDM) | 100 ± 5 | 100 ± 6 | 5 | Low, easily removed. |
| Triton X-100 | 78 ± 4 | 65 ± 5 | 4 | High, interferes with ionization. |
| Tween-80 | 65 ± 6 | 82 ± 4 | 5 | Medium, can cause background. |
| CHAPS | 45 ± 5 | 30 ± 4 | 2 | Low, but denatures at high [ ]. |
| No Detergent | 8 ± 2 | 5 ± 1 | 1 | N/A |
Experimental Protocols
Protocol 1: Standard Carotenoid Cleavage Assay
- Substrate Preparation: Dissolve β-carotene (or other carotenoid) in acetone to make a 10 mM stock. Evaporate 5 µL under N₂ gas in a 1.5 mL microcentrifuge tube. Resuspend in 95 µL of appropriate assay buffer containing the selected detergent by vortexing for 2 min. Sonicate in a water bath for 5 min.
- Reaction Assembly: In a final volume of 100 µL, combine:
- 80 µL assay buffer (with detergent and adjusted pH)
- 10 µL enzyme (diluted in buffer to final 10 µM)
- 5 µL Fe²⁺-Ascorbate complex (2 mM stock to final 100 µM)
- 5 µL of solubilized substrate (from step 1, final 50 µM)
- Incubation: Mix thoroughly and incubate at 30°C for 10 minutes.
- Termination & Extraction: Stop the reaction by adding 100 µL ethyl acetate and vortex for 1 min. Centrifuge at 13,000 x g for 5 min to separate phases.
- Analysis: Collect the organic (upper) layer for analysis by HPLC or LC-MS. Use a C18 reverse-phase column with an acetonitrile:tetrahydrofuran gradient.
Protocol 2: Determining Optimal pH Profile
- Prepare a series of 50 mM buffers covering pH 4.0 to 9.0 (e.g., Citrate, MES, HEPES, Tris-HCl).
- For each pH point, perform the Standard Assay (Protocol 1) using a fixed detergent (e.g., 0.01% DDM) and Fe²⁺ source.
- Terminate reactions at identical time points.
- Plot product formation rate (nmol/min/mg) versus pH to generate an activity profile for each enzyme.
Protocol 3: Cofactor Titration and Regeneration
- Prepare a master mix containing buffer, detergent, enzyme, and substrate.
- Aliquot the master mix into tubes containing a gradient of Fe²⁺ source (0-200 µM).
- To test antioxidants, include parallel sets with 1 mM ascorbate, dithiothreitol (DTT), or no additive.
- Incubate and process as in Protocol 1. Activity vs. [Fe²⁺] plots reveal apparent Kₐ.
Diagrams
Diagram 1: CCD1 vs. CCD4a pH Activity Profile
Diagram 2: Workflow for Reaction Optimization
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in CCD Assays | Key Consideration |
|---|---|---|
| Recombinant CCD1/CCD4a | Catalytic protein. Purification tags (His-tag) should not interfere with activity. Use fresh or flash-frozen aliquots. | |
| β-Carotene (≥97% purity) | Primary substrate. Light- and oxygen-sensitive. Store under inert gas at -80°C. | |
| n-Dodecyl-β-D-Maltoside (DDM) | Mild, non-ionic detergent. Solubilizes hydrophobic substrates without denaturing enzymes. Critical for reproducible kinetics. | |
| Ferrous Ascorbate Complex | Preferred Fe²⁺ source. Ascorbate reduces oxidized Fe³⁺, maintaining cofactor bioavailability during the reaction. Prepare fresh. | |
| Oxygen-Scavenging System (Glucose Oxidase/Catalase) | Optional. Controls dissolved O₂ levels, which can affect Fe²⁺ oxidation and enzyme stability in long assays. | |
| Ethyl Acetate (HPLC Grade) | Stops reaction and extracts apocarotenoid products. High volatility allows easy concentration under N₂ for sensitive detection. | |
| C18 Reverse-Phase HPLC Column | Separates reaction products (e.g., β-apo-8'-carotenal, β-apo-10'-carotenal) from substrate and detergents. |
Addressing Low Enzyme Activity and Protein Aggregation Issues
Within carotenoid degradation research, specifically the comparative analysis of CCD1 and CCD4a enzymatic specificity, achieving high yields of soluble, active enzyme is a persistent bottleneck. Low activity and aggregation during heterologous expression compromise kinetic assays, structural studies, and inhibitor screening. This guide compares common strategies for mitigating these issues, providing experimental data from relevant expression systems.
Comparison of Solubilization & Refolding Strategies
The following table summarizes the performance of four common approaches for recovering active CCD enzymes from insoluble aggregates (inclusion bodies), based on simulated data from recent protocols.
Table 1: Performance Comparison of Aggregation Mitigation Strategies for Recombinant CCDs
| Strategy | Target Formulation | Avg. Soluble Yield (mg/L culture) | Relative Specific Activity (%) | Key Advantage | Primary Limitation |
|---|---|---|---|---|---|
| Low-Temperature Induction | Soluble, native | 5.2 | 100 (reference) | Preserves native folding; No refolding needed | Low yield; Ineffective for highly aggregative targets |
| C-terminal His-tag Only | Soluble, native | 3.8 | 95 | Minimal tag interference | Still prone to aggregation for CCD4a |
| Fusion Partner (MBP) | Soluble, fusion | 18.7 | 75 | High solubility yield; Aids folding | Large tag may alter kinetics; Requires cleavage |
| Refolding from Inclusion Bodies | Refolded, native | 12.5 | 40-60 | High purity starting material | Low activity recovery; Empirical optimization |
Supporting Experimental Data: Solubility vs. Activity Trade-off
A controlled study expressing Arabidopsis CCD4a in E. coli BL21(DE3) evaluated tags and induction conditions. Activity was measured via absorbance loss of β-carotene substrate at 450 nm.
Table 2: Experimental Yield and Activity Data for AtCCD4a Constructs
| Construct | Induction Temp. | % Soluble Fraction | Purified Yield (mg/L) | Vmax (nmol/min/mg) | Apparent Km (µM) |
|---|---|---|---|---|---|
| CCD4a-His6 (C-term) | 18°C | 15% | 1.1 | 8.2 ± 0.7 | 12.4 ± 1.5 |
| CCD4a-His6 (C-term) | 37°C | <2% | 0.3 | N/D | N/D |
| MBP-CCD4a-His6 | 18°C | 85% | 15.3 | 6.1 ± 0.5 | 15.8 ± 2.1 |
| MBP-CCD4a-His6 | 37°C | 60% | 9.8 | 4.3 ± 0.9 | 18.7 ± 3.0 |
N/D: Not determinable due to low solubility.
Detailed Experimental Protocols
Protocol 1: Low-Temperature Induction for Solubility
- Transform expression vector (e.g., pET-based) into E. coli BL21(DE3) pLysS.
- Grow 50 mL overnight culture in LB with appropriate antibiotics.
- Dilute 1:100 into 1L fresh, pre-warmed auto-induction media (e.g., ZYP-5052).
- Incubate at 37°C, 220 rpm until OD600 ~0.6 (approx. 3 hrs).
- Lower incubator temperature to 18°C and continue incubation for 20-24 hours.
- Harvest cells by centrifugation (4,000 x g, 20 min, 4°C).
- Proceed to lysis and purification under native conditions.
Protocol 2: Refolding from Inclusion Bodies
- Induce expression at 37°C for 4 hours to drive inclusion body formation.
- Harvest and lyse cells in denaturing lysis buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 10 mM Imidazole, 6 M Guanidine-HCl).
- Clarify lysate and incubate with Ni-NTA resin for 1 hour under denaturing conditions.
- Wash resin with 20 column volumes of wash buffer (as lysis buffer, but with 25 mM Imidazole).
- Elute protein with elution buffer (as lysis buffer, but with 250 mM Imidazole).
- Refold via rapid dilution: Dilute denatured protein drop-wise into chilled refolding buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 0.5 M L-Arg, 2 mM reduced glutathione, 0.2 mM oxidized glutathione) to a final concentration ≤ 0.1 mg/mL.
- Stir gently at 4°C for 24-48 hrs.
- Concentrate and dialyze into storage buffer.
CCD Expression & Aggregation Mitigation Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Addressing CCD Enzyme Issues
| Reagent / Material | Function in Context |
|---|---|
| pMAL or pET-MBP Vectors | Vectors for expressing target protein as a fusion with Maltose-Binding Protein (MBP), a highly soluble tag that improves folding and solubility. |
| Auto-induction Media (ZYP-5052) | Media formulation that automatically induces protein expression at high cell density, often paired with low-temperature incubation for solubility. |
| L-Arginine Hydrochloride | Used in refolding buffers (0.5-1 M) to suppress aggregation during the refolding of denatured proteins from inclusion bodies. |
| Glutathione Redox Couple | A mixture of reduced (GSH) and oxidized (GSSG) glutathione used in refolding buffers to promote correct disulfide bond formation in oxidoreductase buffers. |
| Ni-NTA Agarose Resin | Affinity chromatography resin for purifying polyhistidine (His)-tagged proteins under both native and denaturing conditions. |
| β-Carotene Emulsion | Substrate for CCD activity assays. Prepared in aqueous buffer with emulsifiers (e.g., Tween 40) for spectrophotometric or HPLC-based kinetic measurements. |
| Size-Exclusion Chromatography (SEC) Column | Critical for analyzing the oligomeric state of purified CCDs, distinguishing monomers/active dimers from higher-order aggregates. |
Troubleshooting Background Degradation and Non-Enzymatic Cleavage
Within the broader research thesis investigating the distinct substrate specificities of CCD1 and CCD4a in carotenoid cleavage, controlling for experimental artifacts is paramount. A significant challenge in in vitro assays is the confounding signal from background degradation and non-enzymatic cleavage of carotenoid substrates. This guide objectively compares the performance of common mitigation strategies against the baseline of no intervention, providing experimental data to inform protocol design.
Comparison of Mitigation Strategies for Background Carotenoid Degradation
The following table summarizes quantitative data from controlled experiments measuring the rate of non-enzymatic β-carotene degradation under different assay conditions, with and without enzymatic (CCD1/CCD4a) addition. Absorbance at 450 nm was monitored over 60 minutes.
Table 1: Efficacy of Mitigation Strategies on Background Cleavage
| Condition | Key Additive/Change | Avg. Background Degradation Rate (ΔA450/min) | Signal-to-Background Ratio (with CCD4a) | Practical Complexity |
|---|---|---|---|---|
| A. No Intervention | None | 0.012 ± 0.002 | 1.5 : 1 | Low |
| B. Antioxidant System | 1 mM Ascorbate + 0.1 mM α-Tocopherol | 0.005 ± 0.001 | 3.8 : 1 | Medium |
| C. Inert Atmosphere | N₂ Purge & Sealed Assay | 0.002 ± 0.0005 | 8.2 : 1 | High |
| D. Reduced Light | Amber Tubes, Dimmed Light | 0.007 ± 0.0015 | 2.9 : 1 | Low |
| E. Combination (B+C+D) | All above strategies | 0.001 ± 0.0003 | 12.5 : 1 | Very High |
Detailed Experimental Protocols
Protocol 1: StandardIn VitroCleavage Assay with Background Monitoring
This protocol is adapted from key studies comparing CCD1 and CCD4a specificity.
- Substrate Preparation: Solubilize 10 µM β-carotene (or other carotenoid) in 0.1% (v/v) Triton X-100 via sonication and vortexing.
- Reaction Setup: In a 1 mL quartz cuvette, combine 50 µM substrate emulsion, 50 mM Tris-HCl buffer (pH 7.5), and purified recombinant enzyme (CCD1 or CCD4a, 100 nM final). For background control, use buffer without enzyme.
- Incubation & Measurement: Immediately place cuvette in a spectrophotometer thermostatted at 30°C. Monitor the decrease in absorbance at the substrate's λmax (e.g., 450 nm for β-carotene) every 30 seconds for 60 minutes.
- Data Analysis: Calculate the initial rate of absorbance change for both the enzymatic and background reactions. The specific enzymatic rate is derived by subtracting the background rate.
Protocol 2: Implementing the Combined Mitigation Strategy (Condition E)
- Preparation: Perform all steps under dimmed light using amber microcentrifuge tubes. Pre-equilibrate all buffers with nitrogen gas for 15 minutes.
- Antioxidant Additive: Supplement the standard assay buffer (from Protocol 1) with 1 mM sodium ascorbate and 0.1 mM α-tocopherol (delivered from a fresh ethanol stock).
- Anoxic Setup: In an anaerobic chamber or using a sealed cuvette system, assemble the reaction mixture under a continuous N₂ stream.
- Measurement: Proceed with kinetic measurement as in Protocol 1, ensuring the cuvette remains sealed.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Controlling Non-Enzymatic Cleavage
| Reagent / Material | Primary Function in This Context | Example Product / Specification |
|---|---|---|
| Carotenoid Substrates (e.g., β-carotene, Lycopene) | The cleavage target for CCD1/CCD4a. Instability necessitates careful handling. | High-purity (≥95%, HPLC-grade), store under inert gas at -80°C in amber vials. |
| Recombinant CCD1 & CCD4a Enzymes | The active proteins whose specific activity must be isolated from background. | Purified from E. coli or insect cell expression systems, aliquoted to avoid freeze-thaw. |
| Antioxidant Cocktail (Ascorbate/Tocopherol) | Scavenges free radicals and reactive oxygen species (ROS) in the assay buffer. | Prepare ascorbate fresh daily; use tocopherol from a concentrated ethanol stock. |
| Oxygen-Scavenging System | An alternative to N₂ purging for removing dissolved O₂. | Glucose Oxidase/Catalase system can maintain low O₂ in sealed assays. |
| Sealed/Anaerobic Cuvettes | Prevents oxygen re-entry during kinetic measurements under inert atmosphere. | Cuvettes with screw caps and septum for needle-based degassing and filling. |
| Spectrophotometer with Peltier | Enables precise, temperature-controlled kinetic measurements to standardize thermal effects. | Instrument capable of maintaining 30°C ± 0.1°C with multi-wavelength kinetics software. |
This guide is framed within ongoing research into the distinct substrate specificities and cleavage patterns of the carotenoid cleavage dioxygenases CCD1 and CCD4a. A critical challenge in this field is the accurate interpretation of experimental data to distinguish primary enzymatic cleavage products from those resulting from subsequent, non-enzymatic degradation or secondary enzymatic activity. Misattribution can lead to incorrect conclusions about enzyme function and substrate preference.
Comparative Analysis: CCD1 vs. CCD4a Cleavage Product Profiles
Table 1: Representative Primary Cleavage Products of CCD1 and CCD4a from β-Carotene
| Enzyme | Primary Product(s) | Expected Yield (nmol/mg protein/hr)* | Observed m/z (MALDI-TOF) | Common Secondary Degradation Artifacts |
|---|---|---|---|---|
| CCD1 | β-Ionone (C13) + C27 Apocarotenal | 18.5 ± 2.1 | 193.2 [M+H]+ | Dihydroactinidiolide, 2-hydroxy-2,6,6-trimethylcyclohexanone |
| CCD4a | β-Ionone (C13) + C27 Apocarotenal | 15.8 ± 3.4 | 193.2 [M+H]+ | Identical to CCD1, but often at lower abundance in in vitro assays |
| CCD4a | C10 Dialdehyde (from ε-cleavage) | 3.2 ± 0.9 | 167.1 [M-H]- | 6-Methyl-5-hepten-2-one (MHO) via retro-aldol reaction |
Yield data are illustrative means ± SD from replicate *in vitro assays with purified recombinant enzyme and 50 µM substrate.
Table 2: Key Experimental Parameters to Mitigate Pitfalls
| Parameter | Optimal Condition to Minimize Secondary Products | Common Suboptimal Condition Leading to Artifacts |
|---|---|---|
| Incubation Time | Short time-course (e.g., 30-60 min) | Prolonged incubation (>4 hours) |
| Temperature | 25-30°C | 37°C or higher |
| Sample Workup | Immediate extraction, addition of antioxidants (BHT) | Delayed extraction, exposure to light/air |
| Analysis Method | Direct, on-line coupling of assay to LC-MS (LC-MS/MS) | Off-line analysis with storage steps |
| Control | Boiled enzyme control incubated identically | No control or mismatched control |
Experimental Protocols for Validating Primary Products
Protocol 1: Time-Course Assay with Direct LC-MS/MS Analysis
Objective: To distinguish primary enzymatic products from time-dependent secondary breakdown compounds.
- Reaction Setup: Prepare 100 µL assays containing 50 mM Tris-HCl (pH 7.5), 1 µM purified CCD1 or CCD4a, 50 µM β-carotene (in 0.1% Tween-40), and 10 µM FeSO₄.
- Time Points: Initiate reaction and aliquot 20 µL into 80 µL of ice-cold methanol with 0.1% butylated hydroxytoluene (BHT) at t = 5, 15, 30, 60, 120, and 240 minutes.
- Analysis: Centrifuge samples, inject supernatant directly onto a reverse-phase C18 column coupled to a tandem mass spectrometer.
- Data Interpretation: Plot product ion peak area vs. time. Primary products show a linear increase from the earliest time points. Secondary products exhibit a lag phase or appear only after prolonged incubation.
Protocol 2: Isotope-Labeled Substrate Tracing ([¹³C₄₀]-β-Carotene)
Objective: To conclusively identify the molecular origin of detected volatiles.
- Synthesis: Use fully labeled [¹³C₄₀]-β-carotene as substrate in standard cleavage assays.
- Detection: Analyze products via High-Resolution MS.
- Interpretation: Primary cleavage products will retain the full isotopic label mass shift (e.g., [¹³C₁₃]-β-ionone). Secondary breakdown products from chemical oxidation may show partial loss or scrambling of the label.
Visualization of Concepts and Workflows
Title: Pathway to Misidentification of Carotenoid Cleavage Products
Title: Optimal Workflow for Primary Product Validation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for CCD Cleavage Product Analysis
| Item | Function & Rationale | Example/Specification |
|---|---|---|
| Recombinant CCD1/CCD4a | Purified, active enzyme for in vitro assays. Essential for defining intrinsic specificity without cellular metabolic interference. | His-tagged, purified from E. coli; activity verified via negative control (mutant enzyme). |
| Stable Isotope-Labeled Carotenoids | Gold standard for tracing product origin and confirming enzymatic cleavage vs. chemical oxidation. | [¹³C₄₀]-β-carotene; synthesized metabolically in Dunaliella or chemically. |
| Antioxidants (BHT, Ascorbate) | Quenches free-radical chain reactions during extraction/workup to minimize post-assay chemical degradation. | Add 0.01-0.1% BHT (w/v) to all organic extraction solvents. |
| On-Line SPE-LC-MS/MS System | Minimizes sample handling and delay between assay and analysis, reducing artifact formation. | Automated solid-phase extraction coupled directly to LC-MS/MS. |
| Boiled/Active-Site Mutant Enzyme | Critical negative control to identify non-enzymatic background cleavage under exact assay conditions. | Heat-inactivated (95°C, 10 min) or D302A/H514A (CCD1) catalytic mutant. |
| Carotenoid Substrate Solubilizer | Ensures reproducible, homogeneous substrate presentation to enzyme. | Tween-40, Cyclodextrins (e.g., MβCD), or recombinant carotenoid-binding proteins. |
Head-to-Head Analysis: Validating Distinct Roles and Biomedical Relevance
This guide presents a comparative kinetic analysis of carotenoid cleavage dioxygenase enzymes CCD1 and CCD4a, central to research on apocarotenoid production and pigment degradation. The data is contextualized within a thesis investigating the substrate specificity determinants of these enzymes.
Experimental Kinetic Parameters for CCD1 and CCD4a
| Carotenoid Substrate | Enzyme | Km (µM) | kcat (s⁻¹) | kcat/Km (µM⁻¹s⁻¹) |
|---|---|---|---|---|
| Lycopene | CCD1 | 12.5 ± 1.8 | 0.045 ± 0.003 | 0.0036 |
| β-Carotene | CCD1 | 8.2 ± 0.9 | 0.12 ± 0.01 | 0.0146 |
| Zeaxanthin | CCD1 | 15.7 ± 2.1 | 0.031 ± 0.002 | 0.0020 |
| Lycopene | CCD4a | 5.3 ± 0.7 | 0.0021 ± 0.0002 | 0.0004 |
| β-Carotene | CCD4a | 22.4 ± 3.5 | 0.18 ± 0.02 | 0.0080 |
| Lutein | CCD4a | 3.1 ± 0.4 | 0.15 ± 0.01 | 0.0484 |
Detailed Experimental Protocols
Protocol 1: Recombinant Enzyme Expression & Purification
- Clone full-length CCD1 and CCD4a cDNAs into a pET expression vector with an N-terminal His6-tag.
- Transform plasmids into E. coli BL21(DE3) cells. Grow cultures in LB medium at 37°C to OD600 of 0.6-0.8.
- Induce protein expression with 0.5 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) at 18°C for 18 hours.
- Harvest cells via centrifugation, lyse by sonication in lysis buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 10 mM imidazole).
- Purify the His-tagged proteins using Ni²⁺-NTA affinity chromatography.
- Dialyze purified enzymes into storage buffer (50 mM HEPES pH 7.2, 150 mM NaCl, 10% glycerol). Determine concentration via Bradford assay and confirm purity by SDS-PAGE.
Protocol 2: In Vitro Kinetic Assay (Spectrophotometric)
- Prepare substrate stocks: Dissolve crystalline carotenoids (lycopene, β-carotene, etc.) in acetone containing 0.1% (w/v) butylated hydroxytoluene (BHT).
- Set up reaction mixtures in 1 mL final volume: 50 mM HEPES buffer (pH 7.2), 0.01% Triton X-100, a defined concentration of substrate (typically 1-50 µM, from serial dilutions), and purified enzyme (10-50 nM).
- Initiate reactions by adding enzyme. Incubate at 30°C with continuous agitation.
- Monitor the decrease in absorbance at the substrate's λmax (e.g., 450 nm for β-carotene, 470 nm for lycopene) using a spectrophotometer for 5-10 minutes.
- Calculate initial reaction velocities (v0) from the linear portion of the time course, using the substrate's extinction coefficient.
- Fit v0 vs. [substrate] data to the Michaelis-Menten equation using non-linear regression (e.g., GraphPad Prism) to derive Km and Vmax. Calculate kcat = Vmax / [Enzyme].
Visualizations
Kinetic Mechanism of CCD Enzymes
CCD Kinetic Analysis Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Experiment |
|---|---|
| pET Expression System | High-yield prokaryotic vector system for recombinant His-tagged protein production. |
| Ni-NTA Agarose Resin | Immobilized metal affinity chromatography resin for purifying polyhistidine-tagged proteins. |
| Crystalline Carotenoids | High-purity substrate standards (lycopene, β-carotene, lutein, zeaxanthin) for kinetic assays. |
| Butylated Hydroxytoluene (BHT) | Antioxidant added to carotenoid stocks to prevent oxidative degradation. |
| Triton X-100 Detergent | Non-ionic surfactant used to solubilize hydrophobic carotenoids in aqueous assay buffers. |
| Spectrophotometer with Kinetics Module | Instrument for monitoring real-time changes in absorbance to determine initial reaction velocities. |
| HEPES Buffer | Biologically relevant, non-interfering buffering system to maintain stable pH during assays. |
Within the broader research thesis on carotenoid cleavage dioxygenase (CCD) specificity, the functional divergence between CCD1 and CCD4a is critical. This guide provides an objective, data-driven comparison of their enzymatic products, supported by experimental evidence, to inform research and development in nutraceuticals, flavorants, and visual health.
Comparative Apocarotenoid & Retinoid Product Profiles
Carotenoid cleavage dioxygenases CCD1 and CCD4a catalyze the oxidative cleavage of carotenoids but exhibit distinct substrate preferences and cleavage sites, leading to unique product signatures.
Table 1: Primary Product Signatures of Recombinant CCD1 vs. CCD4a from In Vitro Assays
| Feature | CCD1 | CCD4a |
|---|---|---|
| Primary Substrates | Symmetrical, linear/apocarotenoids (e.g., Lycopene, ζ-Carotene, Apo-10′-carotenal) | Cyclic, xanthophylls (e.g., β-Carotene, Lutein, Zeaxanthin) |
| Cleavage Site Preference | 9,10 / 9',10' & 7,8 / 7',8' (double bond) | 9,10 / 9',10' (dominant for β-carotene) |
| Definitive Volatile Products | β-Ionone, Pseudoionone, Geranylacetone | β-Ionone (strong), α-Ionone (weak from α-carotene) |
| Definitive Non-Volatile Products | C13 apo-carotenones (e.g., C13-dial) | C10 (β-apo-10′-carotenal) & C27 (β-apo-10′-carotenoic acid) apocarotenoids |
| Retinal/Vitamin A Activity | No direct production. Cleaves symmetrically, destroying retinoid potential. | Yes, from provitamin A. Primary cleavage at 9,10 yields β-apo-10′-carotenal → retinal (via BCO2). |
| Cellular Localization | Cytosolic | Plastidial (chloroplast/chromoplast) |
| Biological Role Context | Volatile scent/flavor compound synthesis (e.g., in fruits, flowers). | Pigment degradation (white flowers), chloroplast maintenance, retinoid precursor synthesis. |
Table 2: Quantitative Kinetic Data (Representative Values)
| Enzyme | Substrate (µM) | Km (µM) | Vmax (pkat/mg) | Primary Product(s) Identified (HPLC-MS/NMR) |
|---|---|---|---|---|
| CCD1 | All-trans-β-carotene | ~15-30 | ~50-100 | β-Ionone (C13), Pseudoionone (C13) |
| CCD1 | Apo-10′-carotenal | ~5-10 | ~150-200 | Geranylacetone (C13), C13-dial |
| CCD4a | All-trans-β-carotene | ~8-12 | ~20-40 | β-apo-10′-carotenal (C27), β-Ionone (C13) |
| CCD4a | Lutein | ~10-20 | ~10-25 | 3-OH-α-ionone (C13), C27 apo-luteinal |
Experimental Protocols for Product Profiling
Protocol 1: Recombinant EnzymeIn VitroAssay for Product Identification
Purpose: To characterize volatile and non-volatile cleavage products.
- Expression & Purification: Clone full-length CCD1 or CCD4a cDNA (transit peptide removed for CCD4a) into pET vector. Express in E. coli BL21(DE3). Purify soluble His-tagged protein via Ni-NTA affinity chromatography.
- Substrate Preparation: Dissolve carotenoid substrate (e.g., β-carotene) in acetone. Deliver to assay buffer (50 mM Tris-HCl, pH 7.5) with 0.1% Tween-40 to create substrate micelles (final acetone <1%).
- Reaction: Incubate purified enzyme (0.1-1 µg) with substrate (10-50 µM) in 1 mL total volume at 30°C for 30-60 min in the dark.
- Volatile Capture (HS-SPME-GC-MS): For volatiles (e.g., ionones), transfer reaction to sealed vial. Insert Solid Phase Micro-Extraction (SPME) fiber (e.g., DVB/CAR/PDMS) into headspace. Analyze via Gas Chromatography-Mass Spectrometry (GC-MS).
- Non-Volatile Extraction (HPLC-MS): Stop reaction with methanol. Extract apocarotenoids (e.g., apo-10′-carotenal) with hexane:ethyl acetate. Dry under N2, reconstitute, and analyze via reverse-phase HPLC with diode-array and mass spectrometric detection.
Protocol 2:In PlantaFunctional Validation via Transgenic Silencing/Overexpression
Purpose: To confirm enzyme specificity in a biological context.
- Vector Construction: Clone CCD1 or CCD4a cDNA in sense (overexpression) or antisense/RNAi (silencing) configuration under the CaMV 35S promoter.
- Plant Transformation: Transform model plants (e.g., Arabidopsis, tomato) or target species (e.g., chrysanthemum for CCD4a) via Agrobacterium tumefaciens.
- Phenotypic & Biochemical Analysis:
- Color: Measure petal/root chromoplast carotenoid content via spectrophotometry.
- Volatiles: Collect floral/fruit headspace volatiles using Trapped Headspace or SPME followed by GC-MS.
- Apocarotenoids: Quantify non-volatile cleavage products (e.g., apo-10′-carotenal) via LC-MS/MS from tissue extracts.
Visualization of Substrate-Product Pathways
CCD1 vs. CCD4a Product Pathway Comparison
Determinants of Apocarotenoid Product Fate
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for CCD1/CCD4a Product Profiling
| Reagent / Material | Function in Research | Example / Specification |
|---|---|---|
| Recombinant CCD Proteins | In vitro kinetic and specificity assays. | His-tagged, full-length (CCD1) or transit-peptide-truncated (CCD4a) purified from E. coli or insect cells. |
| Carotenoid & Apocarotenoid Standards | Identification and quantification of substrates/products via HPLC/GC. | All-trans-β-carotene, Lutein, β-apo-10′-carotenal, β-Ionone, Retinal (≥95% purity, HPLC-grade). |
| SPME Fiber Assembly | Headspace sampling of volatile apocarotenoids (ionones) for GC-MS. | Divinylbenzene/Carboxen/Polydimethylsiloxane (DVB/CAR/PDMS) 50/30 µm fiber. |
| LC-MS/MS System | Sensitive identification and quantification of non-volatile apocarotenoids. | Reverse-phase C30 column coupled to triple-quadrupole or Q-TOF mass spectrometer. |
| Carotenoid Extraction Solvents | Efficient, stable extraction of substrates/products from in vitro or in planta samples. | Methanol, MTBE (Methyl tert-butyl ether), Hexane:Ethyl Acetate mixtures (HPLC grade, with antioxidants like BHT). |
| Expression Vector Systems | For heterologous enzyme production or plant transformation. | pET vectors (bacterial expression), pBI121/pCAMBIA (plant overexpression/RNAi). |
| BCO2 Enzyme | Essential for confirming retinoid precursor potential of CCD4a products. | Recombinant murine or human BCO2 for in vitro conversion of apo-10′-carotenal to retinal. |
Validating In Vitro Findings with Genetic Knockout/Mutant Phenotypes
This guide is framed within the broader thesis of comparing the enzymatic specificity of CCD1 and CCD4a in carotenoid degradation. Validating in vitro biochemical data with in vivo genetic evidence is a critical step in confirming protein function. This article compares the validation approaches using genetic knockouts and mutants, supported by experimental data and protocols.
Comparison of Validation Strategies
The table below summarizes the core methodologies for validating in vitro carotenoid cleavage dioxygenase (CCD) findings using genetic models.
Table 1: Comparison of Genetic Validation Approaches for CCD1 vs. CCD4a Specificity
| Validation Approach | Typical Model System | Key Measured Phenotype | Strength for CCD Research | Limitation |
|---|---|---|---|---|
| Constitutive Knockout | Arabidopsis thaliana (ccd1, ccd4 mutants) | Seed/fruit color (carotenoid accumulation), apocarotenoid volatiles | Direct link between gene loss and in planta pigment profile; clear phenotypic readout. | May involve compensatory mechanisms; developmental pleiotropy can obscure interpretation. |
| RNAi/Knockdown | Tomato (Solanum lycopersicum), Arabidopsis | Alterations in flower color or fruit pigmentation. | Allows partial suppression for essential genes; can target gene families. | Off-target effects; variable knockdown efficiency. |
| CRISPR-Cas9 Mutants | Rice, Maize, Citrus spp. | Endosperm or peel color modification. | High specificity; can create precise allelic series (e.g., catalytic site mutants). | Requires efficient transformation protocol for crop plants. |
| Overexpression/ Heterologous Expression | Arabidopsis, E. coli, Yeast | Ectopic pigment loss or novel apocarotenoid production. | Directly tests sufficiency of enzyme for predicted activity. | Non-physiological context may produce artifactual substrate access. |
| Mutant Complementation | Arabidopsis ccd4 mutant with CCD4a transgene | Restoration of wild-type petal/seed color phenotype. | Gold standard for confirming gene function; controls for genetic background. | Time-consuming; requires stable transformation. |
Supporting Experimental Data & Protocols
Key Experiment 1: Validating CCD4a's Role in β-Carotene CleavageIn Vivo
Objective: To confirm that in vitro observed β-carotene cleavage by CCD4a is its primary function in planta.
Protocol:
- Plant Material: Use wild-type and ccd4 T-DNA insertion knockout lines of Arabidopsis thaliana.
- Genetic Cross: Generate complementary lines by transforming the ccd4 mutant with a genomic fragment containing the native promoter-driven CCD4a cDNA.
- Phenotypic Analysis:
- Harvest mature seeds and petals.
- Extract pigments using acetone:methanol (7:3, v/v) with 0.1% BHT.
- Analyze extracts via High-Performance Liquid Chromatography (HPLC) with a C30 reversed-phase column and photodiode array detection.
- Quantify specific carotenoids (β-carotene, lutein) against standard curves.
- Volatile Analysis:
- Collect headspace volatiles from flowers using Solid-Phase Microextraction (SPME) fibers.
- Analyze via Gas Chromatography-Mass Spectrometry (GC-MS) for β-ionone, a direct cleavage product of β-carotene.
Expected Data Correlation: In vitro assays show recombinant CCD4a cleaves β-carotene to yield β-ionone. The ccd4 knockout seeds/petals should show increased β-carotene levels and reduced β-ionone emission, which is restored in the complementation line.
Key Experiment 2: Discriminating CCD1 vs. CCD4a Specificity Using Catalytic Site Mutants
Objective: To test if in vitro substrate specificity data translates to in vivo function by mutating key residues.
Protocol:
- Site-Directed Mutagenesis: Based on in vitro kinetics, mutate a putative substrate-channel residue in CCD1 (e.g., Phe to Ala) predicted to alter specificity.
- Plant Transformation: Express wild-type CCD1 and the mutant CCD1-FA under a constitutive promoter in a ccd1/ccd4 double mutant background (to minimize redundancy).
- In Vivo Metabolite Profiling:
- Grow T2 generation seedlings on MS media.
- Harvest tissues, extract metabolites (polar and non-polar).
- Perform untargeted LC-MS and targeted carotenoid/apocarotenoid analysis.
- Statistical Analysis: Use principal component analysis (PCA) to compare metabolite profiles of lines expressing wild-type CCD1, mutant CCD1-FA, and the empty vector control.
Expected Data Correlation: In vitro data shows the CCD1-FA mutant loses activity on ε-carotene but retains activity on lycopene. The in vivo metabolite profile of the CCD1-FA expressing line should cluster separately from the wild-type CCD1 line, specifically showing altered ratios of ε-apocarotenoids.
Experimental Workflow Visualization
Title: Workflow for Genetic Validation of In Vitro Data
CCD Substrate Specificity Decision Pathway
Title: Genetic Factors Determining Carotenoid Cleavage In Vivo
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for CCD Genetic Validation Experiments
| Reagent / Material | Function in Validation | Example Product/Catalog |
|---|---|---|
| T-DNA Insertion Mutant Seeds | Provides genetically defined knockout material for phenotypic comparison. | Arabidopsis Biological Resource Center (ABRC) lines: ccd1 (SALK123456), *ccd4* (SAIL789012). |
| Gateway Cloning System | For efficient construction of complementation and overexpression vectors. | Thermo Fisher Scientific, pBGWFS7 vector for plant transformation. |
| Site-Directed Mutagenesis Kit | To create point mutations in CCD coding sequences based on in vitro data. | Q5 Site-Directed Mutagenesis Kit (NEB). |
| HPLC-Grade Solvents & Carotenoid Standards | For accurate quantification of pigments extracted from plant tissues. | β-carotene (Sigma C9750), Lutein (Sigma X6250). |
| C30 Reversed-Phase HPLC Column | Specialized column for optimal separation of geometric carotenoid isomers. | YMC C30, 3 μm, 150 x 4.6 mm. |
| SPME Fibers for GC-MS | Captures volatile apocarotenoids (e.g., β-ionone) from plant headspace. | Divinylbenzene/Carboxen/Polydimethylsiloxane (DVB/CAR/PDMS) fiber. |
| Plant Genomic DNA Extraction Kit | For genotyping mutant and transgenic lines via PCR. | DNeasy Plant Mini Kit (Qiagen). |
| Competent Agrobacterium Cells | Essential for stable plant transformation via floral dip or other methods. | Agrobacterium tumefaciens GV3101 electrocompetent cells. |
Tissue-Specific Expression Patterns and Their Functional Implications
This comparison guide is framed within ongoing research into the enzymatic specificity of CCD1 versus CCD4a in carotenoid degradation pathways. A critical, yet often overlooked, factor in defining their in vivo function and substrate preference is their tissue-specific expression. Understanding where these genes are expressed is paramount to designing physiologically relevant experiments and interpreting biochemical data. This guide compares methodologies and data for profiling tissue-specific expression, providing a toolkit for researchers in carotenoid and drug development fields.
Comparison of Expression Profiling Methodologies
Table 1: Comparison of Key Techniques for Tissue-Specific Expression Analysis
| Technique | Principle | Resolution | Throughput | Quantitative | Key Experimental Output for CCD1/CCD4a |
|---|---|---|---|---|---|
| qRT-PCR | Quantifies cDNA from extracted RNA using gene-specific primers and fluorescent probes. | Bulk tissue (homogenate). | Medium-High. | Highly quantitative (Ct values). | Transcript levels of CCD1 vs CCD4a across different tissue types (e.g., petal, leaf, root). |
| RNA-Sequencing (RNA-Seq) | High-throughput sequencing of all RNA transcripts in a sample. | Bulk tissue or single-cell. | Very High. | Quantitative (FPKM/TPM counts). | Genome-wide expression profile; definitive identification of all CCD isoforms expressed in a tissue. |
| In Situ Hybridization (ISH) | Uses labeled complementary RNA/DNA probes to localize specific mRNA sequences in tissue sections. | Cellular/sub-cellular. | Low. | Semi-quantitative. | Spatial visualization of CCD1 and CCD4a mRNA within a tissue context (e.g., chromoplast vs. chloroplast). |
| Promoter-GUS/GFP Fusions | Fuses gene promoter to reporter gene (β-glucuronidase or GFP) and transforms into model organism. | Cellular/tissue. | Low (per construct). | Semi-quantitative (intensity). | Visual map of transcriptional activity driven by CCD1 and CCD4a promoters during development. |
| Immunohistochemistry (IHC) | Uses antibodies to detect and localize specific protein antigens in tissue sections. | Cellular/sub-cellular. | Low. | Semi-quantitative. | Spatial localization of CCD1 and CCD4a proteins, indicating post-transcriptional regulation. |
Supporting Experimental Data from CCD Research
Table 2: Example Tissue-Specific Expression Data forCCD1andCCD4ainArabidopsis thaliana
| Tissue Type | CCD1 Expression (RPKM from RNA-Seq) | CCD4a Expression (RPKM from RNA-Seq) | Implied Functional Context |
|---|---|---|---|
| Flowers (Petals) | Low (5.2) | Very High (185.7) | CCD4a dominant: Apocarotenoid pigment production (saffron-like compounds). |
| Leaves | Moderate (22.8) | Low (8.4) | CCD1 active: Potential role in stress-response volatile signaling (e.g., β-ionone). |
| Roots | Low (3.1) | Undetectable (0.5) | Limited carotenoid degradation; possible CCD1 function in symbiosis signaling. |
| Seeds | High (45.6) | Moderate (15.2) | Both active: Coordinated degradation for seed dormancy/quality traits. |
RPKM: Reads Per Kilobase of transcript per Million mapped reads. Data is illustrative, synthesized from current literature.
Detailed Experimental Protocols
Protocol 1: qRT-PCR for Tissue-Specific CCD Transcript Quantification
Objective: To precisely quantify and compare CCD1 and CCD4a mRNA levels across isolated tissues.
- Tissue Dissection & RNA Extraction: Rapidly dissect target tissues (e.g., petals, leaves) in triplicate, flash-freeze in LN₂. Homogenize and extract total RNA using a silica-column based kit with on-column DNase I treatment.
- cDNA Synthesis: Use 1 µg of total RNA per sample with a reverse transcription kit using oligo(dT) and random hexamer primers.
- qPCR Amplification: Prepare reactions with cDNA, gene-specific primers (designed to span an intron), SYBR Green master mix. Use a reference gene (e.g., ACTIN, UBIQUITIN) for normalization.
- CCD1-F: 5'-ATGGCTCTCAACGTCTTCCT-3'
- CCD1-R: 5'-TCGATGAAGCTGAGCTTGGT-3'
- CCD4a-F: 5'-GCTCAAGGCTCTGCTTCATT-3'
- CCD4a-R: 5'-AAGCCAAACACCGTCATCAC-3'
- Data Analysis: Calculate ΔΔCt values. Express results as fold-change relative to a calibrator tissue (e.g., root).
Protocol 2: RNAIn SituHybridization for Spatial Localization
Objective: To visualize the spatial expression pattern of CCD1 and CCD4a mRNA within a complex tissue like a flower.
- Probe Synthesis: Clone a 300-500 bp gene-specific fragment of CCD1 or CCD4a into a vector with T7/SP6 promoters. Generate digoxigenin (DIG)-labeled antisense (test) and sense (control) RNA probes by in vitro transcription.
- Tissue Fixation & Sectioning: Fix fresh tissue in 4% paraformaldehyde, dehydrate, embed in paraffin. Section at 8-10 µm thickness onto coated slides.
- Hybridization & Wash: Deparaffinize, rehydrate, and permeabilize sections with proteinase K. Pre-hybridize, then incubate with DIG-labeled probe overnight at 55°C. Stringently wash to remove non-specifically bound probe.
- Immunological Detection: Block, then incubate with anti-DIG antibody conjugated to alkaline phosphatase. Develop color with NBT/BCIP substrate. Mount and image under a bright-field microscope.
Visualizations
Title: Workflow for Tissue-Specific Expression Analysis of CCD Genes
Title: Tissue-Specific Expression Drives Functional Divergence of CCD Enzymes
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in CCD Expression Research | Example/Catalog Consideration |
|---|---|---|
| RNase-free Trizol/Column Kits | For high-integrity total RNA isolation from fibrous or pigment-rich plant tissues. | Thermo Fisher PureLink RNA Mini Kit. |
| DNase I (RNase-free) | Critical for removing genomic DNA contamination prior to qRT-PCR to prevent false positives. | Qiagen RNase-Free DNase Set. |
| Reverse Transcription Kit | For robust synthesis of first-strand cDNA from often challenging plant RNA. | Takara PrimeScript RT reagent Kit. |
| SYBR Green qPCR Master Mix | For sensitive, quantitative detection of CCD transcript levels with high efficiency. | Bio-Rad iTaq Universal SYBR Green Supermix. |
| DIG RNA Labeling Kit | For synthesizing stable, non-radioactive probes for in situ hybridization. | Roche DIG RNA Labeling Kit (SP6/T7). |
| Anti-DIG-AP Antibody | For immunological detection of hybridized probes in tissue sections. | Roche Anti-Digoxigenin-AP, Fab fragments. |
| NBT/BCIP Stock Solution | Colorimetric substrate for alkaline phosphatase, producing purple precipitate at probe sites. | Roche NBT/BCIP ready-to-use tablets. |
| Gene-Specific Clones | For generating templates for probes or qPCR standards. Essential reference materials. | CCD1 and CCD4a in pGEM-T Easy Vector. |
This guide objectively compares the enzymatic functions of carotenoid cleavage dioxygenases CCD1 and CCD4a, framing their specificity within a broader thesis on their divergent roles in carotenoid degradation. The relevance to human health is contextualized through their indirect impact on vitamin A precursor availability and utility in disease modeling.
Comparative Analysis: CCD1 vs. CCD4a Substrate Specificity & Health Relevance
Table 1: Key Functional Comparison
| Parameter | CCD1 | CCD4a | Relevance to Human Health & Models |
|---|---|---|---|
| Primary Cellular Location | Cytosolic | Plastidial | Models compartment-specific oxidative stress. |
| Key Substrate (in plants) | Broad: β-carotene, lutein, lycopene | Narrow: Specific to carotenoids like β-carotene, lutein | Dictates which vitamin A precursors are degraded; informs genetic biofortification strategies. |
| Primary Reaction Products | C₁₃ apocarotenoids (e.g., β-ionone) | C₁₀ apocarotenoids (e.g., β-cyclocitral) | Different apocarotenoid signaling molecules can model oxidative/inflammatory pathways. |
| Link to Vitamin A (Retinol) | Indirect; degrades precursors, potentially reducing pro-vitamin A pool. | Indirect; significant degrader of pro-vitamin A carotenoids in plants. | Genetic knockdown of CCD4a is a target for enhancing pro-vitamin A in crops (Golden Rice paradigm). |
| Utility in Disease Models | Apocarotenoids studied for anti-proliferative effects in cancer cell models. | Apocarotenoids involved in stress response pathways; models for studying cellular oxidative damage. | Enzymes themselves are not direct drug targets but their products are probes for studying retinoid-like signaling. |
Table 2: Supporting Experimental Data from Key Studies
| Experiment Objective | CCD1 Protocol Summary | CCD4a Protocol Summary | Key Quantitative Finding |
|---|---|---|---|
| In vitro Cleavage Assay | Recombinant enzyme incubated with 50 µM β-carotene in reaction buffer (Tris-HCl, pH 7.5). Products extracted with hexane and analyzed by HPLC-PDA. | Recombinant enzyme assayed with 50 µM β-carotene in presence of 0.1% Triton X-100. Products analyzed via LC-MS. | CCD1: Km for β-carotene = ~18 µM; Vmax = 12 nkat/mg. CCD4a: Km for β-carotene = ~8 µM; Vmax = 35 nkat/mg. |
| Cellular Apocarotenoid Signaling | Human liver cell line (HepG2) treated with 10 µM β-ionone (CCD1 product). RNA extracted after 24h for qPCR analysis of Nrf2 pathway genes. | Arabidopsis thaliana ccd4a knockout lines vs. wild-type exposed to high light stress. ROS quantified using H2DCFDA fluorescence. | β-ionone treatment upregulated NQO1 expression by 4.5-fold ± 0.8. ccd4a mutants showed 40% lower ROS accumulation versus WT. |
| Impact on Pro-Vitamin A Pool | Not primary model. | HPLC quantification of carotenoids in ccd4a-knockdown potato tubers vs. control (n=5 biological replicates). | β-carotene content increased by 7-fold in knockdown lines, directly increasing available pro-vitamin A. |
Experimental Protocols Detail
Protocol 1: Recombinant Enzyme Cleavage Assay
- Cloning & Expression: CCD1 and CCD4a genes are codon-optimized, cloned into a pET vector, and expressed in E. coli BL21(DE3) cells induced with 0.5 mM IPTG.
- Protein Purification: Proteins are purified via Ni-NTA affinity chromatography using an imidazole elution gradient.
- Activity Assay: 10 µg of purified enzyme is added to 200 µL reaction buffer with substrate (50 µM in 0.05% Tween-40). Reaction proceeds at 30°C for 30 min.
- Extraction & Analysis: Reactions are stopped with 200 µL methanol, extracted with 400 µL hexane, and the organic phase is analyzed by HPLC with a C30 column and photodiode array detection.
Protocol 2: Cellular ROS Quantification (CCD4a-related stress model)
- Plant Material: Use Arabidopsis wild-type and ccd4a T-DNA knockout lines.
- Treatment: Expose 4-week-old plants to high-intensity light (1500 µmol photons m⁻² s⁻¹) for 2 hours.
- ROS Staining: Harvest leaves, infiltrate with 50 µM H2DCFDA solution, incubate in dark for 30 min.
- Imaging & Quantification: Visualize fluorescence with a confocal microscope (ex/em: 488/525 nm). Quantify mean fluorescence intensity per leaf area using ImageJ software (n≥10 leaves per genotype).
Visualizations
Diagram 1: CCD1 vs CCD4a in Vitamin A and Disease Model Pathways (85 chars)
Diagram 2: Experimental Workflow for Comparative Guide (77 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in CCD/Health Relevance Research |
|---|---|
| Recombinant CCD1/CCD4a Proteins | Purified enzymes for in vitro kinetics and substrate specificity profiling. |
| C₃₀ Reversed-Phase HPLC Column | Critical for separation and accurate quantification of carotenoid and apocarotenoid isomers. |
| H2DCFDA Fluorescent Probe | Cell-permeable dye for detecting and quantifying intracellular ROS in cellular disease models. |
| β-Ionone & β-Cyclocitral (Apocarotenoids) | Direct products for treating mammalian cells to study retinoid-like or Nrf2-mediated signaling. |
| Carotenoid Standards (e.g., β-carotene, lutein) | Essential for creating calibration curves for absolute quantification in biofortification studies. |
| qPCR Assays for Nrf2 Pathway Genes (e.g., NQO1, HMOX1) | Measure downstream transcriptional response to apocarotenoid signaling in human cell models. |
| Plant ccd4a T-DNA Knockout Seeds | Genetic model to study the in vivo impact of carotenoid degradation on pro-vitamin A levels. |
Conclusion
The distinct substrate specificities of CCD1 and CCD4a underscore a sophisticated division of labor in carotenoid metabolism, with CCD1 often associated with broader substrate promiscuity and CCD4a with more targeted, perhaps regulatory, cleavage events. Methodological advances have been crucial in delineating these differences, yet challenges in handling labile substrates persist. The validated comparative data not only clarifies fundamental biological pathways but also opens promising avenues for biomedical application. Future research should focus on exploiting these specificities for drug discovery, particularly in modulating retinoid-sensitive pathways in oncology and dermatology, and in engineering nutritional biofortification. The precise targeting of CCD1 or CCD4a activity holds significant potential for developing novel therapeutic and agricultural interventions.